Abstract
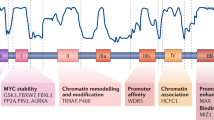
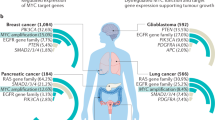
c-MYC is the prototype for oncogene activation by chromosomal translocation. In contrast to the tightly regulated expression of c-myc in normal cells, c-myc is frequently deregulated in human cancers. Herein, aspects of c-myc gene activation and the function of the c-Myc protein are reviewed. The c-myc gene produces an oncogenic transcription factor that affects diverse cellular processes involved in cell growth, cell proliferation, apoptosis and cellular metabolism. Complete removal of c-myc results in slowed cell growth and proliferation, suggesting that while c-myc is not required for cell proliferation, it acts as an integrator and accelerator of cellular metabolism and proliferation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Adams JM, Harris AW, Pinkert CA, Corcoran LM, Alexander WS, Cory S, Palmiter RD, Brinster RL . 1985 Nature 318: 533–538
Alevizopoulos K, Vlach J, Hennecke S, Amati B . 1997 EMBO J. 16: 5322–5333
Amati B, Alevizopoulos K, Vlach J . 1998 Front Biosci. 3: D250–D268
Amundson SA, Zhan Q, Penn LZ, Fornace Jr AJ . 1998 Oncogene 17: 2149–2154
Arcinas M, Boxer LM . 1994 Oncogene 9: 2699–2706
ar-Rushdi A, Nishikura K, Erikson J, Watt R, Rovera G, Croce C . 1983 Science 222: 390–393
Askew DS, Ashmun RA, Simmons BC, Cleveland JL . 1991 Oncogene 6: 1915–1922
Ayer DE, Kretzner L, Eisenman RN . 1993 Cell 72: 211–222
Ayer DE, Laherty CD, Lawrence QA, Armstrong AP, Eisenman RN . 1996 Mol. Cell. Biol. 16: 5772–5781
Bahram F, von der Lehr N, Cetinkaya C, Larsson LG . 2000 Blood 95: 2104–2110
Barr LF, Campbell SE, Bochner BS, Dang CV . 1998 Cancer Res. 58: 5537–5545
Barr LF, Campbell SE, Diette GB, Gabrielson EW, Kim S, Shim H, Dang CV . 2000 Cancer Res. 60: 143–149
Bernard O, Cory S, Gerondakis S, Webb E, Adams JM . 1983 EMBO J. 2: 2375–2383
Beier R, Burgin A, Kiermaier A, Fero M, Karsunky H, Saffrich R, Moroy T, Ansorge W, Roberts J, Eilers M . 2000 EMBO J. 19: 5813–5823
Beijersbergen RL, Hijmans EM, Zhu L, Bernards R . 1994 EMBO J. 13: 4080–4086
Bello-Fernandez C, Packham G, Cleveland JL . 1993 Proc. Natl. Acad. Sci. USA 90: 7804–7808
Bemark M, Neuberger MS . 2000 Oncogene 19: 3404–3410
Bentley DL, Groudine M . 1986 Nature 321: 702–706
Berridge MV, Tan AS, McCoy KD, Kansara M, Rudert F . 1996 J. Immunol. 156: 4092–4099
Bhatia K, Huppi K, Spangler G, Siwarski D, Iyer R, Magrath I . 1993 Nat. Genet. 5: 56–61
Blackwood EM, Eisenman RN . 1991 Science 251: 1211–1217
Bornkamm GW, Polack A, Eick D . 1988 c-myc Deregulation by Chromosomal Translocation in Burkitt's Lymphoma New York: M. Dekker, Inc
Bouchard C, Thieke K, Maier A, Saffrich R, Hanley-Hyde J, Ansorge W, Reed S, Sicinski P, Bartek J, Eilers M . 1999 EMBO J. 18: 5321–5333
Brakebusch C, Wennerberg K, Krell HW, Weidle UH, Sallmyr A, Johansson S, Fassler R . 1999 Oncogene 18: 3852–3861
Bradley JF, Rothberg PG, Ladanyi M, Chaganti RSK . 1993 Genes Chrom. Cancer 7: 128–130
Butzler C, Zou X, Popov AV, Bruggemann M . 1997 Oncogene 14: 1383–1388
Cesarman E, Dalla-Favera R, Bentley D, Groudine M . 1987 Science 238: 1272–1275
Chang DW, Claassen GF, Hann SR, Cole MD . 2000 Mol. Cell. Biol. 20: 4309–4319
Chou TY, Hart GW, Dang CV . 1995 J. Biol. Chem. 270: 18961–18965
Claassen GF, Hann SR . 2000 Proc. Natl. Acad. Sci. USA 97: 9498–9503
Cockerill PN, Yuen M-H, Garrard WT . 1987 J. Biol. Chem. 262: 5394–5397
Cole MD, McMahon SB . 1999 Oncogene 18: 2916–2924
Coller HA, Grandori C, Tamayo P, Colbert T, Lander ES, Eisenman RN, Golub TR . 2000 Proc. Natl. Acad. Sci. USA 97: 3260–3265
Conzen SD, Gottlob K, Kandel ES, Khanduri P, Wagner AJ, O'Leary M, Hay N . 2000 Mol. Cell. Biol. 20: 6008–6018
Cory S . 1986 Adv. Cancer Res. 47: 189–234
Dang CV . 1999 Mol. Cell. Biol. 19: 1–11
Dang CV, Dolde C, Gillison ML, Kato GJ . 1992 Proc. Natl. Acad. Sci. USA 89: 599–602
Dang CV, Lee WM . 1988 Mol. Cell. Biol. 8: 4048–4054
Dang CV, Semenza GL . 1999 Trends Biochem. Sci. 24: 68–72
Davis AC, Wims M, Spotts GD, Hann SR, Bradley A . 1993 Genes Dev. 7: 671–682
de Alboran IM, O'Hagan RC, Gartner F, Malynn B, Davidson L, Rickert R, Rajewsky K, DePinho RA, Alt FW . 2001 Immunity 14: 45–55
Drach J, Kaufmann H, Urbauer E, Schreiber S, Ackermann J, Huber H . 2000 J. Cancer Res. Clin. Onc. 126: 441–447
Dyson PJ, Littlewood TD, Forster A, Rabbitts TH . 1985 EMBO J. 4: 2885–2891
Dyson PJ, Rabbitts TH . 1985 Proc. Natl. Acad. Sci. USA 82: 1984–1988
Eick D, Bornkamm GW . 1986 Nucl. Acids Res. 14: 8331–8345
Eilers M, Schirm S, Bishop JM . 1991 EMBO J. 10: 133–141
Erikson J, ar-Rushdi A, Drwinga HL, Nowell PC, Croce CM . 1983 Proc. Natl. Acad. Sci. USA 80: 820–824
Evan GI, Wyllie AH, Gilbert CS, Littlewood TD, Land H, Brooks M, Waters CM, Penn LZ, Hancock DC . 1992 Cell 69: 119–128
Felsher DW, Zetterberg A, Zhu J, Tlsty T, Bishop JM . 2000 Proc. Natl. Acad. Sci. USA 97: 10544–10548
Ferre-D'Amare AR, Pognonec P, Roeder RG, Burley SK . 1994 EMBO J. 13: 180–189
Ferre-D'Amare AR, Prendergast GC, Ziff EB, Burley SK . 1993 Nature 363: 38–45
Flinn EM, Busch CM, Wright AP . 1998 Mol. Cell. Biol. 18: 5961–5969
Galaktionov K, Chen X, Beach D . 1996 Nature 382: 511–517
Gallant P, Shiio Y, Cheng PF, Parkhurst SM, Eisenman RN . 1996 Science 274: 1523–1527
Gandarillas A, Davies D, Blanchard JM . 2000 Oncogene 19: 3278–3289
Gardner LB, Li Q, Park MS, Flanagan WM, Semenza GL, Dang CV . 2001 J. Biol. Chem. 276: 7919–7926
Gaubatz S, Imhof A, Dosch R, Werner O, Mitchell P, Buettner R, Eilers M . 1995 EMBO J. 14: 1508–1519
Geltinger C, Hortnagel K, Polack A . 1996 Gene Exp. 6: 113–127
Gerbitz A, Mautner J, Geltinger C, Hortnagel K, Christoph B, Asenbauer H, Klobeck G, Polack A, Bornkamm GW . 1999 Oncogene 18: 1745–1753
Gibson AW, Ye R, Johnston RN, Browder LW . 1992 Oncogene 7: 2363–2367
Grandori C, Cowley SM, James LP, Eisenman RN . 2000 Ann. Rev. Cell. Dev. Biol. 16: 653–699
Greasley P, Bonnard C, Amati B . 2000 Nucleic Acids Res. 28: 446–453
Greenberg RA, O'Hagan RC, Deng H, Xiao Q, Hann SR, Adams RR, Lichtsteiner S, Chin L, Morin GB, DePinho RA . 1999 Oncogene 18: 1219–1226
Gregory MA, Hann SR . 2000 Mol. Cell. Biol. 20: 2423–2435
Gu W, Bhatia K, Magrath IT, Dang CV, Dalla-Favera R . 1994 Science 264: 251–254
Guo QM, Malek RL, Kim S, Chiao C, He M, Ruffy M, Sanka K, Lee NH, Dang CV, Liu ET . 2000 Cancer Res. 60: 5922–5928
Hahn WC, Counter CM, Lundberg AS, Beijersbergen RL, Brooks MW, Weinberg RA . 1999 Nature 400: 464–468
Hann SR, Sloan-Brown K, Spotts GD . 1992 Genes Dev. 6: 1229–1240
Hatton KS, Mahon K, Chin L, Chiu FC, Lee HW, Peng D, Morgenbesser SD, Horner J, DePinho RA . 1996 Mol. Cell. Biol. 16: 1794–1804
Hayday AC, Gillies SD, Saito H, Wood C, Wiman D, Hayward WS, Tonegawa S . 1984 Nature 307: 334–340
He TC, Sparks AB, Rago C, Hermeking H, Zawel L, da Costa LT, Morin PJ, Vogelstein B, Kinzler KW . 1998 Science 281: 1509–1512
Henriksson M, Luscher B . 1996 Adv. Cancer Res. 68: 109–182
Hermeking H, Rago C, Schuhmacher M, Li Q, Barrett JF, Obaya AJ, O'Connell BC, Mateyak MK, Tam W, Kohlhuber F, et al. 2000 Proc. Natl. Acad. Sci. USA 97: 2229–2234
Hoang AT, Lutterbach B, Lewis BC, Yano T, Chou TY, Barrett JF, Raffeld M, Hann SR, Dang CV . 1995 Mol. Cell. Biol. 15: 4031–4042
Hueber AO, Zornig M, Lyon D, Suda T, Nagata S, Evan GI . 1997 Science 278: 1305–1309
Inghirami G, Grignani F, Sternas L, Lombardi L, Knowles DM, Dalla-Favera R . 1990 Science 250: 682–686
Iritani BM, Eisenman RN . 1999 Proc. Natl. Acad. Sci. USA 96: 13180–13185
Janz A, Sevignani C, Kenyon K, Ngo CV, Thomas-Tikhonenko A . 2000 Nucleic Acids Res. 28: 2268–2275
Ji L, Arcinas M, Boxer LM . 1994 Mol. Cell. Biol. 14: 7967–7974
Ji L, Arcinas M, Boxer LM . 1995 J. Biol. Chem. 270: 13392–13398
Johnston LA, Prober DA, Edgar BA, Eisenman RN, Gallant P . 1999 Cell 98: 779–790
Judde J-G, Max EE . 1992 Mol. Cell. Biol. 12: 5206–5216
Juin P, Hueber AO, Littlewood T, Evan G . 1999 Genes Dev. 13: 1367–1381
Kan O, Baldwin SA, Whetton AD . 1994 J. Exp. Med. 180: 917–923
Kanda K, Hu H-M, Zhang L, Grandchamps J, Boxer LM . 2000 J. Biol. Chem. 275: 32338–32346
Kasibhatla S, Beere HM, Brunner T, Echeverri F, Green DR . 2000 Curr. Biol. 10: 1205–1208
Kato GJ, Barrett J, Villa-Garcia M, Dang CV . 1990 Mol. Cell. Biol. 10: 5914–5920
Kato GJ, Lee WM, Chen LL, Dang CV . 1992 Genes Dev. 6: 81–92
Kessler DJ, Spicer DB, La Rosa FA, Sonenshein G . 1992 Oncogene 7: 2447–2453
Kim S, Li Q, Dang CV, Lee LA . 2000 Proc. Natl. Acad. Sci. USA 97: 11198–11202
Kolligs FT, Hu G, Dang CV, Fearon ER . 1999 Mol. Cell. Biol. 19: 5696–5706
Kolligs FT, Kolligs B, Hajra KM, Hu G, Tani M, Cho KR, Fearon ER . 2000 Genes Dev. 14: 1319–1331
Koskinen PJ, Ayer DE, Eisenman RN . 1995 Cell Growth Differ. 6: 623–629
Krumm A, Meulia T, Brunvand M, Groudine M . 1992 Genes Dev. 6: 2201–2213
Kyo S, Takakura M, Taira T, Kanaya T, Itoh H, Yutsudo M, Ariga H, Inoue M . 2000 Nucleic Acids Res. 28: 669–677
Lahoz EG, Xu L, Schreiber-Agus N, DePinho RA . 1994 Proc. Natl. Acad. Sci. USA 91: 5503–5507
Land H, Chen AC, Morgenstern JP, Parada LF, Weinberg RA . 1986 Mol. Cell. Biol. 6: 1917–1925
La Rosa FA, Pierce JW, Sonenshein GE . 1994 Mol. Cell. Biol. 14: 1039–1044
Lasorella A, Noseda M, Beyna M, Iavarone A . 2000 Nature 407: 592–598
Leder P, Battey J, Lenoir G . 1983 Science 222: 765–771
Lee LA, Dolde C, Barrett J, Wu CS, Dang CV . 1996 J. Clin. Invest. 97: 1687–1695
Lee TC, Li L, Philipson L, Ziff EB . 1997 Proc. Natl. Acad. Sci. USA 94: 12886–12891
Lewis BC, Shim H, Li Q, Wu CS, Lee LA, Maity A, Dang CV . 1997 Mol. Cell. Biol. 17: 4967–4978
Li LH, Nerlov C, Prendergast G, MacGregor D, Ziff EB . 1994 EMBO J. 13: 4070–4079
Li Q, Dang CV . 1999 Mol. Cell. Biol. 19: 5339–5351
Lombardi L, Newcomb EW, Dalla-Favera R . 1987 Cell 49: 161–170
Lutterbach B, Hann SR . 1994 Mol. Cell. Biol. 14: 5510–5522
Madisen L, Groudine M . 1994 Genes Dev. 8: 2212–2226
Madisen L, Krumm A, Hebber TR, Groudine M . 1998 Mol. Cell. Biol. 18: 6281–6292
Magrath I . 1990 Adv. Cancer Res. 55: 133–270
Marcu KB, Bossone SA, Patel AJ . 1992 Ann. Rev. Biochem. 61: 809–860
Mai S, Jalava A . 1994 Nucleic Acids Res. 22: 2264–2273
Malynn BA, de Alboran IM, O'Hagan RC, Bronson R, Davidson L, DePinho RA, Alt FW . 2000 Genes Dev. 14: 1390–1399
Mateyak MK, Obaya AJ, Adachi S, Sedivy JM . 1997 Cell Growth Differ. 8: 1039–1048
Mateyak MK, Obaya AJ, Sedivy JM . 1999 Mol. Cell. Biol. 19: 4672–4683
McMahon SB, Van Buskirk HA, Dugan KA, Copeland TD, Cole MD . 1998 Cell 94: 363–374
McMahon SB, Wood MA, Cole MD . 2000 Mol. Cell. Biol. 20: 556–562
Michaelson JS, Giannini SL, Birshtein BK . 1995 Nucl. Acids Res. 23: 975–981
Miltenberger RJ, Sukow KA, Farnham PJ . 1995 Mol. Cell. Biol. 15: 2527–2535
Mink S, Mutschler B, Weiskirchen R, Bister K, Klempnauer KH . 1996 Proc. Natl. Acad. Sci. USA 93: 6635–6640
Mitchell KO, Ricci MS, Miyashita T, Dicker DT, Jin Z, Reed JC, El-Deiry WS . 2000 Cancer Res. 60: 6318–6325
Muller B, Stappert H, Reth M . 1990 Eur. J. Immunol. 20: 1409–1411
Muller D, Bouchard C, Rudolph B, Steiner P, Stuckmann I, Saffrich R, Ansorge W, Huttner W, Eilers M . 1997 Oncogene 15: 2561–2576
Nepveu A, Marcu KB . 1986 EMBO J. 5: 2859–2865
Nesbit CE, Tersak JM, Prochownik EV . 1999 Oncogene 18: 3004–3016
Ngo CV, Gee M, Akhtar N, Yu D, Volpert O, Auerbach R, Thomas-Tikhonenko A . 2000 Cell Growth Differ. 11: 201–210
Nishikura K, ar-Rushdi A, Erikson J, Watt R, Rovera G, Croce CM . 1983 Proc. Natl. Acad. Sci. USA 80: 4822–4826
Nussenzweig MD, Schmidt E, Shaw AC, Sinn E, Campos-Torres J, Mathey-Prevot B, Pattengale PK, Leder P . 1988 Nature 336: 446–450
O'Hagan RC, Ohh M, David G, de Alboran IM, Alt FW, Kaelin WG, DePinho RA . 2000a Genes Dev. 14: 2185–2191
O'Hagan RC, Schreiber-Agus N, Chen K, David G, Engelman JA, Schwab R, Alland L, Thomson C, Ronning DR, Sacchettini JC. et al . 2000b Nat. Genet. 24: 113–119
Oster SK, Marhin WW, Asker C, Facchini LM, Dion PA, Funa K, Post M, Sedivy JM, Penn LZ . 2000 Mol. Cell. Biol. 20: 6768–6778
Osthus RC, Shim H, Kim S, Li Q, Reddy R, Mukherjee M, Xu Y, Wonsey D, Lee LA, Dang CV . 2000 J. Biol. Chem. 275: 21797–21800
Packham G, Cleveland JL . 1994 Mol. Cell. Biol. 14: 5741–5747
Palomo C, Zou X, Nicholson IC, Butzler C, Bruggemann M . 1999 Cancer Res. 59: 5625–5628
Pelengaris S, Littlewood T, Khan M, Elia G, Evan G . 1999 Mol. Cell. 3: 565–577
Pelicci PG, Knowles DM, Magrath I, Dalla-Favera R . 1986 Proc. Natl. Acad. Sci. USA 83: 2984–2988
Perez-Roger I, Kim SH, Griffiths B, Sewing A, Land H . 1999 EMBO J. 18: 5310–5320
Perez-Roger I, Solomon DL, Sewing A, Land H . 1997 Oncogene 14: 2373–2381
Pettersson S, Cook GP, Bruggemann M, Williams GT, Neuberger MS . 1990 Nature 344: 165–167
Polack A, Feederle R, Klobeck G, Hortnagel K . 1993 EMBO J. 12: 3913–3920
Prendergast GC, Ziff EB . 1992 Trends Genet. 8: 91–96
Prochownik EV, Eagle Grove L, Deubler D, Zhu XL, Stephenson RA, Rohr LR, Yin X, Brothman AR . 1998 Genes Chromosomes Cancer 22: 295–304
Pusch O, Bernaschek G, Eilers M, Hengstschlager M . 1997a Oncogene 15: 649–656
Pusch O, Soucek T, Hengstschlager-Ottnad E, Bernaschek G, Hengstschlager M . 1997b DNA Cell. Biol. 16: 737–747
Queva C, Hurlin PJ, Foley KP, Eisenman RN . 1998 Oncogene 16: 967–977
Rabbits TH, Forster A, Hamlyn P, Baer R . 1984 Nature 309: 592–597
Reisman D, Elkind NB, Roy B, Beamon J, Rotter V . 1993 Cell Growth Differ. 4: 57–65
Rohn JL, Hueber AO, McCarthy NJ, Lyon D, Navarro P, Burgering BM, Evan GI . 1998 Oncogene 17: 2811–2818
Rosenwald IB, Rhoads DB, Callanan LD, Isselbacher KJ, Schmidt EV . 1993 Proc. Natl. Acad. Sci. USA 90: 6175–6178
Roussel MF . 1998 Adv. Cancer Res. 74: 1–24
Roussel MF, Ashmun RA, Sherr CJ, Eisenman RN, Ayer DE . 1996 Mol. Cell. Biol. 16: 2796–2801
Saleque S, Singh M, Little RD, Giannini SL, Michaelson JS, Birshtein BK . 1997 J. Immunol. 158: 4780–4787
Saphire AC, Bark SJ, Gerace L . 1998 J. Biol. Chem. 273: 29764–29769
Schmidt EV . 1999 Oncogene 18: 2988–2996
Schreiber-Agus N, Chin L, Chen K, Torres R, Rao G, Guida P, Skoultchi AI, DePinho RA . 1995 Cell 80: 777–786
Schreiber-Agus N, Meng Y, Hoang T, Hou Jr. H, Chen K, Greenberg R, Cordon-Cardo C, Lee HW, DePinho RA . 1998 Nature 393: 483–487
Schreiber-Agus N, Stein D, Chen K, Goltz JS, Stevens L, DePinho RA . 1997 Proc. Natl. Acad. Sci. USA 94: 1235–1240
Schuhmacher M, Kohlhuber F, Holzel M, Kaiser C, Burtscher H, Jarsch M, Bornkamm GW, Laux G, Polack A, Weidle UH, Eick D . 2001 Nucleic Acids Res. 29: 397–406
Schuhmacher M, Staege MS, Pajic A, Polack A, Weidle UH, Bornkamm GW, Eick D, Kohlhuber F . 1999 Curr. Biol. 9: 1255–1258
Schuldiner O, Eden A, Ben-Yosef T, Yanuka O, Simchen G, Benvenisty N . 1996 Proc. Natl. Acad. Sci. USA 93: 7143–7148
Sears R, Leone G, DeGregori J, Nevins JR . 1999 Mol. Cell. 3: 169–179
Serrano M, Lin AW, McCurrach ME, Beach D, Lowe SW . 1997 Cell 88: 593–602
Shim H, Chun YS, Lewis BC, Dang CV . 1998 Proc. Natl. Acad. Sci. USA 95: 1511–1516
Shim H, Dolde C, Lewis BC, Wu CS, Dang G, Jungmann RA, Dalla-Favera R, Dang CV . 1997 Proc. Natl. Acad. Sci. USA 94: 6658–6663
Shiramizu B, Barriga F, Neequaye J, Jafri A, Dalla-Favera R, Neri A, Guttierez M, Levine P, Magrath I . 1991 Blood 77: 1516–1526
Shou Y, Martelli ML, Gabrea A, Qi Y, Brents LA, Rosckle A, Deward G, Kirsch IR, Bergsagel PL, Kuehl WM . 2000 Proc. Natl. Acad. Sci. USA 97: 228–233
Siebenlist U, Henninghausen L, Battey J, Leder P . 1984 Cell 37: 381–391
Spencer CA, Groudine M . 1991 Adv. Cancer Res. 56: 1–47
Spotts GD, Patel SV, Xiao Q, Hann SR . 1997 Mol. Cell. Biol. 17: 1459–1468
Stanton BR, Perkins AS, Tessarollo L, Sassoon DA, Parada LF . 1992 Genes Dev. 6: 2235–2247
Strobl LJ, Eick D . 1992 EMBO J. 11: 3307–3314
Strobl LJ, Kohlhuber F, Mautner J, Polack A, Eick D . 1993 Oncogene 8: 1437–1447
Taub R, Moulding C, Battey J, Murphy W, Vasicek T, Lenoir GM, Leder P . 1984 Cell 36: 339–348
Vlach J, Hennecke S, Alevizopoulos K, Conti D, Amati B . 1996 EMBO J. 15: 6595–6604
Wang J, Xie LY, Allan S, Beach D, Hannon GJ . 1998 Genes Dev. 12: 1769–1774
Wang X, Scott E, Sawyers CL, Friedman AD . 1999 Blood 94: 560–571
Wechsler DS, Shelly CA, Petroff CA, Dang CV . 1997 Cancer Res. 57: 4905–4912
Whitehurst C, Henney HR, Max EE, Schroeder HWJ, Stuber F, Siminovitch KA, Garrard WT . 1992 Nucl. Acids Res. 20: 4929–4930
Wittekindt NE, Hortnagel K, Geltinger C, Polack A . 2000 Nucl. Acids Res. 28: 800–808
Wood LJ, Mukherjee M, Dolde CE, Xu Y, Maher JF, Bunton TE, Williams JB, Resar LM . 2000 Mol. Cell. Biol. 20: 5490–5502
Wu K, Grandori C, Amacker M, Simon-Vermot N, Polack A, Lingner J, Dalla-Favera R . 1999a Nat. Genet. 21: 220–224
Wu KJ, Polack A, Dalla-Favera R . 1999b Science 283: 676–679
Yang BS, Geddes TJ, Pogulis RJ, de Crombrugghe B, Freytag SO . 1991 Mol. Cell. Biol. 11: 2291–2295
Yang BS, Gilbert JD, Freytag SO . 1993 Mol. Cell. Biol. 13: 3093–3102
Yano T, Sander CA, Clark HM, Dolezal MV, Jaffe ES, Raffeld M . 1993 Oncogene 8: 2741–2748
Yin XY, Grove L, Datta NS, Long MW, Prochownik EV . 1999 Oncogene 18: 1177–1184
Yu BW, Ichinose I, Bonham MA, Zajac-Kaye M . 1993 J. Biol. Chem. 268: 19586–19592
Zindy F, Eischen CM, Randle DH, Kamijo T, Cleveland JL, Sherr CJ, Roussel MF . 1998 Genes Dev. 12: 2424–2433
Acknowledgements
Supported by National Institutes of Health Grant CA69322 (LM Boxer), CA51497 (CV Dang) and CA57431 (CV Dang). We thank E Emison, L Gardner, LA Lee and D Wechsler for comments.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Boxer, L., Dang, C. Translocations involving c-myc and c-myc function. Oncogene 20, 5595–5610 (2001). https://doi.org/10.1038/sj.onc.1204595
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1204595
Keywords
This article is cited by
-
Defining the landscape of circular RNAs in neuroblastoma unveils a global suppressive function of MYCN
Nature Communications (2023)
-
Long noncoding RNA: a dazzling dancer in tumor immune microenvironment
Journal of Experimental & Clinical Cancer Research (2020)
-
RIG-I aggravates interstitial fibrosis via c-Myc-mediated fibroblast activation in UUO mice
Journal of Molecular Medicine (2020)
-
A combination of LMO2 negative and CD38 positive is useful for the diagnosis of Burkitt lymphoma
Diagnostic Pathology (2019)
-
A three-platelet mRNA set: MAX, MTURN and HLA-B as biomarker for lung cancer
Journal of Cancer Research and Clinical Oncology (2019)