Abstract
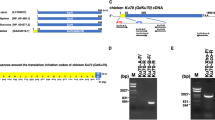
B-Myb is a highly conserved vertebrate member of the Myb transcription factor family, which is expressed in virtually all proliferating cells. A large body of evidence suggests that B-Myb plays an important role in cell cycle regulation; however, the exact nature of its function has not yet been clarified. We have used gene targeting in chicken DT40 cells, a cell line exhibiting very high rates of homologous recombination, to create cells expressing endogenous B-myb in a doxycyclin-dependent manner. We find that the cells proliferate well in the absence of B-Myb, suggesting that B-Myb is not essential for cell proliferation per se. However, cells lacking B-Myb are more sensitive to DNA-damage induced by UV-irradiation and alkylation. Our work provides the first direct evidence for a novel function of B-Myb in the response to DNA-damage. The cells described here should be a useful model to characterize this function in more detail.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Appl H and Klempnauer K-H . (2002). Oncogene, 21, 3076–3081.
Beall EL, Manak JR, Zhou S, Bell M, Lipsick JS and Botchan MR . (2002). Nature, 420, 833–837.
Buerstedde JM and Takeda S . (1991). Cell, 67, 179–188.
Cervellera M, Raschella G, Santilli G, Tanno B, Ventura A, Mancini C, Sevignani C, Calabretta B and Sala A . (2000). J. Biol. Chem., 275, 21055–21060.
Cervellera MN and Sala A . (2000). J. Biol. Chem., 275, 10692–10696.
Davidson C, Tirouvanziam R, Herzenberg L and Lipsick J . (2004). Genetics, 169, 215–229.
Fitzpatrick CA, Sharkov NV, Ramsay G and Katzen AL . (2002). Development, 129, 4497–4507.
Foos G, Grimm S and Klempnauer K-H . (1992). EMBO J., 11, 4619–4629.
Foos G, Grimm S and Klempnauer K-H . (1994). Oncogene, 9, 2481–2488.
Grassilli E, Salomoni P, Perrotti D, Franceschi C and Calabretta B . (1999). Cancer Res., 59, 2451–2456.
Huber A, Bai P, de Murcia JM and de Murcia G . (2004). DNA Repair, 3, 1103–1108.
Joaquin M, Bessa M, Saville MK and Watson RJ . (2002). Oncogene, 21, 7923–7932.
Joaquin M and Watson RJ . (2003). Cell Mol. Life Sci., 60, 2389–2401.
Kamano H, Burk B, Noben-Trauth K and Klempnauer K-H . (1995). Oncogene, 11, 2575–2582.
Katzen AL, Jackson J, Harmon BP, Fung SM, Ramsay G and Bishop JM . (1998). Genes Dev., 12, 831–843.
Klempnauer K-H and Bishop JM . (1983). J. Virol., 4, 565–572.
Lam EW-F and Watson RJ . (1993). EMBO J., 12, 2705–2713.
Lane S, Farlie P and Watson R . (1997). Oncogene, 14, 2445–2453.
Lang D, Dohle F, Terstesse M, Bangen P, August C, Pauels HG and Heidenreich S . (2002). J. Immunol., 168, 6152–6158.
Lang G, Gombert WM and Gould HJ . (2005). Immunology, 114, 25–36.
Manak JR, Mitiku N and Lipsick JS . (2002). Proc. Natl. Acad. Sci. USA, 99, 7438–7443.
Mucenski ML, McLain K, Kier AB, Swerdlow SH, Schreiner CM, Miller TA, Pietryga DW, Scott Jr WJ and Potter SS . (1991). Cell, 65, 677–689.
Neiman PE, Ruddell A, Jasoni C, Loring G, Thomas SJ, Brandvold KA, Lee R, Burnside J and Delrow J . (2001). Proc. Natl. Acad. Sci. USA, 98, 6378–6383.
Nyberg KA, Michelson RJ, Putnam CW and Weinert TA . (2002). Annu. Rev. Genet., 36, 617–656.
Sitzmann J, Noben-Trauth K, Kamano H and Klempnauer K-H . (1996). Oncogene, 12, 1889–1894.
Tanaka Y, Patestos NP, Maekawa T and Ishii S . (1999). J. Biol. Chem., 274, 28067–28070.
Toscani A, Mettus RV, Coupland R, Simpkins H, Litvin J, Orth J, Hatton KS and Reddy EP . (1997). Nature, 386, 713–717.
Zhu W, Giangrande PH and Nevins JR . (2004). EMBO J., 23, 4615–4626.
Ziebold U, Bartsch O, Marais R, Ferrari S and Klempnauer K-H . (1997). Curr. Biol., 7, 253–260.
Acknowledgements
We thank D Wenning for excellent technical assistance and J Delrow and the staff of the microarray facility of the Fred Hutchinson Cancer Research Center for performing the microarray experiments. This work was supported by the DFG (KL461/9-2).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ahlbory, D., Appl, H., Lang, D. et al. Disruption of B-myb in DT40 cells reveals novel function for B-Myb in the response to DNA-damage. Oncogene 24, 7127–7134 (2005). https://doi.org/10.1038/sj.onc.1208869
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1208869
Keywords
This article is cited by
-
Characterization of the zinc finger proteins ZMYM2 and ZMYM4 as novel B-MYB binding proteins
Scientific Reports (2020)
-
MYBL2 (B-Myb): a central regulator of cell proliferation, cell survival and differentiation involved in tumorigenesis
Cell Death & Disease (2017)
-
Interplay with the Mre11-Rad50-Nbs1 complex and phosphorylation by GSK3β implicate human B-Myb in DNA-damage signaling
Scientific Reports (2017)
-
B-Myb switches from Cyclin/Cdk-dependent to Jnk- and p38 kinase-dependent phosphorylation and associates with SC35 bodies after UV stress
Cell Death & Disease (2013)
-
Isolation and functional assessment of common, polymorphic variants of the B-MYB proto-oncogene associated with a reduced cancer risk
Oncogene (2008)