Abstract
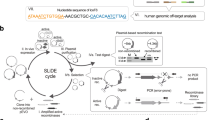
A de novo insertion of an Alu repeated DNA element was found within exon V of the factor IX gene in a patient with severe haemophilia B. The element interrupts the reading frame of the mature factor IX at glutamic acid 96 resulting in a stop codon within the inserted sequence. The Alu repeat is 322 bp long, and the 5′ region is shortened by 38 bp. The insertion created a target site duplication of 15 bp consistent with retro-position, and contains a pure polyadenine tract of at least 78 residues at the 3′ end. The nucleotide sequence agrees with a consensus for an Alu subfamily which is evolutionarily the most recently inserted, suggesting that it is an exact copy of a putative source gene. These observations indicate that retro-position of Alu elements is a continual process and a mechanism for generating human genetic defects.
Similar content being viewed by others
Introduction
The Alu family is the predominant short interspersed repetitive DNA element (SINE) in primates, with more than 500,000 copies in the haploid human genome [1]. An Alu repeat is approximately 300 bp long, and is separated into two halves by a short A-rich region. Alu elements also contain an A-rich 3′ end of variable length, and are flanked by short direct repeats at the sites of integration into the genome [1]. They are thought to have evolved from the polymerase-III-transcribed 7SL RNA gene [2]. Alu repeats have been divided into subfamilies of related sequences [3–7]. The sequence divergence of the subfamilies suggests that they have been inserted into the genome at different evolutionary periods. A group of Alu elements refered to as the ‘new’ [8], human-specific (HS) [9] or predicted variant (PV) [10] Alu subfamily is among the most recently inserted Alu repeats because many members are present only in the human genome and some are polymorphic insertions in the human population. We will utilize the HS terminology when referring to this subgroup of Alu elements.
Alu repeats are considered to be retroposons, sequences that transpose through the reverse transcription of a RNA intermediate [11]. It has been difficult, however, to directly demonstrate in vivo transcription or transposition of an Alu element. A mRNA corresponding to the consensus sequence of the HS subfamily members has been identified, suggesting that there is transcription of an active Alu source gene [10]. LINE-1 or L1 long interspersed repetitive elements may provide the reverse transcriptase to transpose Alu elements in the mammalian genome [12]. A recent report of a new insertion of a HS Alu element in an intron of the type 1 neurofibromatosis (NF1) gene [13] indicates that transposition of Alu elements is a continual process. We describe here the first de novo insertion of an Alu element belonging to the HS subfamily within the coding region of a gene. The insertion was found in the factor IX gene of a patient with severe haemophilia B, an X-linked inherited disorder involving defects in the blood coagulation factor IX.
Materials and Methods
Patient
The patient (HB-7) suffers from severe haemophilia B with factor IX coagulation and antigen < 1 U/dl. He is the only haemophiliac in the family and factor IX levels in the other members are normal, except in his mother who has low factor IX activity and antigen of 45 and 52% of normal values, respectively.
Southern Blotting
Genomic DNA from members of the family, HB-7 and normal controls were isolated from white blood cells [14]. Ten µg of DNA were digested to completion with the restriction enzymes TaqI, XmnI, HindIII, EcoRI and BcII according to the manufacturer’s recommendation (Boehringer, Mannheim, FRG). Southern blots were performed essentially as described in Sambrook et al. [15]. The blots of TaqI- and XmnI-digested DNA were hybridized with a 2.5-kb HindIII-EcoRI-digested genomic fragment (probe pVIII) [16] containing exon IV of the human factor IX gene and flanking introns. The blots of HincIII-, EcoRI- and BcII-digested DNA were hybridized with a full-length factor IX cDNA [16] which is 1,981 bp long and corresponds to the coding region and about half of the 3′ non-translated region of the factor IX mRNA.
Polymerase Chain Reaction (PCR)
The primer sets pHB-12 (5′-CCCAATGTATATTTGACCCA-3′; nucleotides 17569–17589 [17]) and pHB-11 (5′-TGCTGAAGTTTCAGATACAG-3′; nucleotides 17830–17850) were used to amplify exon V and flanking introns by PCR [18] from nucleotides 17569 to 17850 in the factor IX gene [17]. Reaction mixtures (100 µl) contained approximately 250 ng genomic DNA, 10 pmol of each oligonucleotide primer, 2.5 U Taq polymerase (Perkin Elmer-Cetus, Emeryville, Calif., USA), 10 mM Tris (pH 8.3), 50 mM KCl, 1.5 mM MgCl2, 200 µM of each dNTP and 0.01% gelatin. Samples were denatured at 94°C for 5 min and amplified for 30 cycles of 94°C, 55°C and 72°C, each for 60 s. The final 72°C incubation was extended to 7 min. Aliquots (10 µl) of the products were analysed on a 6.5% Polyacrylamide gel stained with ethidium bromide.
Nucleotide Sequence Analysis
The DNA fragment from the patient HB-7 amplified with primers pHB-11 and pHB-12 was eluted from the Polyacrylamide gel and 50 ng were subjected to 30 asymmetric PCR cycles [19]. The single-stranded product was purified and concentrated in a Centricon-100 column (Amicon, Danvers, Mass., USA) and 7 µl were sequenced by the Sanger dideoxy procedure using the Sequenase kit (US Biochemical Corporation, Cleveland, Ohio, USA). Sequencing was done with primers pHB-11 and pHB-12 as well as two internal primers pHB-22 (5′-CATGTAACATTAACAAAT-3′; nucleotides 17675–17695) and pHB-23 (5′-TACAGGAGCAAACCACCTTG; nucleotides 17749–17789).
Results and Discussion
Three generations of a French family with a single severe haemophilia B member (HB-7) were studied to determine the carrier status of the haemophiliac’s aunts. Genomic DNA from the family was digested with the enzymes TaqI and XmnI, and restriction-fragment-length polymorphisms (RFLPs) were examined by Southern blotting. DNA digested with TaqI shows polymorphic bands at 1.8 (A) and 1.3 (a) kb, and XmnI at 11.5 (B) and 6.5 (b) kb. The haplotype of HB-7 was aB and segregation of the alleles demonstrated that the haplotype arrangement was inherited from his grandfather (data not shown). The mutation is suspected to have originated in the grandfather’s gametes because the mother’s phenotype suggests she is carrying the defective gene.
We then searched for rearrangements of the factor IX gene. Southern blotting of Eco-RI-, HindIII- and BcII-digested DNA from HB-7 and a normal individual were hybrizided with a complete factor IX cDNA probe. A 5.2-kb HindIII fragment, a 4.8-kb EcoRI fragment as well as a 1.7-kb BclI fragment include exon V of the factor IX gene. These fragments were enlarged by approximately 300 bp of DNA in HB-7 (data not shown).
Exon V was amplified from genomic DNA of HB-7 by PCR. The normal 282-bp fragment is replaced by one of approximately 600 bp (fig. 1). PCR analysis of DNA from the family confirmed that the alteration is present only in HB-7 and his mother, indicating that the insertion is de novo and the causative mutation. Nucleotide sequencing demonstrated that the inserted element is 322 bp long and is a member of the Alu family repeats (fig. 2). The insertion is in the sense direction and interrupts the reading frame at Glu 96 of the mature factor IX resulting in an inframe stop codon (TAA) at nucleotides 77–79 within the Alu element. The alteration of the reading frame is probably the cause of the disease in HB-7, however, the Alu element in the intron of the NF1 gene resulted in aberrant splicing [13] but was in the antisense direction. Many antisense Alu elements have potential cryptic acceptor sites which may introduce new splice sites and affect the processing of the primary transcript [20].
Exon V of the factor IX gene amplified from genomic DNA from members of family HB-7. Nucleotides 17569 to 17850 in the factor IX gene were amplified by the PCR reaction with primers pHB-11 and pHB-12. Products of the PCR reaction were electrophoresed on a 6.5% Polyacrylamide gel and the pattern visualized by staining the gel with ethidium bromide. The normal 282-bp band seen in III2 is replaced by a 617-bp fragment in DNA from HB-7 (III1). The mother (III) displays two bands indicating that she is a carrier. Lane M = HaeIII-digested φX174 phage DNA.
Direct genomic sequencing and characteristics of the inserted Alu element HB-7. The sequence of the insertion using primer pHB-12 is shown. From bottom to top: the 5′ part of exon V including one of the 15-nucleotide repeats of the factor IX gene, followed by the Alu left monomer truncated by 38 bp at its 5′ end, a middle A-rich region, the Alu right monomer and at least 78 adenine residues.
The structure of Alu element HB-7 favours insertion by retrotransposition. The element is flanked by perfect 15-bp duplications of the factor IX target site sequence (fig. 3) characteristic of insertions of mobile elements into staggered single-stranded nicks at new genomic locations [21]. Although Alu HB-7 is truncated at the 5′ terminus of 38 bp, the target site duplication abuts the 5′ end, indicating premature termination of reverse transcription, or transcription of an incomplete RNA. The direct repeats are not A-T rich as is the case for most of the recently inserted Alu elements [22]. Nevertheless, the sequence surrounding the repeats is predominantly of A and T residues (fig. 3), agreeing with the preferential insertion of Alu elements into short A-T-rich regions [21]. Furthermore, the direct repeats have the sequence 5′-GANx-3′ that has been shown to be a highly specific target site for insertion of many members from the HS subfamily [23].
Location of the Alu insertion in exon V of the factor IX gene. A schematic of the 5’end of exon V is shown at the top. The 15-bp target site is boxed and the A-T-rich sequences flanking the target site are indicated. Below the nucleotide sequence is the amino acid sequence starting with Asp 85 in the mature factor IX. Alu element HB-7 inserted between nucleotides 17688 and 17702 of exon V resulting in 15-bp direct repeats. The arrow indicates the 5′-3′ orientation of the Alu insertion. The sequence of Alu element HB-7 begins at position 39 of a complete Alu sequence and is followed at position 282 by the poly(A) tract. The insert interrupts the CAG codon for Glu 96 in the mature factor IX resulting in an inframe stop codon (TAA) at nucleotides 77 to 79 within the sequence of Alu element HB-7.
Alu element HB-7 has the sequence diagnostic of the HS Alu subfamily (fig. 4). Members of the HS subfamily share five nucleotide substitutions that clearly segregate this subgroup from other Alu elements [8, 10, 22, 23]. The homogeneity of the HS Alu subfamily indicates that a consensus should match the sequence of a putative transcriptionally active source gene. Alu element HB-7 differs from the sequence only by one additional adenine residue in the middle A-rich region (fig. 4), indicating that Alu HB-7 is an exact copy of a source gene. This additional adenine residue in the sequence of HB-7 suggests multiple or dimorphic source genes. The only other report of a de novo insertion of a HS subfamily member [13] was also nearly an identical match to the consensus sequence (fig. 4) and contained the same additional adenine residue, implying that active retroposition is restricted to a very small set of closely related source genes. However, the existence of other distinct source genes is suggested by the identification of recently inserted Alu elements in the human C1 inhibitor locus and Cholinesterase gene [24, 25] which clearly are not members of the HS subfamily.
Comparison of Alu element HB-7 with Alu consensus sequences and the new Alu insertion in the NF1 gene. The Alu consensus in the first line is based on 168 human Alu sequences [6]. Alu class IV is a consensus sequence for an evolutionarily recent branch of Alu elements [7]. The HS Alu consensus sequence is derived from the most recently formed subfamily of Alu sequences [8–10]. Sequences for the de novo inserted Alu element in the NF1 (Alu NF1) [13] and factor IX gene (Alu HB7) are shown in the bottom lines. Identical nucleotides are indicated by dashes, absent nucleotides by asterisks and substitutions with the appropriate nucleotide. Dots refer to the 5 diagnostic nucleotide substitutions that separate the HS subfamily members from other Alu subgroups. The additional adenine residue in the middle A-rich region found in elements HB-7 and NF1 is underlined.
HS Alu elements differ from other Alu subfamilies in that the majority of HS members have only adenine residues, varying from 11 to 37, at the 3′ end, suggesting their recent origin [9, 21, 23]. The new Alu element in the NF1 gene has a pure 3′-poly(A) stretch of > 40 residues [13], and Alu element HB-7 has at least 78 adenine residues at its 3′ end, these being the longest poly(A) tails reported for members of the HS subfamily. Because there are no known polyadenylation signals in Alu elements [26], it has been hypothesized that the A-rich 3′ end is contained in the sequence of the source gene [27, 28]. The different tail lengths are due to random self-priming of the reverse-transcription reaction. The 78 adenine residues found in Alu element HB-7 imply that the 3′ end of the source gene would have to be at least 80 bp long, allowing for a minimal number of residues for self-priming. However, the long, pure poly(A) tract in Alu HB-7 and the considerable variability in the length of these tracts in other HS subfamily members also support the alternative hypothesis that the poly(A) tails are added and processed through post-transcriptional mechanisms [10–11]. Retroposition of Alu elements is considered to be a rare event estimated at 100 to 200 per million years [9]. Alu element HB-7 and the other de novo insertion of a HS subfamily member in the NF1 gene [13] result in deleterious mutations. Such retroposons would not be retained in the population, suggesting that the frequency of retroposition of Alu repeats may be somewhat higher than that predicted by analysis of fixed members of the family. Reports of insertions of L1 elements in the blood coagulation factor VIII gene [29–31] resulting in haemophilia A indicate that retroposition may be a significant but uncommon event for the generation of mutations. For example, the factor VIII, factor IX and cystic-fibrosis-transmembrane-conductance regulatory genes are among the most intensively studied, and over 600 disease-causing mutations have been identified. Retroposition accounts for three of these mutations, including two insertions of L1 elements in the factor VIII gene [29] and the Alu element reported here. The allele which was involved with insertions of both Alu and L1 elements was paternal in origin suggesting that the retrotransposition event occurred in the paternal gametes. Since the tools to efficiently study disease genes have been available for less than a decade it is expected that more cases of retroposition of HS subfamily members will be associated with gene defects.
References
Deininger PL: SINEs: short interspersed repeated DNA elements in higher eucaryotes; in Berg DE, Howe MM (eds): Mobile DNA. Washington, American Society for Microbiology, 1989, pp 619–636.
Ullu E, Tschudi C: Alu sequences are processed 7SL RNA genes. Nature 1984;312:171–172
Labuda D, Striker G: Sequence conservation in Alu evolution. Nucleic Acids Res 1989;17:2477–2491
Jurka J, Smith T: A fundamental division in the Alu family of repeated sequences. Proc Natl Acad Sci USA 1988;85:4775–4778
Slagel V, Flemington E, Traina-Dorge V, Bradshaw H Jr, Deininger PL: Clustering and subfamily relationships of the Alu family in the human genome. Mol Biol Evol 1987;4:19–29
Quentin Y: The Alu family developed through successive waves of fixation closely connected with primate lineage history. J Mol Evol 1988;27:194–202
Britten RJ, Baron WF, Stout DB, Davidson EH: Sources and evolution of human Alu repeated sequences. Proc Natl Acad Sci USA 1988;85:4770–4774
Deininger PL, Slagel VK: Recently amplified Alu family members share a common parental Alu sequence. Mol Cell Biol 1988;8:4566–4569
Batzer MA, Deininger PL: A human-specific subfamily of Alu sequences. Genomics 1991;9:481–487
Matera AG, Hellmann U, Schmid CW: A transpositionally and transcriptionally competent Alu subfamily. Mol Cell Biol 1990;10:5424–5432
Rogers JH: The origin and evolution of retroposons. Int Rev Cytol 1985;93:187–279
Mathias SL, Scott AF, Kazazian HH Jr, Boeke JD, Gabriel A: Reverse transcriptase encoded by a human transposable element. Science 1991;254:1808–1810
Wallace MR, Andersen LB, Saulino AM, Gregory PE, Glover TW, Collins FS: A de novo Alu insertion results in neurofibromatosis type 1. Nature 1991;353:864–866
Goossens M, Kan YW: DNA analysis in the diagnosis of hemoglobin disorders; in Antonini E, Rossi-Bernardi L, Chiancone E (eds): Methods in Enzymology. New York, Academic Press, 1981, vol 76, pp 805–817.
Sambrook J, Fritsch EF, Maniatis T: Molecular Cloning. A Laboratory Manual, ed 2. New York, Cold Spring Harbor, 1989, pp 9.31–9.35.
Anson DS, Choo KH, Rees DJG, Giannelli F, Gould K, Huddleston JA, Brownlee GG: Gene structure of human anti-haemophilic factor IX. EMBO J 1984;3:1053–1060
Yoshitake S, Schach BG, Foster DC, Davie EW, Kurachi K: Nucleotide sequence of the gene for human factor IX (antihemophilic factor B). Biochemistry 1985;24:3736–3750
Saiki RK, Gelfand DH, Stoffel S, Scharf SJ, Higuchi R, Horn GT, Mullis KB, Erlich HA: Primer directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science 1988;239:487–491
Gyllensten UB, Erlich HA: Generation of single-stranded DNA by the polymerase chain reaction and its application to direct sequencing of the HLA-DQA locus. Proc Natl Acad Sci USA 1988;85:7652–7656
Mitchell GA, Labuda D, Fontaine G, Saudubray JM, Bonnefont JP, Lyonnet S, Brody LC, Steel G, Obie C, Valle D: Splice-mediated insertion of an Alu sequence inactivates ornithine 2δ-aminotransferase: A role for Alu elements in human mutation. Proc Natl Acad Sci USA 1991;88:815–819
Daniels GR, Deininger PL: Integration site preferences of the Alu family and similar repetitive DNA sequences. Nucleic Acids Res 1985;13:8939–8954
Batzer MA, Kilroy GE, Richard PE, Shaikh TH, Desselle TD, Hoppens CL, Deininger PL: Structure and variability of recently inserted Alu family members. Nucleic Acids Res 1990;18:6793–6798
Matera AG, Hellmann U, Hintz MF, Schmid CW: Recently transposed Alu repeats result from multiple source genes. Nucleic Acids Res 1990;18:6019–6023
Stoppa-Lyonnet D, Carter PE, Meo T, Tosi M: Clusters of intragenic Alu repeats predispose the human C1 inhibitor locus to deleterious rearrangements. Proc Natl Acad Sci USA 1990;87:1551–1555
Muratani K, Hada T, Yamamoto Y, Kaneko T, Shigeto Y, Ohue T, Furuyama J, Higashino K: Inactivation of the Cholinesterase gene by Alu insertion: Possible mechanism for human gene transposition. Proc Natl Acad Sci USA 1991;88:11315–11319
Proudfoot N: Poly (A) signals. Cell 1991;64:671–674
Jagadeeswaran P, Forget BG, Weissman SM: Short, interspersed repetitive DNA elements in eucaryotes: Transposable DNA elements generated by reverse transcription of RNA pol III transcripts? Cell 1981;26:141–142
VanArsdell SW, Dension RA, Bernstein LB, Weiner AM, Manser T, Gesteland RF: Direct repeats flank three small nuclear RNA pseudogenes in the human genome. Cell 1981;26:11–17
Kazazian HH Jr, Wong C, Youssoufian H, Scott AF, Philips DG, Antonarakis SE: Haemophilia A resulting from de novo insertion of L1 sequences represents a novel mechanism for mutation in man. Nature 1988;332:164–166
Woods-Samuels P, Wong C, Mathias SL, Scott AF, Kazazian HH Jr, Antonarakis SE: Characterization of a non deleterious L1 insertion in an intron of the human factor VIII gene and further evidence of open reading frames in functional L1 elements. Genomics 1989;4:290–296
Dombroski BA, Mathias SL, Nanthakumar E, Scott AF, Kazazian HH Jr: Isolation of an active human transposable element. Science 1991;254:1805–1808
Acknowledgements
We thank Josiane Martin for oligonucleotide synthesis.
This work was supported by grants from INSERM, the CNRS and the CNAMTS. BRB was a recipient of a grant from the Philippe Foundation, Paris and New York.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Vidaud, D., Vidaud, M., Bahnak, B.R. et al. Haemophilia B Due to a De Novo Insertion of a Human-Specific Alu Subfamily Member within the Coding Region of the Factor IX Gene. Eur J Hum Genet 1, 30–36 (1993). https://doi.org/10.1159/000472385
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1159/000472385
Key Words
This article is cited by
-
The L1-dependant and Pol III transcribed Alu retrotransposon, from its discovery to innate immunity
Molecular Biology Reports (2021)
-
A systematic analysis of LINE-1 endonuclease-dependent retrotranspositional events causing human genetic disease
Human Genetics (2005)
-
Aberrant caspase-activated DNase (CAD) transcripts in human hepatoma cells
British Journal of Cancer (2003)
-
An Alu-mediated rearrangement as cause of exon skipping in Hunter disease
Human Genetics (2003)
-
Mutations in a human homologue of Drosophila crumbs cause retinitis pigmentosa (RP12)
Nature Genetics (1999)