Abstract
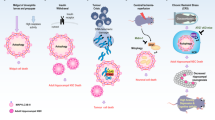
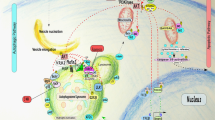
For many years apoptosis research has focused on caspases and their putative role as sole executioners of programmed cell death. Accumulating information now suggests that lysosomal cathepsins are also pivotally involved in this process, especially in pathological conditions. In particular, the role of lysosomes and lysosomal enzymes in initiation and execution of the apoptotic program has become clear in several models, to the point that the existence of a ‘lysosomal pathway of apoptosis’ is now generally accepted. This pathway of apoptosis can be activated by death receptors, lipid mediators, and photodamage. Lysosomal proteases can be released from the lysosomes into the cytosol, where they contribute to the apoptotic cascade upstream of mitochondria. This review focuses on the players and the molecular mechanisms involved in the lysosomal pathway of apoptosis as well as on the importance of this pathway in development and pathology.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Abbreviations
- CNS:
-
central nervous system
- EPM1:
-
progressive myoclonus epilepsy
- FAN:
-
factor associated with neutral sphingomyelinase
- PLA2:
-
phospholipase A2
- ROS:
-
reactive oxygen species
- TNF-α:
-
tumor necrosis factor-alpha
- TNF-R1:
-
tumor necrosis factor-receptor 1
- TRAIL:
-
tumor necrosis factor-related apoptosis-inducing ligand (TRAIL)
References
Antunes F, Cadenas E and Brunk UT . (2001). Biochem. J., 356, 549–555.
Ashkenazi A, Pai RC, Fong S, Leung S, Lawrence DA, Marsters SA, Blackie C, Chang L, McMurtrey AE, Hebert A, DeForge L, Koumenis IL, Lewis D, Harris L, Bussiere J, Koeppen H, Shahrokh Z and Schwall RH . (1999). J. Clin. Invest., 104, 155–162.
Bahr BA and Bendiske J . (2002). J. Neurochem., 83, 481–489.
Banay-Schwartz M, Dahl D, Hui KS and Lajtha A . (1987). Neurochem. Res., 12, 361–367.
Berg T, Gjoen T and Bakke O . (1995). Biochem. J., 307, 313–326.
Bidere N, Lorenzo HK, Carmona S, Laforge M, Harper F, Dumont C and Senik A . (2003). J. Biol. Chem., 278, 31401–31411.
Boya P, Andreau K, Poncet D, Zamzani N, Perfettini J-L, Metivier D, Ojcius DM, Jaattela M and Kroemer G . (2003a). J. Exp. Med., 197, 1323–1334.
Boya P, Gonzales-Polo R-A, Poncet D, Andreau K, Vieira HLA, Roumier T, Perfettini J-L and Kroemer G . (2003b). Oncogene, 22, 3927–3936.
Brunk UT and Svensson I . (1999). Redox Rep., 4, 3–11.
Bursch W . (2001). Cell Death Differ, 8, 569–581.
Campo E, Munoz J, Miquel R, Palacin A, Cardesa A, Sloane BF and Emmert-Buck MR . (1994). Am. J. Pathol., 145, 301–309.
Canbay A, Guicciardi ME, Higuchi H, Feldstein A, Bronk SF, Rydzewski R, Taniai M and Gores GJ . (2003). J. Clin. Invest., 112, 152–159.
Cantor AB and Kornfeld S . (1992). J. Biol. Chem., 267, 23357–23363.
Castiglioni T, Merino MJ, Elsner B, Lah TT, Sloane BF and Emmert-Buck MR . (1994). Hum. Pathol., 25, 857–862.
Chapman HA, Riese JP and Shi G-P . (1997). Annu. Rev. Physiol., 59, 63–88.
Chi S, Kitanaka C, Noguchi K, Mochizuki T, Nagashima Y, Shirouzu M, Fujita H, Yoshida M, Chen W, Asai A, Himeno M, Yokoyama S and Kuchino Y . (1999). Oncogene, 18, 2281–2290.
Cirman T, Oresic K, Droga-Mazovec G, Turk V, Reed JC, Myers RM, Salvesen GS and Turk B . (2004). J. Biol. Chem., 279, 3578–3587.
Claus V, Jahraus A, Tjelle T, Berg T, Kirschke H, Faulstich H and Griffiths GM . (1998). J. Biol. Chem., 273, 9842–9851.
Cuervo AM and Dice LF . (2000). Exp. Gerontol., 35, 119–131.
Dare E, Li W, Zhivotovsky B, Yuan X and Ceccatelli S . (2001). Free Radical Biol. Med., 30, 1347–1356.
Deiss LP, Galinka H, Berissi H, Cohen O and Kimchi A . (1996). EMBO J., 15, 3861–3870.
Demchik LL, Sameni M, Nelson K, Mikkelsen T and Sloane BF . (1999). Int. J. Dev. Neurosci., 17, 483–494.
Demoz M, Castino R, Cesaro P, Baccino FM, Bonelli G and Isidoro C . (2002). Biol. Chem., 383, 1237–1248.
Demoz M, Castino R, Dragonetti A, Raiteri E, Baccino FM and Isidoro C . (1999). J. Cell. Biochem., 73, 370–378.
Faubion WA, Guicciardi ME, Miyoshi H, Bronk SF, Roberts PJ, Svingen PA, Kaufmann SH and Gores GJ . (1999). J. Clin. Invest., 103, 137–145.
Felbor U, Kessler B, Mothes W, Goebel HH, Ploegh HL, Bronson RT and Olsen BR . (2002). Proc. Natl. Acad. Sci. USA, 99, 7883–7888.
Ferri KF and Kroemer G . (2001a). Nat. Cell Biol., 2, E63–E64.
Ferri KF and Kroemer G . (2001b). Nat. Cell Biol., 3, E255–63.
Foghsgaard L, Lademann U, Wissing D, Poulsen B and Jaattela M . (2002). J. Biol. Chem., 277, 39499–39506.
Foghsgaard L, Wissing D, Mauch D, Lademann U, Bastholm L, Boes M, Elling F, Leist M and Jaattela M . (2001). J. Cell Biol., 153, 999–1009.
Guicciardi ME, Deussing J, Miyoshi H, Bronk SF, Svingen PA, Peters C, Kaufmann SH and Gores GJ . (2000). J. Clin. Invest., 106, 1127–1137.
Guicciardi ME, Miyoshi H, Bronk SF and Gores GJ . (2001). Am. J. Pathol., 159, 2045–2054.
Halangk W, Lerch MM, Brandt-Nedelev B, Roth W, Ruthenbuerger M, Reinheckel T, Domschke W, Lippert H, Peters C and Deussing J . (2000). J. Clin. Invest., 106, 773–781.
Heinrich M, Wickel M, Schneider-Brachert W, Sandberg C, Gahr J, Schwandner R, Weber T, Brunner J, Kronke M and Schutze S . (1999). EMBO J., 18, 5252–5263.
Hishita T, Tada-Oikawa S, Tohyama K, Miura Y, Nishihara T, Tohyama Y, Yoshida Y, Uchiyama T and Kawanishi S . (2001). Cancer Res., 61, 2878–2884.
Houseweart MK, Pennacchio LA, Vilaythong A, Peters C, Noebels JL and Myers RM . (2003a). J. Neurobiol., 56, 315–327.
Houseweart MK, Vilaythong A, Yin XM, Turk B, Noebels JL and Myers RM . (2003b). Cell Death Differ, 10, 1329–1335.
Ishidoh K and Kominami E . (2002). Biol. Chem., 383, 1827–1831.
Ishisaka R, Kanno T, Akiyama J, Yoshioka T, Utsumi K and Utsumi T . (2001). J. Biochem., 129, 35–41.
Ishisaka R, Utsumi T, Kanno T, Arita K, Katunuma N, Akiyama J and Utsumi K . (1999). Cell Struct. Funct., 24, 465–470.
Jaattela M . (2002). Ann. Med., 34, 480–488.
Jaattela M, Benedict M, Tewari M, Shayman JA and Dixit VM . (1995). Oncogene, 10, 2297–2305.
Johansson A-C, Steen H, Ollinger K and Roberg K . (2003). Cell Death Differ., 10, 1253–1259.
Jones B, Roberts PJ, Faubion WA, Kominami E and Gores GJ . (1998). Am J Physiol, 275, G723–30.
Kagedal K, Johansson U and Ollinger K . (2001a). FASEB J, 15, 1592–1594.
Kagedal K, Zhao M, Svensson I and Brunk UT . (2001b). Biochem. J., 359, 335–343.
Katunuma N, Matsui A, Le QT, Utsumi K, Salvesen G and Ohashi A . (2001). Adv. Enzyme Regul., 41, 237–250.
Katunuma N, Matsunaga Y, Himeno K and Hayashi Y . (2003). Biol. Chem., 384, 883–890.
Koblinski JE, Ahram M and Sloane BF . (2000). Clin. Chim. Acta, 291, 113–135.
Koike M, Nakanishi H, Saftig P, Ezaki J, Isahara K, Ohsawa Y, Schulz-Schaeffer W, Watanabe T, Waguri S, Kametaka S, Shibata M, Yamamoto K, Kominami E, Peters C, von Figura K and Uchiyama T . (2000). J. Neurosci., 20, 6898–6906.
Leist M and Jaattela M . (2001a). Nat. Rev. Mol. Cell Biol., 2, 589–598.
Leist M and Jaattela M . (2001b). Cell Death Differ., 8, 324–326.
Levicar N, Nutall RK and Lah TT . (2003). Acta Neurochir., 145, 825–838.
Li W, Yuan X, Nordgren G, Dalen H, Dubowchik GM, Firestone RA and Brunk UT . (2000). FEBS Lett., 470, 35–39.
Li W, Yuan XM, Ivanova S, Tracey KJ, Eaton JW and Brunk UT . (2003). Biochem. J., 371, 429–436.
Lieuallen K, Pennacchio LA, Park M, Myers RM and Lennon GG . (2001). Hum. Mol. Genet., 10, 1867–1871.
Liu N, Raja SM, Zazzeroni F, Metkar SS, Shah R, Zhang M, Wang Y, Bromme D, Russin WA, Lee JC, Peter ME, Froelich CJ, Franzoso G and Ashton-Rickardt PG . (2003). EMBO J., 22, 5313–5322.
Los M, Wesselborg S and Schulze-Osthoff K . (1999). Immunity, 10, 629–639.
Madge LA, Li JH, Choi J and Pober JS . (2003). J. Biol. Chem., 278, 21295–21306.
Manna SK, Zhang HJ, Yan T, Oberley LW and Aggarwal BB . (1998). J. Biol. Chem., 273, 13245–13254.
Matus A and Green GD . (1987). Biochemistry, 26, 8083–8086.
Meyer A, Fritz E, Fortelny R, Kofler K and Ludwig H . (1997). Cancer Invest., 15, 106–110.
Nakagawa T, Roth W, Wong P, Nelson A, Farr A, Deussing J, Villadangos JA, Ploegh HL, Peters C and Rudensky AY . (1998). Science, 280, 450–453.
Nakanishi H, Amano T, Sastradipura DF, Yoshimine Y, Tsukuba T, Tanabe K, Hirotsu I, Ohono T and Yamamoto K . (1997). J. Neurochem., 68, 739–749.
Nishimura Y, Sameni M and Sloane BF . (1998). Pathol. Oncol. Res., 4, 283–296.
Ollinger K . (2000). Arch. Biochem. Biophys., 373, 346–351.
Ono K, Kim SO and Han J . (2003). Mol. Cell. Biol., 23, 665–676.
Persson HL, Yu Z, Tirosh O, Eaton JW and Brunk UT . (2003). Free Radical Biol. Med., 34, 1295–1305.
Raha S and Robinson BH . (2000). Trends Biochem. Sci., 25, 502–508.
Reiners JJJ, Caruso JA, Mathieu P, Chelladurai B, Yin XM and Kessel D . (2002). Cell Death Differ., 9, 934–944.
Reinheckel T, Deussing J, Roth W and Peters C . (2001). Biol. Chem., 382, 735–741.
Rempel SA, Rosemblum ML, Mikkelsen T, Yan PS, Ellis KD, Golembieski WA, Sameni M, Rozhin J, Ziegler G and Sloane BF . (1994). Cancer Res., 54, 6027–6031.
Roberg K, Johansson U and Ollinger K . (1999). Free Radical Biol. Med., 27, 1228–1237.
Roberg K and Ollinger K . (1998). Am. J. Pathol., 152, 1151–1156.
Roberts LR, Adjei PN and Gores GJ . (1999). Cell Biochem. Biophys., 30, 71–88.
Roberts LR, Kurosawa H, Bronk SF, Fesmier PJ, Agellon LB, Leung WY, Mao F and Gores GJ . (1997). Gastroenterology, 113, 1714–1726.
Roth W, Deussing J, Botchkarev VA, Pauly-Evers M, Saftig P, Hafner A, Schmidt P, Schmahl W, Scherer J, Anton-Lamprecht I, von Figura K, Paus R and Peters C . (2000). FASEB J., 14, 2075–2086.
Saftig P, Hetman M, Schmahl W, Weber K, Heine L, Mossmann H, Koster A, Hess B, Evers M, von Figura K and Peters C . (1995). EMBO J., 15, 3599–3608.
Schotte P, Van Criekinge W, Van de Craen M, Van Loo G, Desmedt M, Grooten J, Cornelissen M, De Ridder L, Vandekerckhove J, Fiers W, Vandenabeele P and Beyaert R . (1998). Biochem. Biophys. Res. Commun., 251, 379–387.
Schutze S, Machliedt T, Adam D, Schwander R, Weigmann K, Kruse M-L, Heinrich M, Wickel M and Kronke M . (1999). J. Biol. Chem., 274, 10203–10212.
Shibata M, Kanamori S, Isahara K, Ohsawa Y, Konishi A, Kametaka S, Watanabe T, Ebisu S, Ishido K, Kominami E and Uchiyama Y . (1998). Biochem. Biophys. Res. Commun., 251, 199–203.
Shuja S and Murnane MJ . (1996). Int. J. Cancer, 66, 420–426.
Sinha AA, Wilson MJ, Gleason DF, Reddy PK, Sameni M and Sloane BF . (1995). Prostate, 26, 171–178.
Smolen JE, Stoehr SJ and Boxer LA . (1986). Biochem. Biophys. Acta., 886, 1–17.
Stoka VV, Turk B, Schendel SL, Kim TH, Cirman T, Snipas SJ, Ellerby LM, Bredesen D, Freeze H, Abrahamson M, Bromme D, Krajewski S, Reed JC, Yin XM, Turk V and Salvesen GS . (2001). J. Biol. Chem., 276, 3149–3157.
Sukoh N, Abe S, Ogura S, Isobe H, Takekawa H, Inoue K and Kawakami Y . (1994). Cancer, 74, 46–51.
Suzuki YJ, Forman HJ and Sevanian A . (1997). Free Radical Biol. Med., 22, 269–285.
Tobin DJ, Foitzik K, Reinheckel T, Mecklenburg L, Botchkarev VA, Peters C and Paus R . (2002). Am. J. Pathol., 160, 1807–1821.
Tsuchiya K, Kohda Y, Yoshida M, Zhao L, Ueno T, Yamashida J, Yoshioka T, Kominami E and Yamashima T . (1999). Exp. Neurol., 155, 187–194.
Turk B, Bieth JG, Bjork I, Dolenc I, Turk D, Cimerman N, Kos J, Colic A, Stoka V and Turk V . (1995). Biol. Chem. Hoppe. Seyler, 376, 225–230.
Turk B, Dolenc I, Turk V and Bieth JG . (1993). Biochemistry, 32, 375–380.
Turk B, Stoka V, Rozman-Pungercar J, Cirman T, Droga-Mazovec G, Oresic K and Turk V . (2002). Biol. Chem., 383, 1035–1044.
Turk B, Turk D and Turk V . (2000). Biochim. Biophys. Acta, 1477, 98–111.
Turk B, Turk V and Turk D . (1997). Biol. Chem., 378, 141–150.
Vancompernolle K, Van Herreweghe F, Pynaert G, Van de Craen M, De Vos K, Totty N, Sterling A, Fiers W, Vandenabeele P and Grooten J . (1998). FEBS Lett., 438, 150–158.
Walczak H, Miller RE, Ariail K, Gliniak B, Griffith TS, Kubin M, Chin W, Jones J, Woodward A, Le T, Smith C, Smolak P, Goodwin RG, Rauch CT, Schuh JCL and Lynch DH . (1999). Nat. Med., 5, 157–163.
Wang JH, Redmond HP, Watson RW and Bouchier-Hayes D . (1997). Am. J. Physiol., 272, C1543–C1551.
Watanabe M, Higashi T, Watanabe A, Osawa T, Sato Y, Kimura Y, Tominaga S, Hashimoto N, Yoshida Y, Morimoto S, Shito S, Hasinoto M, Kobayashi M, Tomoda J and Tsuji T . (1989). Biochem. Med. Metab. Biol., 42, 21–29.
Welss T, Sun J, Irving JA, Blum R, Smith AI, Whisstock JC, Pike RN, von Mikecz A, Ruzicka T, Bird PI and Abts HF . (2003). Biochemistry, 42, 7381–7389.
Werneburg NW, Guicciardi ME, Bronk SF and Gores GJ . (2002). Am. J. Physiol. Gastrointest Liver Physiol., 283, G947–G956.
Wu GS, Saftig P, Peters C and El-Deiry WS . (1998). Oncogene, 16, 2177–2183.
Xing R, Wu F and Mason RW . (1998). Cancer Res., 58, 904–909.
Yamashima T, Kohda Y, Tsuchiya K, Ueno T, Yamashita J, Yoshioka T and Kominami E . (1998). Eur. J. Neurosci., 10, 1723–1733.
Zdolsek J, Zhang H, Roberg K and Brunk UT . (1993). Free Radical Res. Commun., 18, 71–85.
Zhao M, Antunes F, Eaton JW and Brunk UT . (2003). Eur. J. Biochem., 270, 3778–3786.
Zhao M, Brunk UT and Eaton JW . (2001a). FEBS Lett., 509, 399–404.
Zhao M, Eaton JW and Brunk UT . (2000). FEBS Lett., 485, 104–108.
Zhao M, Eaton JW and Brunk UT . (2001b). FEBS Lett., 509, 405–412.
Acknowledgements
This work was supported by NIH Grant (DK63947 to G.J.G.), the Palumbo Foundation, and the Mayo Foundation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Guicciardi, M., Leist, M. & Gores, G. Lysosomes in cell death. Oncogene 23, 2881–2890 (2004). https://doi.org/10.1038/sj.onc.1207512
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1207512
Keywords
This article is cited by
-
The identification of a two-gene prognostic model based on cisplatin resistance-related ceRNA network in small cell lung cancer
BMC Medical Genomics (2023)
-
RIPK1 inhibition contributes to lysosomal membrane stabilization in ischemic astrocytes via a lysosomal Hsp70.1B-dependent mechanism
Acta Pharmacologica Sinica (2023)
-
Diversity and complexity of cell death: a historical review
Experimental & Molecular Medicine (2023)
-
The romantic history of signaling pathway discovery in cell death: an updated review
Molecular and Cellular Biochemistry (2023)
-
Synthesis, antitumor activities and functional mechanism of purine derivatives harboring phenyl moieties through three carbon bridges
Medicinal Chemistry Research (2023)