Abstract
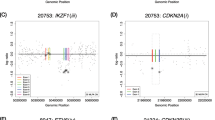
Chromosomal loss of heterozygosity (LOH) is a common mechanism for the inactivation of tumor suppressor genes in human epithelial cancers. Hybridization to single-nucleotide polymorphism (SNP) arrays is an efficient method to detect genome-wide cancer LOH. Here, we survey LOH patterns in a panel of 33 human lung cancer cell lines using SNP array hybridization containing 1500 SNPs. We compared the LOH patterns generated by SNP array hybridization to those previously obtained by 399 microsatellite markers and find a high degree of concordance between the two methods. A novel informatics platform, dChipSNP, was used to perform hierarchical tumor clustering based on genome-wide LOH patterns. We demonstrate that this method can separate non-small-cell and small-cell lung cancer samples based on their shared LOH. Furthermore, we analysed seven human lung cancer cell lines using a novel 10 000 SNP array and demonstrate that this is an efficient and reliable method of high-density allelotyping. Using this array, we identified small regions of LOH that were not detected by lower density SNP arrays or by standard microsatellite marker panels.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Alizadeh AA, Eisen MB, Davis RE, Ma C, Lossos IS, Rosenwald A, Boldrick JC, Sabet H, Tran T, Yu X, Powell JI, Yang L, Marti GE, Moore T, Hudson Jr J, Lu L, Lewis DB, Tibshirani R, Sherlock G, Chan WC, Greiner TC, Weisenburger DD, Armitage JO, Warnke R, Levy R, Wilson W, Grever MR, Byrd JC, Botstein D, Brown PO and Staudt LM . (2000). Nature, 403, 503–511.
Beer DG, Kardia SL, Huang CC, Giordano TJ, Levin AM, Misek DE, Lin L, Chen G, Gharib TG, Thomas DG, Lizyness ML, Kuick R, Hayasaka S, Taylor JM, Iannettoni MD, Orringer MB and Hanash S . (2002). Nat. Med., 8, 816–824.
Bhattacharjee A, Richards WG, Staunton J, Li C, Monti S, Vasa P, Ladd C, Beheshti J, Bueno R, Gillette M, Loda M, Weber G, Mark EJ, Lander ES, Wong W, Johnson BE, Golub TR, Sugarbaker DJ and Meyerson M . (2001). Proc. Natl. Acad. Sci. USA, 98, 13790–13795.
Dumur CI, Dechsukhum C, Ware JL, Cofield SS, Best AM, Wilkinson DS, Garrett CT and Ferreira-Gonzalez A . (2003). Genomics, 81, 260–269.
Eisen MB, Spellman PT, Brown PO and Botstein D . (1998). Proc. Natl. Acad. Sci. USA, 95, 14863–14868.
Fong KM, Zimmerman PV and Smith PJ . (1994). Genes Chromosomes Cancer, 10, 183–189.
Fong KM, Zimmerman PV and Smith PJ . (1995). Cancer Res., 55, 220–223.
Garber ME, Troyanskaya OG, Schluens K, Petersen S, Thaesler Z, Pacyna-Gengelbach M, van de Rijn M, Rosen GD, Perou CM, Whyte RI, Altman RB, Brown PO, Botstein D and Petersen I . (2001). Proc. Natl. Acad. Sci. USA, 98, 13784–13789.
Girard L, Zochbauer-Muller S, Virmani AK, Gazdar AF and Minna JD . (2000). Cancer Res., 60, 4894–4906.
Golub TR, Slonim DK, Tamayo P, Huard C, Gaasenbeek M, Mesirov JP, Coller H, Loh ML, Downing JR, Caligiuri MA, Bloomfield CD and Lander ES . (1999). Science, 286, 531–537.
Hoque MO, Lee CC, Cairns P, Schoenberg M and Sidransky D . (2003). Cancer Res., 63, 2216–2222.
Janne PA, Tanenbaum DM, Beheshti J, Mark EJ, Johnson BE and Meyerson ML . (2002). Proc. Am. Assoc. Can. Res., 43, 3147a.
Jemal A, Murray T, Samuels A, Ghafoor A, Ward E and Thun MJ . (2003). CA Cancer J. Clin., 53, 5–26.
Kennedy GC, Matsuzaki H, Dong S, Liu WM, Huang J, Liu G, Su X, Cao M, Chen W, Zhang J, Liu W, Yang G, Di X, Ryder T, He Z, Surti U, Phillips MS, Boyce-Jacino MT, Fodor SP and Jones KW . (2003). Nat. Biotechnol., 21, 1233–1237.
Li C and Wong WH . (2001). Proc. Natl. Acad. Sci. USA, 98, 31–36.
Lieberfarb ME, Lin M, Lechpammer M, Li C, Tanenbaum DM, Febbo PG, Wright RL, Shim J, Kantoff PW, Loda M, Meyerson M and Sellers WR . (2003). Cancer Res., 63, 4781–4785.
Lin M, Wei L-J, Sellers WR, Lieberfarb M, Wong WH and Li C . (2003). Bioinformatics, (in press).
Lindblad-Toh K, Tanenbaum DM, Daly MJ, Winchester E, Lui WO, Villapakkam A, Stanton SE, Larsson C, Hudson TJ, Johnson BE, Lander ES and Meyerson M . (2000). Nat. Biotechnol., 18, 1001–1005.
Mei R, Galipeau PC, Prass C, Berno A, Ghandour G, Patil N, Wolff RK, Chee MS, Reid BJ and Lockhart DJ . (2000). Genome Res., 10, 1126–1137.
Mitsudomi T, Oyama T, Nishida K, Ogami A, Osaki T, Sugio K, Yasumoto K, Sugimachi K and Gazdar AF . (1996). Clin. Cancer Res., 2, 1185–1189.
Osada H and Takahashi T . (2002). Oncogene, 21, 7421–7434.
Primdahl H, Wikman FP, von der Maase H, Zhou XG, Wolf H and Orntoft TF . (2002). J. Natl. Cancer Inst., 94, 216–223.
Reich DE, Gabriel SB and Altshuler D . (2003). Nat. Genet., 33, 457–458.
Sachidanandam R, Weissman D, Schmidt SC, Kakol JM, Stein LD, Marth G, Sherry S, Mullikin JC, Mortimore BJ, Willey DL, Hunt SE, Cole CG, Coggill PC, Rice CM, Ning Z, Rogers J, Bentley DR, Kwok PY, Mardis ER, Yeh RT, Schultz B, Cook L, Davenport R, Dante M, Fulton L, Hillier L, Waterston RH, McPherson JD, Gilman B, Schaffner S, Van Etten WJ, Reich D, Higgins J, Daly MJ, Blumenstiel B, Baldwin J, Stange-Thomann N, Zody MC, Linton L, Lander ES and Altshuler D . (2001). Nature, 409, 928–933.
Sekido Y, Fong KM and Minna JD . (2003). Annu. Rev. Med., 54, 73–87.
Stanton SE, Shin SW, Johnson BE and Meyerson M . (2000). Genes Chromosomes Cancer, 27, 323–331.
Tseng JE, Kemp BL, Khuri FR, Kurie JM, Lee JS, Zhou X, Liu D, Hong WK and Mao L . (1999). Cancer Res., 59, 4798–4803.
Virmani AK, Fong KM, Kodagoda D, McIntire D, Hung J, Tonk V, Minna JD and Gazdar AF . (1998). Genes Chromosomes Cancer, 21, 308–319.
Wang ZC, Lin M, Wei L-J, Li C, Miron A, Lodeiro G, Harris L, Ramaswamy S, Tanenbaum DM, Meyerson M, Iglehart JD and Richardson A . (2003). Cancer Res., (in press).
Zabarovsky ER, Lerman MI and Minna JD . (2002). Oncogene, 21, 6915–6935.
Acknowledgements
This work is supported by grants from the National Cancer Institute R01CA92824 (DC), the National Institutes of Health 2P30 CA06516-39 (CL) and 1K12CA87723-01 (PAJ), the American Cancer Society RSG-03-240-01-MGO (MM), the Pew Scholars in the Biomedical Sciences (MM), the Flight Attendant Medical Research Institute (MM), the Dunkin Donuts Rising Star Award (PAJ), The Arthur and Rochelle Belfer Foundation (CL and MM), Lung Cancer SPORE P50CA70907 (JM) and The Longenbaugh Foundation (JM).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on Oncogene website (http://www.nature.com/onc) and also at research.dfci.harvard.edu/meyersonlab/snp/janne-oncogene2004
Supplementary information
Rights and permissions
About this article
Cite this article
Jänne, P., Li, C., Zhao, X. et al. High-resolution single-nucleotide polymorphism array and clustering analysis of loss of heterozygosity in human lung cancer cell lines. Oncogene 23, 2716–2726 (2004). https://doi.org/10.1038/sj.onc.1207329
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1207329
Keywords
This article is cited by
-
Genome wide SNP comparative analysis between EGFR and KRAS mutated NSCLC and characterization of two models of oncogenic cooperation in non-small cell lung carcinoma
BMC Medical Genomics (2008)
-
Virtual karyotyping with SNP microarrays reduces uncertainty in the diagnosis of renal epithelial tumors
Diagnostic Pathology (2008)
-
SNPExpress: integrated visualization of genome-wide genotypes, copy numbers and gene expression levels
BMC Genomics (2008)
-
Whole genome SNP arrays as a potential diagnostic tool for the detection of characteristic chromosomal aberrations in renal epithelial tumors
Modern Pathology (2008)
-
Genome-wide DNA copy number analysis in pancreatic cancer using high-density single nucleotide polymorphism arrays
Oncogene (2008)