Abstract
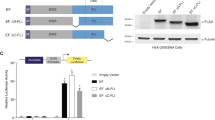
Chromosomal fusion of the N-terminal region of the Ewings Sarcoma Oncogene (EWS-activation-domain, EAD) to the DNA-binding domains of a variety of cellular transcription factors produce oncogenic proteins (EWS-fusion proteins (EFPs)) that cause distinct malignancies. In EFPs, the EAD acts as a potent transcriptional activation domain and this ability is repressed in the context of normal, non-tumorigenic, EWS. Trans-activation by the EAD is therefore a specific characteristic of EFPs and it is thought that EFPs induce tumorigenesis via improper transcriptional activation of cellular genes. Functional elements required for transcriptional activation are dispersed throughout the EAD, as are thirty-one copies of a Degenerate Hexapeptide Repeat (DHR, consensus SYGQQS). This suggests that the EAD contains a highly reiterated functional element related to DHRs. Here we show that in the context of EWS/ATF1, the EFP that causes malignant melanoma of soft parts, trans-cooperation by small regions of the EAD (∼30 residues) results in potent transcriptional activation dependent on the conserved tyrosine residues present in DHRs. These findings provide the first evidence for a role of DHRs in EAD-mediated trans-activation and demonstrate that the EAD represents a novel tyrosine-dependent transcriptional activation domain.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Benuck ML, Li Z, Childs G . 1999 J. Biol. Chem. 274: 25419–25425
Bertolotti A, Melot T, Acker J, Vigneron M, Delattre O, Tora L . 1998 Mol. Cell. Biol. 18: 1489–1497
Blair WS, Bogerd HP, Madore SJ, Cullen BR . 1994 Mol. Cell. Biol. 14: 7226–7234
Brown AD, Lopez-Terrada D, Denny CT, Lee KAW . 1995 Oncogene 10: 1749–1756
Carey M . 1998 Cell 92: 5–8
Carey M, Kolman J, Katz DA, Gradoville L, Barberis L, Miller G . 1992 J. Virol. 66: 4803–4813
Delattre O, Zucman J, Plougastel B, Desmaze C, Melot T, Peter M, Kovar H, Joubert I, de Jong P, Rouleau G, Aurias A, Thomas G . 1992 Nature 359: 162–165
Deloulme JC, Prichard L, Delattre O, Storm DR . 1997 J. Biol. Chem. 272: 27369–27377
Felsch JS, Lane WS, Peralta EG . 1999 Curr. Biol. 9: 485–488
Fujimura Y, Ohno T, Siddique H, Lee L, Rao VN, Reddy ESP . 1996 Oncogene 12: 159–167
Gill G, Pascal E, Tseng ZH, Tjian R . 1994 Proc. Natl. Acad. Sci. 91: 192–196
Guinamard R, Fougereau M, Seckinger P . 1997 Scand. J. Immunol. 45: 587–595
Kim TK, Roeder RG . 1994 Nucleic Acids Res. 22: 251
Kim J, Lee KAW, Pelletier J . 1998 Oncogene 16: 1021–1030
Kim J, Lee JM, Branton PE, Pelletier J . 1999 Proc. Natl. Acad. Sci., USA 96: 14300–14305
Kovar H, Aryee D, Zoubek A . 1999 Curr. Op. Oncol. 11: 275–284
Krajewski W, Lee KAW . 1994 Mol. Cell. Biol. 14: 7204–7210
Lee KAW, Masson N . 1993 Biochem. Biophys. Acta. 1174: 221–233
Lessnick SL, Braun BS, Denny CT, May WA . 1995 Oncogene 10: 423–431
Li KKC, Lee KAW . 2000 J. Biol. Chem. 275: 23053–23058
MacArthur H, Walter G . 1984 J. Virol. 52: 483–491
May WA, Lessnick SL, Braun BS, Klemsz M, Lewis BC, Lunsford LB, Hromas R, Denny CT . 1993 Mol. Cell. Biol. 13: 7393–7398
May WA, Denny CT . 1997 Curr. Top. Micro. Immun. 220: 143–150
Mayer BJ, Jackson PK, Van ER, Baltimore D . 1992 Mol. Cell. Biol. 12: 609–618
Montminy MR, Sevarino KA, Wagner JA, Mandel G, Goodman RH . 1986 Proc. Natl. Acad. Sci., USA 83: 6682–6686
Netzer WJ, Hartl FU . 1997 Nature 388: 343–349
Nyanguile O, Uesugi M, Austin DJ, Verdine GL . 1997 Proc. Natl. Acad. Sci., USA 94: 13402–13406
Ohno T, Rao VN, Reddy ESP . 1993 Cancer Res. 53: 5859–5863
Pan S, Ming KY, Dunn TA, Li KKC, Lee KAW . 1998 Oncogene 16: 1625–1631
Petermann R, Mossier BM, Aryee DNT, Khazak V, Golemis EA, Kovar H . 1998 Oncogene 17: 603–610
Pollock R, Gilman M . 1997 Proc. Natl. Acad. Sci., USA 94: 13388–13389
Prasad DDK, Ouchida M, Lee L, Rao VN, Reddy ESP . 1994 Oncogene 9: 3717–3729
Rauscher III FJ . 1997 Curr. Top. Micro. Immun. 220: 151–162
Regier JL, Shen F, Triezenberg SJ . 1993 Proc. Natl. Acad. Sci., USA 90: 883–887
Ribeiro A, Brown AD, Lee KAW . 1994 J. Biol. Chem. 269: 31124–31128
Ron D . 1997 Curr. Top. Micro. Immun. 220: 131–142
Sanchez-Garcia I, Rabbitts TH . 1994 Proc. Natl. Acad. Sci., USA 91: 7869–7873
Seipel K, Georgiev O, Schaffner W . 1992 EMBO J. 11: 4961–4968
Songyang Z, Shoelson SE, Chaudhuri M, Gish G, Pawson T, Haser WG, King F, Roberts T, Ratnofsky S, Lechleider RJ, Neel BG, Birge RB, Fajardom JE, Chou MM, Hanafusa H, Schaffhausen B, Cantley LC . 1993 Cell 72: 767–778
Songyang Z, Carraway III KL, Eck MJ, Harrison SC, Feldman RA, Mohammadi M, Schlessinger J, Hubbard SR, Smith DP, Eng C, Lorenzo MJ, Ponder BAJ, Mayer BJ, Cantley LC . 1995 Nature 373: 536–539
Tanaka M, Herr W . 1994 Mol. Cell. Biol. 14: 6056–6067
Triezenberg SJ . 1995 Curr. Opin. Genet. Dev. 5: 190–196
Zhang D, Paley AJ, Childs G . 1998 J. Biol. Chem. 273: 18086–18091
Zhou H, Lee KAW . 2001 Oncogene 20: 1519–1524
Zucman J, Delattre O, Desmaze C, Epstein AL, Stenman G, Speleman F, Fletchers CDM, Aurias A, Thomas G . 1993 Nature Genetics 4: 341–345
Acknowledgements
We thank Dr Zhang Mingjie for many discussions during the course of this work, Dr Chris Rock for helpful comments on the manuscript and Kim KC Li for excellent technical assistance. This work was supported by a Hong Kong Government Research Grants Council grant (award HKUST 6106/98M) to KAW Lee.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Feng, L., Lee, K. A repetitive element containing a critical tyrosine residue is required for transcriptional activation by the EWS/ATF1 oncogene. Oncogene 20, 4161–4168 (2001). https://doi.org/10.1038/sj.onc.1204522
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1204522
Keywords
This article is cited by
-
The melanocyte inducing factor MITF is stably expressed in cell lines from human clear cell sarcoma
British Journal of Cancer (2003)
-
Homotypic and heterotypic interactions of EWS, FLI1 and their oncogenic fusion protein
Oncogene (2003)