Abstract
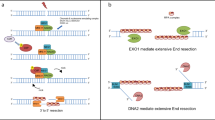
DNA nucleases catalyze the cleavage of phosphodiester bonds. These enzymes play crucial roles in various DNA repair processes, which involve DNA replication, base excision repair, nucleotide excision repair, mismatch repair, and double strand break repair. In recent years, new nucleases involved in various DNA repair processes have been reported, including the Mus81 : Mms4 (Eme1) complex, which functions during the meiotic phase and the Artemis : DNA-PK complex, which processes a V(D)J recombination intermediate. Defects of these nucleases cause genetic instability or severe immunodeficiency. Thus, structural biology on various nuclease actions is essential for the elucidation of the molecular mechanism of complex DNA repair machinery. Three-dimensional structural information of nucleases is also rapidly accumulating, thus providing important insights into the molecular architectures, as well as the DNA recognition and cleavage mechanisms. This review focuses on the three-dimensional structure-function relationships of nucleases crucial for DNA repair processes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Aravind L, Koonin EV . 1998a Trends Biochem. Sci. 23: 17–19
Aravind L, Koonin EV . 1998b Nucleic Acids Res. 26: 3746–3752
Aravind L, Walker DR, Koonin EV . 1999 Nucleic Acids Res. 27: 1223–1242
Beernink PT, Segelke BW, Hadi MZ, Erzberger JP, Wilson III DM, Rupp B . 2001 J. Mol. Biol. 307: 1023–1034
Beese LS, Steitz TA . 1991 EMBO J. 10: 25–33
Blunt T, Finnie NJ, Taccioli GE, Smith GC, Demengeot J, Gottlieb TM, Mizuta R, Varghese AJ, Alt FW, Jeggo PA . 1995 Cell, 80: 813–823
Boddy MN, Gaillard PH, McDonald WH, Shanahan P, Yates III JR, Russell P . 2001 Cell 107: 537–548
Boddy MN, Lopez-Girona A, Shanahan P, Interthal H, Heyer WD, Russell P . 2000 Mol. Cell. Biol. 20: 8758–8766
D'Amours D, Jackson SP . 2002 Nat. Rev. Mol. Cell. Biol. 3: 317–327
de Boer J, Hoeijmakers JH . 2000 Carcinogenesis 21: 453–460
Farber GK, Petsko GA . 1990 Trends Biochem. Sci. 15: 228–234
Fijalkowska IJ, Schaaper RM . 1996 Proc. Natl. Acad. Sci. USA 93: 2856–2861
Galburt EA, Chevalier B, Tang W, Jurica MS, Flick KE, Monnat Jr RJ, Stoddard BL . 1999 Nat. Struct. Biol. 6: 1096–1099
Goedken ER, Marqusee S . 2001 J. Biol. Chem. 276: 7266–7271
Goldsby RE, Lawrence NA, Hays LE, Olmsted EA, Chen X, Singh M, Preston BD . 2001 Nat. Med. 7: 638–639
Gorman MA, Morera S, Rothwell DG, de La Fortelle E, Mol CD, Tainer JA, Hickson ID, Freemont PS . 1997 EMBO J. 16: 6548–6558
Griffith JP, Kim JL, Kim EE, Sintchak MD, Thomson JA, Fitzgibbon MJ, Fleming MA, Caron PR, Hsiao K, Navia MA . 1995 Cell 82: 507–522
Hadden JM, Declais AC, Phillips SE, Lilley DM . 2002 EMBO J. 21: 3505–3515
Hamdan S, Carr PD, Brown SE, Ollis DL, Dixon NE . 2002 Structure (Camb) 10: 535–546
Hopfner KP, Karcher A, Craig L, Woo TT, Carney JP, Tainer JA . 2001 Cell 105: 473–485
Hosfield DJ, Mol CD, Shen B, Tainer JA . 1998 Cell 95: 135–146
Hosfield DJ, Guan Y, Haas BJ, Cunningham RP, Tainer JA . 1999 Cell 98: 397–408
Hwang KY, Baek K, Kim HY, Cho Y . 1998 Nat. Struct. Biol. 5: 707–713
Interthal H, Heyer WD . 2000 Mol. Gen. Genet. 263: 812–827
Jencks WP . 1969 Catalysis in Chemistry and Enzymology New York: McGraw Hill pp 111–115
Kaliraman V, Mullen JR, Fricke WM, Bastin-Shanower SA, Brill SJ . 2001 Genes Dev. 15: 2730–2740
Katayanagi K, Okumura M, Morikawa K . 1993 Proteins 17: 337–346
Kirchgessner CU, Patil CK, Evans JW, Cuomo CA, Fried LM, Carter T, Oettinger MA, Brown JM . 1995 Science 267: 1178–1183
Lamers MH, Perrakis A, Enzlin JH, Winterwerp HH, de Wind N, Sixma TK . 2000 Nature 407: 711–717
Levin JD, Shapiro R, Demple B . 1991 J. Biol. Chem. 266: 22893–22898
Lieber MR . 1997 Bioessays 19: 233–240
Ma Y, Pannicke U, Schwarz K, Lieber MR . 2002 Cell 108: 781–794
Modrich P, Lahue R . 1996 Annu. Rev. Biochem. 65: 101–133
Mol CD, Hosfield DJ, Tainer JA . 2000a Mutat. Res. 460: 211–229
Mol CD, Izumi T, Mitra S, Tainer JA . 2000b Nature 403: 451–456
Mol CD, Kuo CF, Thayer MM, Cunningham RP, Tainer JA . 1995 Nature 374: 381–386
Morrison A, Johnson AL, Johnston LH, Sugino A . 1993 EMBO J. 12: 1467–1473
Moshous D, Callebaut I, de Chasseval R, Corneo B, Cavazzana-Calvo M, Le Deist F, Tezcan I, Sanal O, Bertrand Y, Philippe N, Fischer A, de Villartay JP . 2001 Cell 105: 177–186
Mullen JR, Kaliraman V, Ibrahim SS, Brill SJ . 2001 Genetics 157: 103–118
Murzin AG, Brenner SE, Hubbard T, Chothia C . 1995 J. Mol. Biol. 247: 536–540
Obmolova G, Ban C, Hsieh P, Yang W . 2000 Nature 407: 703–710
Parikh SS, Putnam CD, Tainer JA . 2000 Mutat. Res. 460: 183–199
Petit C, Sancar A . 1999 Biochimie 81: 15–25
Pingoud A, Jeltsch A . 2001 Nucleic Acids Res. 29: 3705–3727
Prakash S, Prakash L . 2000 Mutat. Res. 451: 13–24
Raaijmakers H, Vix O, Toro I, Golz S, Kemper B, Suck D . 1999 EMBO J. 18: 1447–1458
Ramotar D, Popoff SC, Gralla EB, Demple B . 1991 Mol. Cell. Biol. 11: 4537–4544
Roberts RJ, Cheng X . 1998 Annu. Rev. Biochem. 67: 181–198
Shamoo Y, Steitz TA . 1999 Cell 99: 155–166
Sharples GJ . 2001 Mol. Microbiol. 39: 823–834
Shevelev IV, Hubscher U . 2002 Nat. Rev. Mol. Cell. Biol. 3: 364–376
Stewart GS, Maser RS, Stankovic T, Bressan DA, Kaplan MI, Jaspers NG, Raams A, Byrd PJ, Petrini JH, Taylor AM . 1999 Cell 99: 577–587
Suck D, Lahm A, Oefner C . 1988 Nature 332: 464–468
Tsutakawa SE, Jingami H, Morikawa K . 1999a Cell 99: 615–623
Tsutakawa SE, Muto T, Kawate T, Jingami H, Kunishima N, Ariyoshi M, Kohda D, Nakagawa M, Morikawa K . 1999b Mol. Cell 3: 621–628
Tsutakawa SE, Morikawa K . 2001 Nucleic Acids Res. 19: 3775–3783
Vassylyev D, Morikawa K . 1997 Curr. Opin. Struct. Biol. 7: 103–109
Wilson III DM, Thompson LH . 1997 Proc. Natl. Acad. Sci. USA 94: 12754–12757
Xanthoudakis S, Miao G, Wang F, Pan YC, Curran T . 1992 EMBO J. 11: 3323–3335
Yamagata A, Kakuta Y, Masui R, Fukuyama K . 2002 Proc. Natl. Acad. Sci. USA 99: 5908–5912
Yang W . 2000 Mutat. Res. 460: 245–256
Yoshikawa M, Iwasaki H, Shinagawa H . 2001 J. Biol. Chem. 276: 10432–10436
Acknowledgements
We regret that the limit of space may have not allowed us to site all works in the field. We thank Kayoko Komori for critical reading of the manuscript and helpful comments. T Nishino is a research fellow of the Japan society for the promotion of sciences. This research was partly supported by NEDO (New Energy and Industrial Technology Development Organization).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Nishino, T., Morikawa, K. Structure and function of nucleases in DNA repair: shape, grip and blade of the DNA scissors. Oncogene 21, 9022–9032 (2002). https://doi.org/10.1038/sj.onc.1206135
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1206135
Keywords
This article is cited by
-
LAP2α preserves genome integrity through assisting RPA deposition on damaged chromatin
Genome Biology (2022)
-
Identification and characterization of DNA endonucleases in Plasmodium falciparum 3D7 clone
Malaria Journal (2018)
-
The Sulfolobus solfataricus RecQ-like DNA helicase Hel112 inhibits the NurA/HerA complex exonuclease activity
Extremophiles (2018)
-
ExoMeg1: a new exonuclease from metagenomic library
Scientific Reports (2016)
-
Phosphodiester hydrolysis computed for cluster models of enzymatic active sites
Theoretical Chemistry Accounts (2016)