Abstract
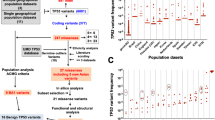
Last decade has led to the accumulation of large amounts of data on cancer genetics, opening an unprecedented access to the mapping of cancer genes in the human genome. Single-nucleotide polymorphisms (SNPs), the most common form of DNA variation in humans, emerge as an invaluable tool for cancer association studies. These genotypic markers can be used to assay how alleles of candidate genes correlate with the malignant phenotype, and may provide new clues into the genetic modifications that characterize cancer onset. In this cancer-oriented study, we detail an SNP mining strategy based on the analysis of expressed sequence tags among publicly available databases. Our whole-genome approach provides a comprehensive and unbiased description of nonsynonymous SNPs (nsSNPs) in tumoral versus normal tissues. To gain further insights into the possible relationships between genetic variation and altered phenotype, locations of a subset of nsSNPs were mapped onto protein domains known to be critical for protein function. Computational methods were also used to predict the potential impact of these cancer-associated nsSNPs on protein structure and function. We illustrate our approach through the detailed biochemical and structural characterization of a previously unknown cancer-associated mutation (G79C) affecting the 8 kDa dynein light chain (DNCL1).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W and Lipman DJ . (1997). Nucleic Acids Res., 25, 3389–3402.
Ayer DE, Kretzner L and Eisenman RN . (1993). Cell, 72, 211–222.
Bassen R, Brichory F, Caulet-Maugendre S, Delaval P and Dazord L . (2000). Bull. Cancer, 87, 703–707.
Boon K, Caron HN, van Asperen R, Valentijn L, Hermus MC, van Sluis P, Roobeek I, Weis I, Voute PA, Schwab M and Versteeg R . (2001). EMBO J., 20, 1383–1393.
Buetow KH, Edmonson M, MacDonald R, Clifford R, Yip P, Kelley J, Little DP, Strausberg R, Koester H, Cantor CR and Braun A . (2001). Proc. Natl. Acad. Sci. USA, 98, 581–584.
Cargill M, Altshuler D, Ireland J, Sklar P, Ardlie K, Patil N, Shaw N, Lane CR, Lim EP, Kalyanaraman N, Nemesh J, Ziaugra L, Friedland L, Rolfe A, Warrington J, Lipshutz R, Daley GQ and Lander ES . (1999). Nat. Genet., 22, 231–238.
Cerni C, Skrzypek B, Popov N, Sasgary S, Schmidt G, Larsson LG, Luscher B and Henriksson M . (2002). Oncogene, 21, 447–459.
Chakravarti A . (1998). Nat. Genet., 19, 216–217.
Chasman D and Adams RM . (2001). J. Mol. Biol., 307, 683–706.
Coller HA, Grandori C, Tamayo P, Colbert T, Lander ES, Eisenman RN and Golub TR . (2000). Proc. Natl. Acad. Sci. USA, 97, 3260–3265.
Collins FS, Guyer MS and Charkravarti A . (1997). Science, 278, 1580–1581.
Dick T, Ray K, Salz HK and Chia W . (1996). Mol. Cell Biol., 16, 1966–1977.
Draptchinskaia N, Gustavsson P, Andersson B, Pettersson M, Willig TN, Dianzani I, Ball S, Tchernia G, Klar J, Matsson H, Tentler D, Mohandas N, Carlsson B and Dahl N . (1999). Nat. Genet., 21, 169–175.
Fan J, Zhang Q, Tochio H, Li M and Zhang M . (2001). J. Mol. Biol., 306, 97–108.
Fan JS, Zhang Q, Li M, Tochio H, Yamazaki T, Shimizu M and Zhang M . (1998). J. Biol. Chem., 273, 33472–33481.
Farmer II BT, Constantine KL, Goldfarb V, Friedrichs MS, Wittekind M, Yanchunas Jr J, Robertson JG and Mueller L . (1996). Nat. Struct. Biol., 3, 995–997.
Fleming MA, Potter JD, Ramirez CJ, Ostrander GK and Ostrander EA . (2003). Proc. Natl. Acad. Sci. USA, 100, 1151–1156.
Fuhrmann JC, Kins S, Rostaing P, El Far O, Kirsch J, Sheng M, Triller A, Betz H and Kneussel M . (2002). J. Neurosci., 22, 5393–5402.
Gorelik E, Galili U and Raz A . (2001). Cancer Metastasis Rev., 20, 245–277.
Gouy M, Gautier C, Attimonelli M, Lanave C and di Paola G . (1985). Comput. Appl. Biosci., 1, 167–172.
Hanahan D and Weinberg RA . (2000). Cell, 100, 57–70.
Iida A, Saito S, Sekine A, Mishima C, Kitamura Y, Kondo K, Harigae S, Osawa S and Nakamura Y . (2002). J. Hum. Genet., 47, 285–310.
Imyanitov EN, Togo AV and Hanson KP . (2004). Cancer Lett., 204, 3–14.
Jaffrey SR and Snyder SH . (1996). Science, 274, 774–777.
Jin H and Varner J . (2004). Br. J. Cancer, 90, 561–565.
Johnson JP . (1999). Cancer Metast. Rev., 18, 345–357.
Liang J, Jaffrey SR, Guo W, Snyder SH and Clardy J . (1999). Nat. Struct. Biol., 6, 735–740.
Lo KW, Naisbitt S, Fan JS, Sheng M and Zhang M . (2001). J. Biol. Chem., 276, 14059–14066.
Menssen A and Hermeking H . (2002). Proc. Natl. Acad. Sci. USA, 99, 6274–6279.
Mohrenweiser HW, Xi T, Vazquez-Matias J and Jones IM . (2002). Cancer Epidemiol Biomarkers Prev., 11, 1054–1064.
Mulder NJ, Apweiler R, Attwood TK, Bairoch A, Barrell D, Bateman A, Binns D, Biswas M, Bradley P, Bork P, Bucher P, Copley RR, Courcelle E, Das U, Durbin R, Falquet L, Fleischmann W, Griffiths-Jones S, Haft D, Harte N, Hulo N, Kahn D, Kanapin A, Krestyaninova M, Lopez R, Letunic I, Lonsdale D, Silventoinen V, Orchard SE, Pagni M, Peyruc D, Ponting CP, Selengut JD, Servant F, Sigrist CJ, Vaughan R and Zdobnov EM . (2003). Nucleic Acids Res., 31, 315–318.
Naisbitt S, Valtschanoff J, Allison DW, Sala C, Kim E, Craig AM, Weinberg RJ and Sheng M . (2000). J. Neurosci., 20, 4524–4534.
Naora H, Takai I and Adachi M . (1998). J. Cell Biol., 141, 741–753.
Ng PC and Henikoff S . (2002). Genome Res., 12, 436–446.
Ng PC and Henikoff S . (2003). Nucleic Acids Res., 31, 3812–3814.
Nomoto S, Haruki N, Takahashi T, Masuda A, Koshikawa T, Fujii Y and Osada H . (1999). Oncogene, 18, 7180–7183.
Phillis R, Statton D, Caruccio P and Murphey RK . (1996). Development, 122, 2955–2963.
Poteete AR, Rennell D and Bouvier SE . (1992). Proteins, 13, 38–40.
Puthalakath H, Huang DC, O'Reilly LA, King SM and Strasser A . (1999). Mol. Cell, 3, 287–296.
Puthalakath H and Strasser A . (2002). Cell Death Differ., 9, 505–512.
Puthalakath H, Villunger A, O'Reilly LA, Beaumont JG, Coultas L, Cheney RE, Huang DC and Strasser A . (2001). Science, 293, 1829–1832.
Qiu P, Wang L, Kostich M, Ding W, Simon JS and Greene JR . (2004). BMC Cancer, 4, 4.
Ramensky V, Bork P and Sunyaev S . (2002). Nucleic Acids Res., 30, 3894–3900.
Reich DE, Gabriel SB and Altshuler D . (2003). Nat. Genet., 33, 457–458.
Rosenwald IB . (1996). Cancer Lett., 102, 113–123.
Rosenwald IB, Rhoads DB, Callanan LD, Isselbacher KJ and Schmidt EV . (1993). Proc. Natl. Acad. Sci. USA, 90, 6175–6178.
Roussel MF, Ashmun RA, Sherr CJ, Eisenman RN and Ayer DE . (1996). Mol. Cell Biol., 16, 2796–2801.
Ruggero D and Pandolfi PP . (2003). Nat. Rev. Cancer, 3, 179–192.
Sachidanandam R, Weissman D, Schmidt SC, Kakol JM, Stein LD, Marth G, Sherry S, Mullikin JC, Mortimore BJ, Willey DL, Hunt SE, Cole CG, Coggill PC, Rice CM, Ning Z, Rogers J, Bentley DR, Kwok PY, Mardis ER, Yeh RT, Schultz B, Cook L, Davenport R, Dante M, Fulton L, Hillier L, Waterston RH, McPherson JD, Gilman B, Schaffner S, Van Etten WJ, Reich D, Higgins J, Daly MJ, Blumenstiel B, Baldwin J, Stange-Thomann N, Zody MC, Linton L, Lander ES and Altshuler D . (2001). Nature, 409, 928–933.
Sass PM . (1998). Cancer Invest., 16, 322–328.
Schnorrer F, Bohmann K and Nusslein-Volhard C . (2000). Nat. Cell Biol., 2, 185–190.
Stitziel NO, Tseng YY, Pervouchine D, Goddeau D, Kasif S and Liang J . (2003). J. Mol. Biol., 327, 1021–1030.
Strausberg RL, Simpson AJ and Wooster R . (2003). Nat. Rev. Genet., 4, 409–418.
Sunyaev S, Lathe III W and Bork P . (2001a). Curr. Opin. Struct. Biol., 11, 125–130.
Sunyaev S, Ramensky V and Bork P . (2000). Trends Genet., 16, 198–200.
Sunyaev S, Ramensky V, Koch I, Lathe III W, Kondrashov AS and Bork P . (2001b). Hum. Mol. Genet., 10, 591–597.
Syvanen AC, Landegren U, Isaksson A, Gyllensten U and Brookes A . (1999). Eur. J. Hum. Genet., 7, 98–101.
Turk V, Kos J and Turk B . (2004). Cancer Cell, 5, 409–410.
Vadlamudi RK, Bagheri-Yarmand R, Yang Z, Balasenthil S, Nguyen D, Sahin AA, den Hollander P and Kumar R . (2004). Cancer Cell, 5, 575–585.
Wall SJ, Jiang Y, Muschel RJ and DeClerck YA . (2003). Cancer Res., 63, 4750–4755.
Wang W, Lo KW, Kan HM, Fan JS and Zhang M . (2003). J. Biol. Chem., 278, 41491–41499.
Wang Z and Moult J . (2001). Hum. Mutat., 17, 263–270.
Xi T, Jones IM and Mohrenweiser HW . (2004). Genomics, 83, 970–979.
Acknowledgements
We thank Pr Germain Gillet, Adel Khelifi and Cristina Vieira-Heddi for their valuable comments. VN is supported by a grant from INRA. AA is recipient of a fellowship from the ARC. Research in MZ's laboratory was supported by grants from the Research Grants Council of Hong Kong. MZ is a Croucher Foundation Senior Research Fellow.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on Oncogene website (http://www.nature.com/onc)
Supplementary information
Rights and permissions
About this article
Cite this article
Aouacheria, A., Navratil, V., Wen, W. et al. In silico whole-genome scanning of cancer-associated nonsynonymous SNPs and molecular characterization of a dynein light chain tumour variant. Oncogene 24, 6133–6142 (2005). https://doi.org/10.1038/sj.onc.1208745
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1208745
Keywords
This article is cited by
-
Dynein light chain 1 functions in somatic cyst cells regulate spermatogonial divisions in Drosophila
Scientific Reports (2011)
-
Predicting cancer involvement of genes from heterogeneous data
BMC Bioinformatics (2008)
-
In silico whole-genome screening for cancer-related single-nucleotide polymorphisms located in human mRNA untranslated regions
BMC Genomics (2007)
-
Mining expressed sequence tags identifies cancer markers of clinical interest
BMC Bioinformatics (2006)
-
Bioinformatic screening of human ESTs for differentially expressed genes in normal and tumor tissues
BMC Genomics (2006)