Abstract
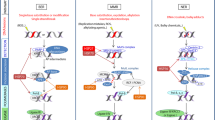
Heat-shock proteins are produced in response to different types of stress conditions making cells resistant to stress-induced cell damage. Under normal conditions, heat-shock proteins play numerous roles in cell function, including modulating protein activity by changing protein conformation, promoting multiprotein complex assembly/disassembly, regulating protein degradation within the proteasome pathway, facilitating protein translocation across organellar membranes, and ensuring proper folding of nascent polypeptide chains during protein translation. When cells are stressed, a common response is to undergo cell death by one of two pathways, either ‘necrosis’ or ‘apoptosis’. Recently, both routes to cell death have been revealed to share similar mechanisms, with heat-shock proteins and their cofactors responsible for inhibiting both apoptotic and necrotic pathways. We therefore briefly summarize recent reports showing molecular evidence of cell death regulation by heat-shock proteins and their cochaperones.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Antoku K, Maser RS, Scully Jr WJ, Delach SM and Johnson DE . (2001). Biochem. Biophys. Res. Commun., 286, 1003–1010.
Auluck PK, Chan HY, Trojanowski JQ, Lee VM and Bonini NM . (2002). Science, 295, 858–865.
Ballinger CA, Connell P, Wu Y, Hu Z, Thompson LJ, Yin LY and Patterson C . (1999). Mol. Cell. Biol., 19, 4535–4545.
Basso AD, Solit DB, Chiosis G, Giri B, Tsichlis P and Rosen N . (2002). J. Biol. Chem., 277, 39858–39866.
Beere HM, Wolf BB, Cain K, Mosser DD, Mahboubi A, Kuwana T, Tailor P, Morimoto RI, Cohen GM and Green DR . (2000). Nat. Cell. Biol., 2, 469–475.
Benn SC, Perrelet D, Kato AC, Scholz J, Decosterd I, Mannion RJ, Bakowska JC and Woolf CJ . (2002). Neuron, 36, 45–56.
Biggs III WH, Meisenhelder J, Hunter T, Cavenee WK and Arden KC . (1999). Proc. Natl. Acad. Sci. USA, 96, 7421–7426.
Bimston D, Song J, Winchester D, Takayama S, Reed JC and Morimoto RI . (1998). EMBO J., 17, 6871–6878.
Bodine SC, Latres E, Baumhueter S, Lai VK, Nunez L, Clarke BA, Poueymirou WT, Panaro FJ, Na E, Dharmarajan K, Pan ZQ, Valenzuela DM, DeChiara TM, Stitt TN, Yancopoulos GD and Glass DJ . (2001). Science, 294, 1704–1708.
Bonini NM . (2002). Proc. Natl. Acad. Sci. USA, 99, 16407–16411.
Bruey JM, Ducasse C, Bonniaud P, Ravagnan L, Susin SA, Diaz-Latoud C, Gurbuxani S, Arrigo AP, Kroemer G, Solary E and Garrido C . (2000a). Nat. Cell. Biol., 2, 645–652.
Bruey JM, Paul C, Fromentin A, Hilpert S, Arrigo AP, Solary E and Garrido C . (2000b). Oncogene, 19, 4855–4863.
Cardone MH, Roy N, Stennicke HR, Salvesen GS, Franke TF, Stanbridge E, Frisch S and Reed JC . (1998). Science, 282, 1318–1321.
Charette SJ, Lavoie JN, Lambert H and Landry J . (2000). Mol. Cell. Biol., 20, 7602–7612.
Chen G, Cao P and Goeddel DV . (2002). Mol. Cell., 9, 401–410.
Chen F, Chang R, Trivedi M, Capetanaki Y and Cryns VL . (2003). J. Biol. Chem., 278, 6848–6853.
Chittenden T . (2002). Cancer Cell., 2, 165–166.
Clevenger CV, Thickman K, Ngo W, Chang WP, Takayama S and Reed JC . (1997). Mol. Endocrinol., 11, 608–618.
Danen-van Oorschot AAAM, den Hollander A, Takayama S, Reed JC, van der Eb AJ and Noteborn MHM . (1997). Apoptosis, 2, 395–402.
Datta SR, Brunet A and Greenberg ME . (1999). Genes Dev., 13, 2905–2927.
Datta SR, Dudek H, Tao X, Masters S, Fu H, Gotoh Y and Greenberg ME . (1997). Cell, 91, 231–241.
Demonacos C, Krstic-Demonacos M and La Thangue NB . (2001). Mol. Cell., 8, 71–84.
Dix DJ, Allen JW, Collins BW, Mori C, Nakamura N, Poorman-Allen P, Goulding EH and Eddy EM . (1996). Proc. Natl. Acad. Sci. USA, 93, 33264–33268.
Doong H, Price J, Kim YS, Gasbarre C, Probst J, Liotta LA, Blanchette J, Rizzo K and Kohn E . (2000). Oncogene, 19, 4385–4395.
Eversole-Cire P, Concepcion F.A, Simon M.I, Takayama S, Reed JC and Chen J . (2000). Investigative Ophthalmology and Visual Science, 41, 1953–1961.
Gabai VL, Yaglom JA, Volloch V, Meriin AB, Force T, Koutroumanis M, Massie B, Mosser DD and Sherman MY . (2000). Mol. Cell. Biol., 20, 6826–6836.
Gao T and Newton AC . (2002). J. Biol. Chem., 277, 31585–31592.
Garrido C, Bruey JM, Fromentin A, Hammann A, Arrigo AP and Solary E . (1999). FASEB J., 13, 2061–2070.
Gjoerup O, Chao H, DeCaprio JA and Roberts TM . (2000). J. Virol., 74, 864–874.
Gomes MD, Lecker SH, Jagoe RT, Navon A and Goldberg AL . (2001). Proc. Natl. Acad. Sci. USA, 98, 14440–14445.
Gross A, Yin XM, Wang K, Wei MC, Jockel J, Milliman C, Erdjument-Bromage H, Tempst P and Korsmeyer SJ . (1999). J. Biol. Chem., 274, 1156–1163.
Gupta S and Knowlton AA . (2002). Circulation, 106, 2727–2733.
Imai Y, Soda M, Hatakeyama S, Akagi T, Hashikawa T, Nakayama KI and Takahashi R . (2002). Mol. Cell., 10, 55–67.
Imai Y, Soda M, Inoue H, Hattori N, Mizuno Y and Takahashi R . (2001). Cell, 105, 891–902.
Iwaki K, Chi SH, Dillmann WH and Mestril R . (1993). Circulation, 87, 2023–2032.
Jaattela M, Wissing D, Kokholm K, Kallunki T and Egeblad M . (1998). EMBO J., 17, 6124–6134.
Jiang J, Ballinger CA, Wu Y, Dai Q, Cyr DM, Hohfeld J and Patterson C . (2001). J. Biol. Chem., 276, 42938–42944.
Jiang Y, Woronicz JD, Liu W and Goeddel DV . (1999). Science, 283, 543–546.
Kamradt MC, Chen F, Sam S and Cryns VL . (2002). J. Biol. Chem., 277, 38731–38876.
Kato K, Ito H, Kamei K, Iwamoto I and Inaguma Y . (2002). FASEB J., 16, 1432–1434.
Kawagoe J, Abe K, Aoki M and Kogure K . (1993). Brain Res., 621, 121–125.
Kazemi-Esfarjani P and Benzer S . (2000). Science, 287, 1837–1840.
Kelley WL . (1998). Trends Biochem. Sci., 23, 222–227.
Kermer P, Digicaylioglu MH, Kaul M, Zapata JM, Krajewska M, Stenner-Liewen F, Takayama S, Krajewski S, Lipton S.A and Reed JC . (2003). Brain Pathology, in press.
Kim S, Nollen EA, Kitagawa K, Bindokas VP and Morimoto RI . (2002). Nat. Cell. Biol., 4, 826–831.
Konishi H, Matsuzaki H, Tanaka M, Takemura Y, Kuroda S, Ono Y and Kikkawa U . (1997). FEBS Lett., 410, 493–498.
Kurisu J, Honma A, Miyajima H, Kondo S, Okumura M and Imaizumi K . (2003). Genes Cells, 8, 189–202.
Lambert H, Charette SJ, Bernier AF, Guimond A and Landry J . (1999). J. Biol. Chem., 274, 9378–9385.
Lee MY, Kim SY, Shin SL, Choi YS, Lee JH and Tsujimoto Y . (2002). Exp. Neurol., 175, 338–346.
Lee JH, Takahashi T, Yasuhara N, Inazawa J, Kamada S and Tsujimoto Y . (1999). Oncogene, 18, 6183–6190.
Lewis J, Devin A, Miller A, Lin Y, Rodriguez Y, Neckers L and Liu ZG . (2000). J. Biol. Chem., 275, 10519–10526.
Li H, Zhu H, Xu CJ and Yuan J . (1998). Cell, 94, 491–501.
Liao Q, Ozawa F, Friess H, Zimmermann A, Takayama S, Reed JC, Kleeff J and Buchler MW . (2001). FEBS Lett., 503, 151–157.
Luo X, Budihardjo I, Zou H, Slaughter C and Wang X . (1998). Cell, 94, 481–490.
Mehlen P, Kretz-Remy C, Preville X and Arrigo AP . (1996a). EMBO J., 15, 2695–2706.
Mehlen P, Schulze-Osthoff K and Arrigo AP . (1996b). J. Biol. Chem., 271, 16510–16514.
Meriin AB, Yaglom JA, Gabai VL, Zon L, Ganiatsas S, Mosser DD and Sherman MY . (1999). Mol. Cell. Biol., 19, 2547–2555.
Miki K and Eddy EM . (2002). Mol. Cell. Biol., 22, 2536–2543.
Mosser DD, Caron AW, Bourget L, Meriin AB, Sherman MY, Morimoto RI and Massie B . (2000). Mol. Cell. Biol., 20, 7146–7159.
Okubo S, Wildner O, Shah MR, Chelliah JC, Hess ML and Kukreja RC . (2001). Circulation, 103, 877–881.
Oliner JD, Pietenpol JA, Thiagalingam S, Gyuris J, Kinzler KW and Vogelstein B . (1993). Nature, 362, 857–860.
Ozes ON, Mayo LD, Gustin JA, Pfeffer SR, Pfeffer LM and Donner DB . (1999). Nature, 401, 82–85.
Pagliuca MG, Lerose R, Cigliano S and Leone A . (2003). FEBS Lett., 541, 11–15.
Pang Q, Christianson TA, Keeble W, Koretsky T and Bagby GC . (2002). J. Biol. Chem., 277, 49638–49643.
Pang Q, Keeble W, Christianson TA, Faulkner GR and Bagby GC . (2001). EMBO J., 20, 4478–4489.
Pfanner N . (1999). Curr. Biol., 9, R720–R724.
Radford NB, Fina M, Benjamin IJ, Moreadith RW, Graves KH, Zhao P, Gavva S, Wiethoff A, Sherry AD, Malloy CR and Williams RS . (1996). Proc. Natl. Acad. Sci. USA, 93, 2339–2342.
Ravagnan L, Gurbuxani S, Susin SA, Maisse C, Daugas E, Zamzami N, Mak T, Jaattela M, Penninger JM, Garrido C and Kroemer G . (2001). Nat. Cell. Biol., 3, 839–843.
Rogalla T, Ehrnsperger M, Preville X, Kotlyarov A, Lutsch G, Ducasse C, Paul C, Wieske M, Arrigo AP, Buchner J and Gaestel M . (1999). J. Biol. Chem., 274, 18947–18956.
Romano M, Festa M, Pagliuca G, Lerose R, Bisogni R, Chiurazzi F, Storti G, Volpe S, Venuta S, Turco M and Leone S . (2003). Cell Death Differ., 10, 383–385.
Sakahira H and Nagata S . (2002). J. Biol. Chem., 277, 3364–3370.
Sakamoto H, Mashima T, Yamamoto K and Tsuruo T . (2002). J. Biol. Chem., 277, 45770–45775.
Saleh A, Srinivasula SM, Balkir L, Robbins PD and Alnemri ES . (2000). Nat. Cell. Biol., 2, 476–483.
Samali A, Cai J, Zhivotovsky B, Jones DP and Orrenius S . (1999). EMBO J., 18, 2040–2048.
Sato S, Fujita N and Tsuruo T . (2000). Proc. Natl. Acad. Sci. USA, 97, 10832–10837.
Suzuki K, Sawa Y, Kaneda Y, Ichikawa H, Shirakura R and Matsuda H . (1997). J. Clin. Invest., 99, 1645–1650.
Syken J, De-Medina T and Munger K . (1999). Proc. Natl. Acad. Sci. USA, 96, 8499–8504.
Takayama S, Bimston DN, Matsuzawa S, Freeman BC, Aime-Sempe C, Xie Z, Morimoto RI and Reed JC . (1997). EMBO J., 16, 4887–4896.
Takayama S, Sato T, Krajewski S, Kochel K, Irie S, Millan JA and Reed JC . (1995). Cell, 80, 279–284.
Takayama S, Xie Z and Reed J . (1999). J. Biol. Chem., 274, 781–786.
Terada S, Fukuoka K, Fujita T, Komatsu T, Takayama S, Reed JC and Suzuki E . (1997). Cytotechnology, 25, 17–23.
Thress K, Song J, Morimoto RI and Kornbluth S . (2001). EMBO J., 20, 1033–1041.
Van Molle W, Wielockx B, Mahieu T, Takada M, Taniguchi T, Sekikawa K and Libert C . (2002). Immunity, 16, 685–695.
Vanden Berghe T, Kalai M, Van Loo G, Declercq W and Vandenabeele P . (2003). J. Biol. Chem., 278, 5622–5629.
Vanhaesebroeck B and Alessi DR . (2000). Biochem. J., 346, 561–576.
Varfolomeev EE, Schuchmann M, Luria V, Chiannilkulchai N, Beckmann JS, Mett IL, Rebrikov D, Brodianski VM, Kemper OC, Kollet O, Lapidot T, Soffer D, Sobe T, Avraham KB, Goncharov T, Holtmann H, Lonai P and Wallach D . (1998). Immunity, 9, 267–276.
Wang X, Osinska H, Klevitsky R, Gerdes AM, Nieman M, Lorenz J, Hewett T and Robbins J . (2001). Circ. Res., 89, 84–91.
Wang HG, Takayama S, Rapp UR and Reed JC . (1996). Proc. Natl. Acad. Sci. USA, 93, 7063–7068.
Warrick JM, Chan HY, Gray-Board GL, Chai Y, Paulson HL and Bonini NM . (1999). Nat. Genet., 23, 425–428.
Xanthoudakis S, Roy S, Rasper D, Hennessey T, Aubin Y, Cassady R, Tawa P, Ruel R, Rosen A and Nicholson DW . (1999). EMBO J., 18, 2049–2056.
Xiang SL, Kumano T, Iwasaki SI, Sun X, Yoshioka K and Yamamoto KC . (2001). Biochem. Biophys. Res. Commun., 287, 932–940.
Yeh WC, Itie A, Elia AJ, Ng M, Shu HB, Wakeham A, Mirtsos C, Suzuki N, Bonnard M, Goeddel DV and Mak TW . (2000). Immunity, 12, 633–642.
Zhang X, Beuron F and Freemont PS . (2002). Curr. Opin. Struct. Biol., 12, 231–238.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Takayama, S., Reed, J. & Homma, S. Heat-shock proteins as regulators of apoptosis. Oncogene 22, 9041–9047 (2003). https://doi.org/10.1038/sj.onc.1207114
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1207114
Keywords
This article is cited by
-
Cytotoxicity of bismuth(III) dithiocarbamate derivatives by promoting a mitochondrial-dependent apoptotic pathway and suppressing MCF-7 breast adenocarcinoma cell invasion
JBIC Journal of Biological Inorganic Chemistry (2024)
-
Cellular protection from H2O2 toxicity by Fv-Hsp70: protection via catalase and gamma-glutamyl-cysteine synthase
Cell Stress and Chaperones (2023)
-
In vitro effects of 5-Hydroxy-L-tryptophan supplementation on primary bovine mammary epithelial cell gene expression under thermoneutral or heat shock conditions
Scientific Reports (2022)
-
Effect of silver nanoparticles on gene transcription of land snail Helix aspersa
Scientific Reports (2022)
-
Carry-over effects of dry period heat stress on the mammary gland proteome and phosphoproteome in the subsequent lactation of dairy cows
Scientific Reports (2022)