Abstract
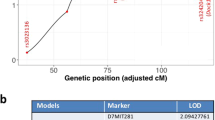
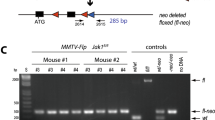
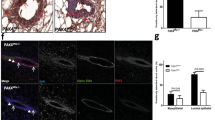
We have examined defects in mammary development and tumorigenesis in a transgenic model expressing the c-myc gene under the MMTV–LTR promoter. The stochastic tumors which arise from hyperplastic ductal and lobular lesions in this model are characterized by high rates both of apoptosis and of chromosomal instability. Since the p53 gene product is thought to be central in the maintenance of genomic integrity, in part due to its ability to induce apoptosis in cells harboring DNA damage, we examined its expression and possible mutation. Initially, we observed that unmutated p53 is strongly expressed in premalignant mammary glands and in mammary tumors derived from the MMTV-c-myc strain. We then mated the MMTV-myc strain to a p53-deficient strain as a means of examining the effect of this lesion on mammary development and tumorigenesis in the context of c-myc overexpression. A lack of both p53 alleles in the presence of c-myc overexpression resulted in a dramatic hyerplastic alteration in mammary gland development. Specifically, in female bitransgenic MMTV-c-myc/p53 null mice (MMTV-myc/p53−/−), lobular hyperplasias were observed at almost every ductal end bud as early as 32 days of age. In contrast, only mild ductal and lobular hyperplasias were seen in MMTV-myc mice that contained both p53 alleles (MMTV-myc/p53+/+); an intermediate phenotype occurred in mice with a single intact (MMTV-myc/p53+/−) p53 allele. Mammary carcinomas arose with a high frequency in MMTV-myc/p53+/− mice; the tumors were comparable in frequency, histology and apoptotic index to the tumors in MMTV-myc/p53+/+ mice. Also, as previously observed (), lymphomas arose with extremely short latency in MMTV-myc/ p53−/− mice, precluding study of the fate of their hyperplastic mammary lesions in situ. The frequency of p53 mutations in MMTV-myc/p53+/+ and MMTV-myc/p53+/− mammary tumors and in cell lines derived from these tumors was examined by direct sequencing. No point mutations or deletions in p53 were observed in mammary tumors or cell lines from either genotype. Finally, a detailed chromosomal analysis using multicolor spectral karyotyping (SKY) revealed that there were multiple chromosomal alterations in the c-myc-overexpressing cells that contained either one or two unmutated p53 alleles. Variable ploidy changes, a common translocation of chromosome 11, and other chromosomal aberrations were observed. Our data thus support an interaction between c-Myc and p53 in mammary development, but suggest that loss of p53 is required neither for c-myc-dependent tumorigenesis nor for c-myc-dependent chromosomal instability.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
McCormack, S., Weaver, Z., Deming, S. et al. Myc/p53 interactions in transgenic mouse mammary development, tumorigenesis and chromosomal instability. Oncogene 16, 2755–2766 (1998). https://doi.org/10.1038/sj.onc.1201804
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1201804
Keywords
This article is cited by
-
IL-25 blockade inhibits metastasis in breast cancer
Protein & Cell (2017)
-
The epigenetic modifier JMJD6 is amplified in mammary tumors and cooperates with c-Myc to enhance cellular transformation, tumor progression, and metastasis
Clinical Epigenetics (2016)
-
A mouse model with T58A mutations in Myc reduces the dependence on KRas mutations and has similarities to claudin-low human breast cancer
Oncogene (2013)
-
MYC gene amplification is often acquired in lethal distant breast cancer metastases of unamplified primary tumors
Modern Pathology (2012)
-
Unlocking the power of cross-species genomic analyses: identification of evolutionarily conserved breast cancer networks and validation of preclinical models
Breast Cancer Research (2008)