Abstract
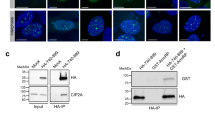
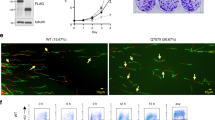
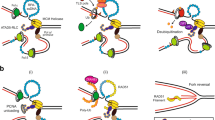
The hHus1 and several hRad proteins are involved in the control of DNA integrity checkpoints, although the mechanisms underlying these processes are unknown. Using a yeast two-hybrid system to detect protein-protein interactions, we found that human proliferating cell nuclear antigen (PCNA), a protein known to function in both DNA replication and repair, interacts with the human checkpoint-related protein Hus1 (hHus1). In human skin fibroblast cells, exposure to ionizing radiation of hydroxyurea triggers translocation of hHus1 from the cytosol to the nucleus, where it associates with PCNA as well as another checkpoint protein, hRad9. This nuclear translocation and the complex formation or hHus1 with PCNA and hRad9 correlate closely with changes in cell cycle distribution in response to radiation exposure. These results suggest that this multi-protein complex may be important for coordinating cell-cycle progression, DNA replication and repair of damaged DNA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
al-Khodairy F and Carr AM. . 1992 EMBO J. 11: 1343–1350.
al-Khodairy F, Fotou E, Sheldrick KS, Griffiths DJ, Lehmann AR and Carr AM. . 1994 Mol. Biol. Cell. 5: 147–160.
Aravind L, Walker DR and Koonin EV. . 1999 Nucleic Acids Res. 27: 1223–1242.
Ayyagari R, Impellizzeri KJ, Yoder BL, Gary SL and Burgers PM. . 1995 Mol. Cell Biol. 15: 4420–4429.
Bravo R and Macdonald-Bravo H. . 1987 J. Cell Biol. 105: 1549–1554.
Caspari T, Dahlen M, Kanter-Smoler G, Lindsay HD, Hofmann K, Papadimitriou K, Sunnerhagen P and Carr AM. . 2000 Mol. Cell Biol., 20: 1254–1262.
Cimprich KA, Shin TB, Keith CT and Schreiber SL. . 1996 Proc. Natl. Acad. Sci. USA 93: 2850–2855.
Eissenberg JC, Ayyagari R, Gomes XV and Burgers PM. . 1997 Mol. Cell Biol. 17: 6367–6378.
Enoch T, Carr AM and Nurse P. . 1992 Genes Dev. 6: 2035–2046.
Estojak J, Brent R and Golemis EA. . 1995 Mol. Cell. Biol. 15: 5820–5829.
Golemis EA, Gyuris J and Brent R. . 1994 In: Current Protocols in Molecular Biology. J. Wiley & Sons, Inc., pp13.14.1–13.14.17.
Griffiths DJ, Barbet NC, McCready S, Lehmann AR and Carr AM. . 1995 EMBO J. 14: 5812–5823.
Hang H and Lieberman HB. . 2000 Genomics 65: 24–33.
Hartwell LH and Kastan MB. . 1994 Science 266: 1821–1828.
Kelman Z and Hurwitz J. . 1998 Trends Biochem. Sci. 23: 236–238.
Komatsu K, Miyashita T, Hang H, Hopkins KM, Zheng W, Cuddeback S, Yamada M, Lieberman HB and Wang HG. . 2000 Nat. Cell. Biol. 2: 1–6.
Kondo T, Matsumoto K and Sugimoto K. . 1999 Mol. Cell Biol. 19: 1136–1143.
Kong XP, Onrust R, O'Donnell M and Kuriyan J. . 1992 Cell 69: 425–437.
Kostrub CF, al-Khodairy F, Ghazizadeh H, Carr AM and Enoch T. . 1997 Mol. Gen. Genet. 254: 389–399.
Kostrub CF, Knudsen K, Subramani S and Enoch T. . 1998 EMBO J. 17: 2055–2066.
Krishna TS, Kong XP, Gary S, Burgers PM and Kuriyan J. . 1994 Cell 79: 1233–1243.
Lieberman HB, Hopkins KM, Nass M, Demetrick D and Davey S. . 1996 Proc. Natl. Acad. Sci. USA 93: 13890–13895.
Nojima H. . 1997 Hum. Cell 10: 221–230.
Parker AE, Van de Weyer I, Laus MC, Oostveen I, Yon J, Verhasselt P and Luyten WH. . 1998 J. Biol. Chem. 273: 18332–18339.
Rowley R, Subramani S and Young PG. . 1992 EMBO J. 11: 1335–1342.
Russell, P. . 1998 Trends Biochem. Sci., 23: 399–402.
St. Onge RP, Udell CM, Casselman R and Davey S. . 1999 Mol. Biol. Cell 10: 1985–1995.
Stillman B. . 1994 Cell 78: 725–728.
Thelen MP, Venclovas C and Fidelis K. . 1999 Cell 96: 769–770.
Tsurimoto T. . 1999 Front Biosci. 4: D849–D858.
Udell CM, Lee SK and Davey S. . 1998 Nucleic Acids Res. 26: 3971–3976.
Venclovas C and Thelen MP. . 2000 Nucleic Acids Res. 28: 2481–2493.
Volkmer E and Karnitz LM. . 1999 J. Biol. Chem. 274: 567–570.
Waga S and Stillman B. . 1998 Annu. Rev. Biochem. 67: 721–751.
Waseem NH, Labib K, Nurse P and Lane DP. . 1992 EMBO J. 11: 5111–5120.
Acknowledgements
We thank Nikola Valkov for assistance with fluorescence confocal microscopy, the Molecular Biology, Flow Cytometry and Molecular Imaging core facilities at the Moffitt Cancer Center and Research Institute, and the NIH (CA82197, CA72694, GM52493, and CA68446) for support.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Komatsu, K., Wharton, W., Hang, H. et al. PCNA interacts with hHus1/hRad9 in response to DNA damage and replication inhibition. Oncogene 19, 5291–5297 (2000). https://doi.org/10.1038/sj.onc.1203901
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1203901
Keywords
This article is cited by
-
Agnoprotein of polyomavirus BK interacts with proliferating cell nuclear antigen and inhibits DNA replication
Virology Journal (2015)
-
The Ron receptor tyrosine kinase activates c-Abl to promote cell proliferation through tyrosine phosphorylation of PCNA in breast cancer
Oncogene (2014)
-
Disruption of the Rad9/Rad1/Hus1 (9–1–1) complex leads to checkpoint signaling and replication defects
Oncogene (2004)
-
Interaction between proliferating cell nuclear antigen and JUN-activation-domain-binding protein 1 in the meristem of rice, Oryza sativa L.
Planta (2003)
-
Colocalization of human Rad17 and PCNA in late S phase of the cell cycle upon replication block
Oncogene (2002)