Abstract
Von Hippel-Lindau (VHL) disease is an autosomal dominantly inherited condition characterized by a predisposition to the development of haemangioblastoma, renal cell carcinoma and phaeochromocytoma. The gene which, when altered, causes the disease was cloned in 1993, and maps within a series of known polymorphic loci in the 3p25–p26 region. To optimize a DNA-based presymptomatic diagnosis, we have selected six highly informative microsatellite loci, closely linked to the VHL gene. Genotyping using a multiplex-PCR approach was performed in 26 affected families including 99 asymptomatic relatives born from an affected parent. Ninety-six subjects were informative with one or more markers, 76 being informative with markers on both sides of the gene. Combination of age-related and DNA-based risk information improved the accuracy of risk assessment for 90 at-risk patients (91%) and allowed attribution of risk with a confidence limit higher than 0.98 in 79 cases (88%).
Similar content being viewed by others
Introduction
Von Hippel-Lindau (VHL) disease is a rare genetic disease with a dominant mode of inheritance, that predisposes affected individuals to a large variety of tumours, the most frequent types being haemangioblastoma of the central nervous system and retina, renal cell carcinoma and phaeochromocytoma [1]. The minimum birth incidence of VHL disease is 1 in 36,000 [2], penetrance is almost complete by 65 years of age, and median actuarial life expectancy is reduced to 49 years, with renal cell carcinoma being the most common cause of death [3]. Because the age of onset is variable and because early tumour detection may reduce morbidity, individuals at risk are advised to enter an early and long-term detection program.
The gene responsible for the VHL disease has been localized on the short arm of chromosome 3 by linkage analysis [4] and a partial cDNA has recently been cloned [5]. The known portion of the gene, divided into three exons, encodes a 284-amino-acid protein. The VHL gene fulfills the characteristics of a tumour suppressor gene: renal cell carcinomas from VHL patients exhibit loss of the wildtype allele inherited from the unaffected parent, and somatic mutations are found in sporadic tumours [6]. Germline alterations have been identified in VHL patients. They can either be large deletions detected by Southern analysis [7] or, more frequently, point mutations found throughout the known coding sequence [8]. However, lacking a functional test, the causal relationship of the detected DNA variation with the VHL disease cannot yet be systematically documented. Furthermore, these combined approaches are unable to reveal DNA variation in approximately 25% of the cases. Thus cosegregation studies should remain a valuable approach to presymptomatic diagnosis in many families.
Several polymorphic markers have been localized close to the VHL gene, and have been ordered with respect to one another and to the VHL gene [9–12]. We selected six highly informative microsatellite polymorphisms that could be genotyped using the PCR technique. Twenty-six families including 99 asymptomatic relatives at 50% prior risk of being gene carriers have been genotyped for these polymorphisms. We report here the result of this analysis. We show that this selection of loci enables a reliable presymptomatic diagnosis of the VHL disease in most families. This approach can be introduced into routine clinical practice.
Material and Methods
Patients
Blood samples were obtained from a total of 268 individuals of 26 apparently unrelated VHL families. The clinical data were collected by SR through patient interviews, hospital notes and autopsy reports of all the referred affected patients and for other family members then identified. All patients satisfied the diagnostic criteria for VHL disease first defined by Melmon and Rosen [13]. In these families, 99 asymptomatic members had an affected parent. Thus, these individuals had a 50% prior risk of being gene carriers.
Genotyping
DNA was extracted from 200 µl of peripheral blood using the Instagene™ Purification Matrix (Bio-Rad). The resulting amount of DNA is sufficient for 100 independent amplifications. Two amplifications were performed by multiplexing (1) amplimers from four loci (D3S1304, D3S1038, D3S1317, D3S1537) and (2) amplimers from two loci (D3S651, D3S656), in final reaction volumes of 20 µl, using the following conditions: initial denaturation at 94°C for 5 min then 35 cycles of denaturation at 94°C (30 s), annealing at 55°C (30 s) and elongation at 72°C (90 s) (PCR9600 from Perkin Elmer). Amplified products (1 µl of each) were electrophoresed in 6% Polyacrylamide sequencing gels (7 M urea and 32% (vol/vol) formamide; acrylamide:bisacrylamide = 29:1), then briefly transferred by capillary blotting on a nylon membrane. Co-hybridization with a 3′-labelled (CA)n oligonucleotide, and an oligonucleotide (5′CTATAAAATGGCTATACCCAG) specific for D3S1537 enables the simultaneous detection of all alleles amplified from the 6 polymorphic loci (table 1).
Risk Analysis
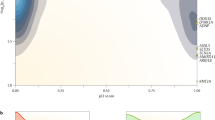
All family members were initially classified as affected, unknown or, when not at risk (spouses), as unaffected. For each at-risk member, the probability of being genetically affected was estimated using the genotyping data and the MLink option of the LINKAGE package. Sex-average recombination fractions between the VHL gene and polymorphic loci that were used for risk estimation were derived from the previously established maps [9–12]: telomere — D3S1537 — 0.06 — D3S1304 — 0.02 — VHL — 0.005 — D3S1317 — 0.01 — D3S1038 — 0.05 — D3S651 — 0.01 — D3S656 — centromere (fig. 1a). When indicated, age-dependent penetrance classes according to Maher et al. [14] were used to derive by Bayesian calculation a final carrier risk estimation combining the age-related and DNA-based risks.
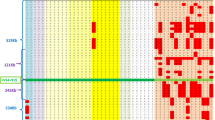
a Genetic map of 6 microsatellite polymorphisms localized in a 15-cM region surrounding the VHL gene. A location map shows the order of selected polymorphic loci with respect to one another and to the VHL gene. Genetic distances were generated using the multipoint linkage analyses in CEPH and VHL families [9–11]. D3S1317 is centromeric to the VHL gene and its distance to the VHL gene is about 50 kb [12]. The corresponding genetic distance was assumed to be 0.5 cM in order to obtain conservative risk estimation, b Genotyping of the VHL region in one affected nuclear family. Filled symbols and question marks correspond to affected and at-risk family members, respectively. Individual III-3 was aged 32 with a completely normal clinical examination (abdominal ultrasonography, ocular fundus examination with angiography, brain magnetic resonance imaging). Individual II-2 had a single renal cyst detected by ultrasonography at age 54. Haplotypes for each of the 4 typed family members have been reconstructed. Experimental data for the 4 flanking loci amplified using a multiplexing procedure are shown. The DNA-based probability of being a gene carrier for the at-risk patient III-3 could be evaluated to 0.0001 using the MLink option of the LINKAGE computer program. Taking into account his age-dependent penetrance (0.67), a combined risk estimation for being a gene carrier was evaluated at 33 × 10−6. The DNA-based probability of being a gene carrier for the at-risk patient II-2 could be evaluated at 0.0008 using MLink. The detection of a renal cyst in this patient prevented the use of the age-dependent risk in the final risk evaluation.
Results
In order to select from among the different polymorphic loci that had been mapped in the vicinity of the VHL gene those loci that would be most useful in the presymptomatic diagnosis of the VHL disease, we applied the following criteria. First, typing of the loci has to be accessible to the PCR technology. Second, they have to be highly informative, i.e. the frequency of heterozygotes has to be 0.5 or higher. Finally, their position with respect to the VHL gene has to be known with precision. Six loci met these criteria. Five of them are associated with a CA repeat of variable length, and the polymorphism of the sixth locus (D3S1537) is due to a variable number of a tetranucleotide repeat. Previous cosegregation analyses have provided a reliable genetic map of the VHL region which included the four most distant loci (D3S1537, D3S1304, D3S651, D3S656) [9, 10]. The positions of the two remaining loci, D3S1317 and D3S1038, relative to the VHL gene have been determined by physical and genetic mapping. They are located on either side of D3S601 and are both centromeric to the currently known portion of the VHL gene [11, 12]. The distance between D3S1317 and the 3′ end of the gene is about 50 kb (fig. 1a). To have a reliable risk evaluation, genetic distances of 0.5 and 1 cM between the VHL gene and the loci D3S1317 and D3S1038, respectively, were assumed.
Genotypic data for these six selected loci were collected on DNAs extracted from all available blood samples of 26 VHL families. In affected individuals, heterozygosity at each locus was not significantly different from that observed in unaffected members. The results were entered into a pedigree datafile in a LINKAGE format. The MLink option of the LINKAGE program was run for each at-risk individual and each typed locus. The genotypic data could not be used in risk estimation for 3 of the 99 at-risk individuals since, in these cases, none of the loci were informative. In contrast, 76 at-risk patients were informative for at least two loci, localized on the centromeric and telomeric sides of the VHL gene. Six of these 76 cases demonstrated a recombination between the closest informative flanking loci. Because it was not possible to decide whether the recombination event occurred centromeric or telomeric to the VHL gene, it was not possible to determine precisely the DNA-based risk of these 6 patients. In the remaining 70 cases, the risk could be calculated with a confidence limit higher than 200:1, solely from the genotyping data (fig. 1b). Twenty cases were informative for a single polymorphic locus or for loci located on a single side of the gene. The lack of information on the other side of the gene led, in many cases, to a confidence limit in the risk estimation that was too low to enable the estimated risk to assist patient management (table 2).
For all asymptomatic individuals born from an affected parent, a combined DNA and age-related risk could be evaluated by a standard Bayesian method using the known age-dependent penetrance classes for the VHL disease [14]. This approach confirmed and increased the precision of the prediction for those at-risk individuals who had a low DNA-based risk (table 2). In contrast, it caused a decrease in the probability of being affected for those asymptomatic individuals with a high DNA-based risk. For this last group, the combined risk of being a gene carrier was not lower than 0.82, indicating that a stringent surveillance protocol should be maintained for every individual belonging to this group (table 2).
Discussion
A multiplex-PCR approach for typing a group of selected polymorphic loci was devised and applied to 99 asymptomatic individuals born from one affected parent. Three individuals were informative for none of the 6 loci and 6 others demonstrated recombination between the two closest informative flanking loci, so that this approach was not useful in risk estimation for a total of 9 individuals (9%). For all other at-risk individuals, a combination of age-related and DNA-based risk information allowed attribution of a risk estimate which effectively assisted decisions concerning optimal patient management. A total of 67 individuals were predicted to be at low risk with a confidence limit higher than 0.98, and although the prediction for the 4 remaining low-risk individuals was less accurate, it remained higher than 0.93 (table 2). Among the asymptomatic individuals predicted as having inherited the genetic defect, 12 cases (63%) were diagnosed with a confidence limit higher than 0.98. For the other 7, this accuracy was not reached because the corresponding individuals were informative on a single locus or on several loci located on a single side of the VHL gene. For these patients, the age-dependent penetrance alone would have suggested that they had a high probability of not having inherited the genetic defect. However, the prediction when DNA-based risk and age-dependent penetrance were combined indicated that these 7 patients had inherited the genetic defect with a confidence limit which was always higher than 0.82. Taken together, these data show that the presently proposed approach improved the accuracy of risk assessment in 90 of 99 cases (91 %). This efficiency compares favourably with previous studies where 70% of cases could be evaluated using a different set of polymorphic loci, none of which was amplifiable using the PCR technique [15].
These results suggest that presymptomatic diagnosis using the six loci selected in this work can be proposed as a systematic approach in the management of families affected with the VHL disease, thus extending the screening program to those relatives who are genetically affected. However, it would be premature to recommend no further follow-up for the relatives found to be at low risk only on the basis of presymptomatic diagnosis combining DNA-based and agerelated risks. More cautious guidelines recommending periodic clinical assessment of all at-risk individuals [1] are indicated, because of the small but real error rate which is inherent to this diagnostic approach [16]. This is also necessary because of the lack of a definitive demonstration that the VHL disease is always associated with an alteration of the VHL gene on chromosome 3p. Although no genetic heterogeneity has yet been observed for the VHL disease, the number and size of the families presently studied remain too small to allow the exlusion of other genetic loci in the aetiology of the VHL disease. However, the proposal can be made that lowrisk individuals enter a reduced screening protocol [1].
Identification of DNA variants in the VHL gene provides an alternate method for presymptomatic diagnosis. When the variation can be convincingly proposed as responsible for the disease, this approach enables a very accurate diagnostic test. It furthermore extends the availability of presymptomatic diagnosis to families with a structure not suitable for linkage-based risk prediction. Nevertheless, genetic diagnosis using flanking DNA markers will continue to be useful in families in which identification of the causal mutation remains dubious or has not been possible. For newly recorded individuals with a well-documented familial history of VHL disease, the diagnostic approach described here takes advantage of effective genotyping methods based on rapid DNA extraction from small quantities of peripheral blood followed by a single-step hybridization on mutiplex-PCR products, thus enabling diagnostic conclusions to be obtained in less than 2 days.
References
Maher ER, Moore AT: Von Hippel-Lindau disease. Br J Opthalmol 1992;76:743–745.
Maher ER, Iselius L, Yates JRW, Littler M, Benjamin C, Harris R, Sampson J, Williams A, Ferguson-Smith MA, Morton N: Von Hippel-Lindau disease: A genetic study. J Med Genet 1991;28:443–447.
Maher ER, Yates JRW, Harries R, Benjamin C, Harris R, Moore AT, Ferguson-Smith MA: Clincial features and natural history of von Hippel-Lindau disease. Q J Med 1990; 283:1151–1163.
Seizinger BR, Rouleau GA, Ozelius LJ, Lane AH, Farmer GE, Lamiell JM, Haines J, Yuen JWM, Collins D, Majoor-Krakauer D, Bonner T, Mathew C, Rubinstein A, Halperin J, McConkie-Rosell A, Green JS, Trofatter JA, Ponder BA, Eierman L, Bowmer MI, Schimke R, Oostra B, Aronin N, Smith DI, Drabkm H, Wasiri MH, Hobbs WJ, Martuza RL, Conneally PM, Hsia YE, Gusella JF: Von Hippel-Lindau disease maps to the region of chromosome 3 associated with renal cell carcinoma. Nature 1988;332:268–269.
Latif F, Tory K, Gnarra J, Yao M, Dun FM, Orcutt ML, Stackhouse T, Kuzmin I, Modi W, Geil L, Schmidt L, Zhou F, Li H, Wei MH, Chen F, Glenn G, Choyke P, Walther MM, Weng Y, Duan DSR, Dean M, Glavak D, Richards FM, Crossey PA, Ferguson-Smith MA, Le Paslier D, Chumakov I, Cohen D, Chinault AC, Maher ER, Lineahn WM, Zbar B, Lerman MI: Identification of the von Hippel-Lindau disease tumor suppressor gene. Science 1993;260: 1317–1320.
Gnarra JR, Tory K, Weng Y, Schmidt L, Wei MH, Li H, Latif F, Liu S, Chen F, Duh FM, Lubensky I, Duan DR, Florence C, Pozzatti R, Walther MM, Bander NH, Grossman HB, Brauch H, Pomer S, Brooks JD, Isaacs WB, Lerman MI, Zbar B, Linehan WM: Mutations of the VHL tumour suppressor gene in renal cell carcinoma. Nat Genet 1994;7:85–89.
Richards FM, Crossey PA, Phipps ME, Foster K, Latif F, Evans G, Sampson J, Larman MI, Zbar B, Affara NA, Ferguson-Smith MA, Maher ER: Detailed mapping of germline deletions of the von Hippel-Lindau disease tumour suppressor gene. Hum Mol Genet 1994;3:595–598.
Crossey PA, Richards FM, Foster K, Green JS, Prowse A, Latif F, Lerman MI, Zbar B, Affara NA, Ferguson-Smith MA, Maher ER: Identification of intragenic mutations in the von Hippel-Lindau disease tumour suppressor gene and correlation with disease phenotype. Hum Mol Genet 1994;3:1303–1308.
Richards FM, Maher ER, Latif F, Phipps ME, Tory K, Lush M, Crossey PA, Oostra B, Gustavson KH, Green J, Turner G, Yates JRW, Linehan WM, Affara NA, Lerman M, Zbar B, Ferguson-Smith MA: Detailed genetic mapping of the von Hippel-Lindau disease tumour suppressor gene. J Med Genet 1993;30: 104–107.
Pericak-Vance MA, Nunes KJ, Whisenant E, Loeb DB, Small KW, Stajich JM, Rimmler JB, Yamaoka LH, Smith DI, Drabkin HA, Vance JM: Genetic mapping of dinucleotide repeat polymorphisms and von Hippel-Lindau disease on chromosome 3p25-p26. J Med Genet 1993;30: 487–491.
Crossey PA, Maher ER, Jones MH, Richards FM, Latif F, Phipps ME, Lush M, Foster K, Tory K, Green JS, Ostra B, Yates JRW, Linehan WM, Affara NA, Lerman M, Zbar B, Nakamura Y, Ferguson-Smith MA: Genetic linkage between von Hippel-Lindau disease and three microsatellite polymorphisms refines the localisation of the VHL locus. Hum Mol Genet 1993,2:279–282.
Kuzmin I, Stackhouse T, Latif F, Duh FM, Geil L, Gnarra J, Yao M, Orcutt ML, Li H, Tory K, Le Paslier D, Chumakov I, Cohen D, Chinault AC, Linehan WM, Lerman MI, Zbar B: One-megabase yeast artificial chromosome and 400-kilobase cosmid-phage contigs containing the von Hippel-Lindau suppressor and Ca2+-transporting adenosine triphosphatase isoform 2 genes. Cancer Res 1994;54:2486–2491.
Melmon KL, Rosen SW: Lindau’s disease. Am J Med 1964;36:595–617.
Maher ER, Bentley E, Payne SJ, Latif F, Richards FM, Chiano M, Hosoe S, Yates JRW, Linehan M, Barton DE, Glenn G, Affara NA, Lerman M, Zbar B, Ferguson-Smith MA: Presymptomatic diagnosis of von Hippel-Lindau disease with flanking DNA markers. J Med Genet 1992;29:902–905.
Seizinger BR, Smith DI. Filling-Katz MR, Neumann H, Green JS, Choyke PL, Anderson KM, Freoman RN, Klauck SM, Whaley J, Decker HJH, Hsia YE, Collins D, Halperin J, Lamiell JM, Oostra B, Wasiri MH, Gorin MB, Scherer G, Drabkin HA, Aronin N, Schinzel A, Martuza RL, Gusella JF, Haines JL: Genetic flanking markers refine diagnostic criteria and provide insights into the genetics of von Hippel-Lindau disease. Science 1991; 88:2864–2868.
Glenn GM, Linehan M, Hosoe S, Latif F, Yao M, Choyke P, Gorin MB, Chew E, Oldfield E, Manolatos C, Orcutt ML, Walther MCM, Weiss GH, Tory K, Jensson O, Lerman MI, Zbar B: Screening for von Hippel-Lindau disease by DNA-polymorphism analysis. JAMA 1992;267:1226–1231.
Acknowledgements
We wish to thank the VHL patients and their families for their cooperation, and clinicians and surgeons for their contribution to the registration of families. This work was supported by the Association Française contre les Myopathies and the Comité du Cher de la Ligue Nationale contre le Cancer
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Olschwang, S., Boisson, C., Richard, S. et al. DNA-Based Presymptomatic Diagnosis for the von Hippel-Lindau Disease by Linkage Analysis. Eur J Hum Genet 3, 108–115 (1995). https://doi.org/10.1159/000472284
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1159/000472284