Abstract
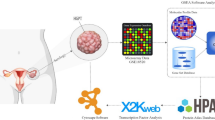
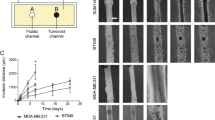
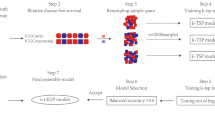
Epithelial ovarian cancer (EOC) is the gynecological disease with the highest death rate. We applied an automatic class discovery procedure based on gene expression profiling to stages III–IV tumors to search for molecular signatures associated with the biological properties and progression of EOC. Using a complementary DNA microarray containing 4451 cancer-related, sequence-verified features, we identified a subset of EOC characterized by the expression of numerous genes related to the extracellular matrix (ECM) and its remodeling, along with elements of the fibroblast growth factor 2 (FGF2) signaling pathway. A total of 10 genes were validated by quantitative real-time polymerase chain reaction, and coexpression of FGF2 and fibroblast growth factor receptor 4 in tumor cells was revealed by immunohistochemistry, confirming the reliability of gene expression by cDNA microarray. Since the functional relationships among these genes clearly suggested involvement of the identified molecular signature in processes related to epithelial–stromal interactions and/or epithelial–mesenchymal cellular plasticity, we applied supervised learning analysis on ovarian-derived cell lines showing distinct cellular phenotypes in culture. This procedure enabled construction of a gene classifier able to discriminate mesenchymal-like from epithelial-like cells. Genes overexpressed in mesenchymal-like cells proved to match the FGF2 signaling and ECM molecular signature, as identified by unsupervised class discovery on advanced tumor samples. In vitro functional analysis of the cell plasticity classifier was carried out using two isogenic and immortalized cell lines derived from ovarian surface epithelium and displaying mesenchymal and epithelial morphology, respectively. The results indicated the autocrine, but not intracrine stimulation of mesenchymal conversion and cohort/scatter migration of cells by FGF2, suggesting a central role for FGF2 signaling in the maintenance of cellular plasticity of ovary-derived cells throughout the carcinogenesis process. These findings raise mechanistic hypotheses on EOC pathogenesis and progression that might provide a rational underpinning for new therapeutic modalities.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Auersperg N, Maines-Bandiera SL, Dyck HG and Kruk PA . (1994). Lab. Invest., 71, 510–518.
Auersperg N, Wong AS, Choi KC, Kang SK and Leung PC . (2001). Endocr. Rev., 22, 255–288.
Aunoble B, Sanches R, Didier E and Bignon YJ . (2000). Int. J. Oncol., 16, 567–576.
Balli S, Fey MF, Hanggi W, Zwahlen D, Berclaz G, Dreher E and Aebi S . (2000). Eur. J. Cancer, 36, 2061–2068.
Bertrand V, Hudson C, Caillol D, Popovici C and Lemaire P . (2003). Cell, 115, 615–627.
Crickard K, Gross JL, Crickard U, Yoonessi M, Lele S, Herblin WF and Eidsvoog K . (1994). Gynecol. Oncol., 55, 277–284.
Darai E, Scoazec JY, Walker-Combrouze F, Mlika-Cabanne N, Feldmann G, Madelenat P and Potet F . (1997). Hum. Pathol., 28, 922–928.
Davies BR, Worsley SD and Ponder BA . (1998). Histopathology, 32, 69–80.
Di Blasio AM, Carniti C, Vigano P and Vignali M . (1995). J. Steroid. Biochem. Mol. Biol., 53, 375–379.
Dyck HG, Hamilton TC, Godwin AK, Lynch HT, Maines-Bandiera SL and Auersperg N . (1996). Int. J. Cancer, 69, 429–436.
Dysvik B and Jonassen I . (2001). Bioinformatics, 17, 369–370.
Feeley KM and Wells M . (2001). Histopathology, 38, 87–95.
Fujimoto J, Ichigo S, Hori M, Hirose R, Sakaguchi H and Tamaya T . (1997). Eur. J. Gynaecol. Oncol., 18, 349–352.
Greenlee RT, Hill-Harmon MB, Murray T and Thun M . (2001). CA Cancer J. Clin., 51, 15–36.
Holschneider CH and Berek JS . (2000). Semin. Surg. Oncol., 19, 3–10.
Hori A, Sasada R, Matsutani E, Naito K, Sakura Y, Fujita T and Kozai Y . (1991). Cancer Res., 51, 6180–6184.
Hosack DA, Dennis Jr G, Sherman BT, Lane HC and Lempicki RA . (2003). Genome Biol., 4, R70.
Hough CD, Cho KR, Zonderman AB, Schwartz DR and Morin PJ . (2001). Cancer Res., 61, 3869–3876.
Hough CD, Sherman-Baust CA, Pizer ES, Montz FJ, Im DD, Rosenshein NB, Cho KR, Riggins GJ and Morin PJ . (2000). Cancer Res., 60, 6281–6287.
Janda E, Lehmann K, Killisch I, Jechlinger M, Herzig M, Downward J, Beug H and Grunert S . (2002). J. Cell Biol., 156, 299–313.
Jazaeri AA, Yee CJ, Sotiriou C, Brantley KR, Boyd J and Liu ET . (2002). J. Natl. Cancer Inst., 94, 990–1000.
Karavanova ID, Dove LF, Resau JH and Perantoni AO . (1996). Development, 122, 4159–4167.
Kinsella MG, Tsoi CK, Jarvelainen HT and Wight TN . (1997). J. Biol. Chem., 272, 318–325.
Kodama J, Hashimoto I, Seki N, Hongo A, Yoshinouchi M, Okuda H and Kudo T . (2001). Anticancer Res., 21, 2983–2987.
Kruk PA, Maines-Bandiera SL and Auersperg N . (1990). Lab. Invest., 63, 132–136.
Kruk PA, Uitto VJ, Firth JD, Dedhar S and Auersperg N . (1994). Exp. Cell Res., 215, 97–108.
Liotta LA and Kohn EC . (2002). J. Natl. Cancer Inst., 94, 1113–1114.
Long CJ, Roth MR, Tasheva ES, Funderburgh M, Smit R, Conrad GW and Funderburgh JL . (2000). J. Biol. Chem., 275, 13918–13923.
Maines-Bandiera SL and Auersperg N . (1997). Int. J. Gynecol. Pathol., 16, 250–255.
Matei D, Graeber TG, Baldwin RL, Karlan BY, Rao J and Chang DD . (2002). Oncogene, 21, 6289–6298.
Matias-Guiu X and Prat J . (1998). Virchows Arch., 433, 103–111.
McShane LM, Radmacher MD, Freidlin B, Yu R, Li MC and Simon R . (2002). Bioinformatics, 18, 1462–1469.
Murdoch AD, Dodge GR, Cohen I, Tuan RS and Iozzo RV . (1992). J. Biol. Chem., 267, 8544–8557.
Obermair A, Speiser P, Reisenberger K, Ullrich R, Czerwenka K, Kaider A, Zeillinger R and Miksche M . (1998). Cancer Lett., 130, 69–76.
Ornitz DM and Marie PJ . (2002). Genes Dev., 16, 1446–1465.
Parrott JA, Nilsson E, Mosher R, Magrane G, Albertson D, Pinkel D, Gray JW and Skinner MK . (2001). Mol. Cell. Endocrinol., 175, 29–39.
Perantoni AO, Dove LF and Karavanova I . (1995). Proc. Natl. Acad. Sci. USA, 92, 4696–4700.
Plotnikov AN, Hubbard SR, Schlessinger J and Mohammadi M . (2000). Cell, 101, 413–424.
Plotnikov AN, Schlessinger J, Hubbard SR and Mohammadi M . (1999). Cell, 98, 641–650.
Powers CJ, McLeskey SW and Wellstein A . (2000). Endocr. Relat. Cancer, 7, 165–197.
Radmacher MD, McShane LM and Simon R . (2002). J. Comput. Biol., 9, 505–511.
Righetti SC, Perego P, Corna E, Pierotti MA and Zunino F . (1999). Cell Growth Differ., 10, 473–478.
Santala M, Simojoki M, Risteli J, Risteli L and Kauppila A . (1999). Clin. Cancer Res., 5, 4091–4096.
Schafer T, Zentgraf H, Zehe C, Brugger B, Bernhagen J and Nickel W . (2004). J. Biol. Chem., 279, 6244–6251.
Schaner ME, Ross DT, Ciaravino G, Sorlie T, Troyanskaya O, Diehn M, Wang YC, Duran GE, Sikic TL, Caldeira S, Skomedal H, Tu IP, Hernandez-Boussard T, Johnson SW, O’Dwyer PJ, Fero MJ, Kristensen GB, Borresen-Dale AL, Hastie T, Tibshirani R, van de RM, Teng NN, Longacre TA, Botstein D, Brown PO and Sikic BI . (2003). Mol. Biol. Cell, 14, 4376–4386.
Schlessinger J, Plotnikov AN, Ibrahimi OA, Eliseenkova AV, Yeh BK, Yayon A, Linhardt RJ and Mohammadi M . (2000). Mol. Cell, 6, 743–750.
Schwartz DR, Kardia SL, Shedden KA, Kuick R, Michailidis G, Taylor JM, Misek DE, Wu R, Zhai Y, Darrah DM, Reed H, Ellenson LH, Giordano TJ, Fearon ER, Hanash SM and Cho KR . (2002). Cancer Res., 62, 4722–4729.
Schwartz DR, Wu R, Kardia SL, Levin AM, Huang CC, Shedden KA, Kuick R, Misek DE, Hanash SM, Taylor JM, Reed H, Hendrix N, Zhai Y, Fearon ER and Cho KR . (2003). Cancer Res., 63, 2913–2922.
Sheng G, dos RM and Stern CD . (2003). Cell, 115, 603–613.
Sherman-Baust CA, Weeraratna AT, Rangel LB, Pizer ES, Cho KR, Schwartz DR, Shock T and Morin PJ . (2003). Cancer Cell, 3, 377–386.
Shridhar V, Lee J, Pandita A, Iturria S, Avula R, Staub J, Morrissey M, Calhoun E, Sen A, Kalli K, Keeney G, Roche P, Cliby W, Lu K, Schmandt R, Mills GB, Bast RCJ, James CD, Couch FJ, Hartmann LC, Lillie J and Smith DI . (2001). Cancer Res., 61, 5895–5904.
Simojoki M, Santala M, Risteli J and Kauppila A . (2001). Gynecol. Oncol., 82, 110–115.
Steele IA, Edmondson RJ, Bulmer JN, Bolger BS, Leung HY and Davies BR . (2001). Oncogene, 20, 5878–5887.
Strutz F, Zeisberg M, Ziyadeh FN, Yang CQ, Kalluri R, Muller GA and Neilson EG . (2002). Kidney Int., 61, 1714–1728.
Sundfeldt K, Piontkewitz Y, Ivarsson K, Nilsson O, Hellberg P, Brännström M, Janson PO, Enerbäck S and Hedin L . (1997). Int. J. Cancer, 74, 275–280.
Taverna S, Ghersi G, Ginestra A, Rigogliuso S, Pecorella S, Alaimo G, Saladino F, Dolo V, Dell’Era P, Pavan A, Pizzolanti G, Mignatti P, Presta M and Vittorelli ML . (2003). J. Biol. Chem., 278, 51911–51919.
Thiery JP . (2002). Nat. Rev. Cancer, 2, 442–454.
Tonin PN, Hudson TJ, Rodier F, Bossolasco M, Lee PD, Novak J, Manderson EN, Provencher D and Mes-Masson AM . (2001). Oncogene, 20, 6617–6626.
Tsao SW, Mok CH, Knapp RC, Oike K, Muto MG, Welch WR, Goodman HM, Sheets EE, Berkowitz RS and Lau CC . (1993). Gynecol. Oncol., 48, 5–10.
Valve E, Martikainen P, Seppanen J, Oksjoki S, Hinkka S, Anttila L, Grenman S, Klemi P and Harkonen P . (2000). Int. J. Cancer, 88, 718–725.
Van Niekerk CC, Ramaekers FCS, Hanselaar AGJM, Aldeweireldt J and Poels LG . (1993). Am. J. Pathol., 142, 157–177.
von Heydebreck A, Huber W, Poustka A and Vingron M . (2001). Bioinformatics, 17, S107–S114.
Walsh FS and Doherty P . (1997). Annu. Rev. Cell Dev. Biol., 13, 425–456.
Welsh JB, Zarrinkar PP, Sapinoso LM, Kern SG, Behling CA, Monk BJ, Lockhart DJ, Burger RA and Hampton GM . (2001). Proc. Natl. Acad. Sci. USA, 98, 1176–1181.
Acknowledgements
Support for this project was provided to INT (Istituto Nazionale Tumori) – LNCIB (Laboratorio Nazionale CIB) by AIRC (Associazione Italiana Ricerca sul Cancro) Coordinated Project at IFOM (FIRC Institute for Molecular oncology) ‘Detection of cancer gene mutation and expression profiling by nanotechnology’ and by CNR/MIUR ‘Progetto strategico Oncologia’ (contract no. 02.00385.ST97 to MA Pierotti and no. 02.00386.ST97 to C Schneider) and grants to S Canevari from AIRC-FIRC and Special Project of Health Ministry.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Supplementary Information accompanies the paper on Oncogene website (http://www.nature.com/onc).
Supplementary information
Rights and permissions
About this article
Cite this article
Cecco, L., Marchionni, L., Gariboldi, M. et al. Gene expression profiling of advanced ovarian cancer: characterization of a molecular signature involving fibroblast growth factor 2. Oncogene 23, 8171–8183 (2004). https://doi.org/10.1038/sj.onc.1207979
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1207979
Keywords
This article is cited by
-
Ubiquitous Neural Cell Adhesion Molecule (NCAM): Potential Mechanism and Valorisation in Cancer Pathophysiology, Drug Targeting and Molecular Transductions
Molecular Neurobiology (2022)
-
Mitochondrial function — gatekeeper of intestinal epithelial cell homeostasis
Nature Reviews Gastroenterology & Hepatology (2018)
-
Multiple signatures of a disease in potential biomarker space: Getting the signatures consensus and identification of novel biomarkers
BMC Genomics (2015)
-
Identification of the IGF1/PI3K/NF κB/ERK gene signalling networks associated with chemotherapy resistance and treatment response in high-grade serous epithelial ovarian cancer
BMC Cancer (2013)
-
An IL6-correlated signature in serous epithelial ovarian cancer associates with growth factor response
BMC Genomics (2013)