Abstract
We ascertained 9 multigeneration Belgian families with pure dominant spastic paraplegia (SPG) for clinical and genetic studies. Linkage was examined using simple tandem repeat (STR) markers located near the 5 loci for familial SPG on chromosomes Xq28 (SPG1), Xq21.3–q22 (SPG2), 2p21–p24 (SPG4), 14q12–q23 (SPG3) and 15q11.1 (SPG6). Positive linkage results were obtained only for markers at the SPG4 locus mapping the SPG4 gene between D2S400 and D2S367, a region of 4 cM. In order to facilitate the positional cloning of the SPG4 gene, we constructed a contiguous YAC map covering the SPG4 candidate region. Our physical mapping data indicate that the SPG4 gene resides within maximal 5 Mb.
Similar content being viewed by others
Introduction
Familial pure dominant spastic paraplegia (SPG) is a clinically and genetically heterogeneous group of neurodegenerative disorders, mainly characterised by spasticity of the lower limbs. Gait deteriorates progressively, with some patients becoming wheelchair bound at an older age. Spasticity is always disproportionate to the degree of weakness, and sensory signs are usually absent on routine neurological examination. Clinically, patients can be diagnosed as pure SPG patients when only symptoms of spasticity are present or as complex SPG patients if additional features are also present. Pure SPG is inherited either as an autosomal dominant or an autosomal recessive trait and rarely as an X-linked trait. In pure autosomal dominant SPG, the age at onset of the disease can vary over a wide range. A classification in early-onset SPG and late-onset SPG with disease onset respectively before and after age 35 has been proposed [1]. However, there is a certain degree of overlap between early-onset and late-onset SPG.
Pathological data on these disorders are scarce. Abnormalities are usually confined to the spinal cord showing degeneration of the crossed pyramidal tracts starting in the cervical region and increasing caudally. Some degenerative changes are frequently present in the posterior columns and the spinocerebellar tracts at the cervical level [2, 3]. The pathogenesis and the underlying molecular biological defect remain unknown. Therefore, the diagnosis of pure SPG is entriely based on detecting unequivocal signs of a pyramidal syndrome on clinical examination. It is difficult to exclude the disease in young asymptomatic at-risk individuals especially in late-onset SPG families. Genetic counselling is further hampered since age at onset and severity of symptoms can vary considerably from patient to patient even within the same family. Genetic linkage studies have demonstrated that pure dominant SPG is genetically heterogeneous with 2 loci on chromosome X, SPG1 at Xq28 and SPG2 at Xq21.3–q22 [4, 5], and 3 loci on chromosomes 2p21–p24 (SPG4), 14q12–q23 (SPG3) and 15q11.1 (SPG6) [6–10].
Materials and Methods
Clinical Studies
On the basis of the clinical examination, neurological signs were scored for gait abnormalities, hyperreflexia, presence of Babinski signs, weakness, and sensation abnormalities (touch, vibration, joint position and pin-prick). Individuals were considered to be definitely affected when the following criteria were met: unequivocally stiff gait or hyperreflexia with extensor plantar reflexes. Obligate gene carriers with an affected parent and an affected offspring were also scored as affected. Some individuals, especially in the younger generations, were scored as probably affected based on hyperreflexia with polykinetic reflexes or clonus but without Babinski sign or unequivocal gait abnormality.
Genetic Linkage Studies
After informed consent was given, 30 ml of blood was obtained for DNA isolation. DNA was isolated from whole blood samples, and the polymerase chain reaction (PCR) amplification of STR DNA markers was carried out using standard procedures. For markers D14S255, D14S269 and D14S281, the polymorphic alleles were separated on a 6% Polyacrylamide sequencing gel and visualised by autoradiography. For all other STR markers we used a fluorescent detection system based on primer end labelling with one of the fluorophores JOE, FAM or TAMRA using the protocol of Applied Biosystems (ABI) Inc.; the polymorphic alleles were separated on a 6% Polyacrylamide sequencing gel in the automated sequencer ABI 373A. The data were collected and analysed using the ABI GENESCAN 672 software.
Two-point lod scores (Z) were calculated including both affected and unaffected individuals using the computer program MLINK of the FASTLINK version 2.3P package [11–14], assuming an autosomal or X-linked dominant disease gene with a frequency of 1/10,000 and equal female and male recombination rates. Phenocopy and mutation rates were set to zero. The unaffected individuals were assigned a disease penetrance class, according to their age. Based on the cumulative risk curve [15], 8 age-dependent penetrance classes were used with a maximal disease penetrance of 90% at the age of 40 years. For each simple tandem repeat (STR) marker, the allele frequencies were calculated from the unrelated individuals in the pedigrees using the computer program PRIMITIV (kindly provided by L. Sandkuijl).
YAC Contig Mapping
By searching 4 different databases: Genome Database: http://gdbwww.gdb.org, The Whitehead Institute for Genome Research: http://www-genome.wi.mit.edu/, Baylor College of Medicine Genome center: http://kiwi.imgen.bcm.tmc.edu:8088/home.html, and the Généthon server: http://www.genethon.fr/genethon_en.html, we selected a total of 27 CEPH mega YAC. YAC clones were obtained from the YAC screening center Leiden, The Netherlands (yscl@ruly46.medfac.leidenuniv.nl). The YAC clones were plated on selective culture plates and from each plate one colony was grown in selective culture media for YAC DNA extraction using standard procedures.
Published primers were used to amplify sequence-tagged sites (STSs) and STR markers from the region using standard PCR conditions. The PCR products were separated on a 1.5% agarose gel and visualised on a UV transilluminator after ethidiumbromide staining.
Results
Clinical Features
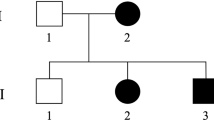
We have ascertained 9 families with pure dominant SPG for clinical and genetic studies (fig. 1). The clinical profile in each kinship is characterised by progressive gait abnormalities, hyperreflexia and Babinski signs. Some severely affected patients had a complete paralysis of the legs. Sensory disturbances were never prominent. The neurological signs were always most marked in the legs and all patients retained normal function of the arms. In many patients the neurological deficits and symptoms came on so insidiously that it was difficult to identify an exact age at onset. In the younger generation, several individuals were unaware of symptoms, although they had an aberrant walking pattern that was readily apparent to their relatives. A striking variability in age at onset and severity was observed in pedigrees SL-1, SL-2 and SL-11 (fig. 1). These families usually contained both early- and late-onset cases with regard to the proposed cut-off age of 35 years. However, we found patients with an early-onset age in all families and therefore all our families can be classified as pure early-onset SPG. The clinical features of SL-10 have been reported previously [16].
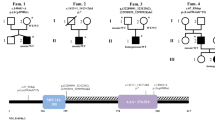
Pedigrees of the SPG families. Symbols: squares = males; circles = females; slashed symbols = deceased individuals; filled symbols = definitely affected patients; half-filled symbols = probably affected individuals. For each affected individual examined, we have indicated at the left of the symbol respectively age at onset and age at examination in years separated by a ‘/’. Also, an indication of the disease severity is given on a scale of 5 with 1 = hyperreflexia without unequivocal gait abnormalities; 2 = slight stiffness when walking; 3 = striking stiffness; 4 = needs walking aid; 5 = wheelchair bound. Family members for which DNA samples are available are indicated with an asterisk above the symbol. The alleles and haplotypes for the STR markers D2S400, D2S352, and D2S367 are given below the symbols for family SL-1, SL-4 and SL-11. The disease haplotype is boxed. When phase was unknown or reconstructed, the alleles were placed between brackets.
Linkage Results
All pedigrees are consistent with an autosomal dominant mode of inheritance (fig. 1). However, in the absence of male-to-male transmission X-linked dominant transmission cannot be ruled out in families SL-6, SL-8 and SL-9. Therefore, we examined STR markers near the SPG1 and SPG2 loci [4, 5]. Two-point lod scores summarized over the 3 families were negative, making linkage to the SPG1 and SPG2 loci unlikely (data available upon request). Next we analysed STR markers located near the SPG3, SPG4 and SPG6 loci. The two-point lod scores (Z) summarised over the 9 families are conclusively negative (Z < −2) for D14S255 (Z = −7.65 at θ = 0.00, exclusion limit: 4.61 cM) and for marker D15S156 (Z = −17.54 at θ = 0.00, exclusion limit: 15.55 cM) excluding linkage to SPG3 and SPG6 loci (data available upon request). At the SPG4 locus, positive lod scores are obtained with all 3 markers D2S400 (Zmax = 13.4, θmax = 0.10), D2S352 (Zmax = 5.69, θmax = 0.05) and D2S367 (Zmax = 8.50, θmax = 0.00), with the majority of the families showing positive lod scores (table 1). One family, SL-11, showed conclusive linkage (Z > 3) in the absence of recombinants with markers D2S367 and D2S352. With D2S400, located telomeric of D2S352, an obligate recombinant is observed in patient IV-7 (fig. 1). In the probably affected individual IV-12, a recombination occurred between markers D2S352 and D2S367. The latter recombinant does not show in the two-point lod scores (table 1), since probably affected individuals were assigned an unknown disease status in the linkage analysis. The SPG4 candidate region defined by the informative recombinants with D2S400 and D2S367 in family SL-11 is in agreement with previous findings [6].
YAC Contig Mapping
DNA of 27 YACs selected for the SPG4 candidate region was examined by PCR amplification for the presence of the STRs D2S400, D2S352 and D2S367 as well as for a number of STSs previously located in the candidate region [17]. Fourteen YACs were positive for one or more STSs/STRs, while 7 YACs contained multiple STSs/STRs (fig. 2). Together the YACs spanned the complete candidate region between D2S400 and D2S367. The order of the STRs/STS is in full agreement with the physical map of Hudson et al. [17], the latter however did not contain D2S367 flanking the SPG4 locus. A maximal physical size of 5 Mb was calculated for the SPG4 candidate region based on the 4 YACs: 895_c_12, 802_a_5, 931_e_1 and 937_d_3, taking into account the insert sizes of these YACs [18].
Discussion
Linkage analyses in 9 Belgian SPG families provided evidence that one family, SL-11, is linked to the SPG4 locus. The majority of the remaining families showed positive lod scores with 2 families, SL-1 and SL-4, exceeding lod scores of 1.5, suggesting that the SPG4 locus is the most common locus for pure autosomal dominant SPG in our families. Analysis of the lod scores obtained with the linked marker D2S352 in the 9 families, using the computer program HOMOG 3.3, provided weak evidence for heterogeneity with 72% of the families linked to SPG4 (α = 0.72, θ = 0.00, χ2 = 2.36). Indeed negative lod scores were obtained in families SL-3 and SL-8 (table 1), making linkage to SPG4 in these families unlikely.
In family SL-11, there was a wide onset age range of 1–60 years (fig. 1). Patients with extremely young onset ages were also more severely affected, with some having a complete paralysis of the legs at older age. Further, when descendants of patient III-21 were examined it was clear that onset age decreased and disease severity increased, suggestive of disease anticipation. The grandson V-2 was severely affected at age 1 year, while both his father IV-11 and grandmother III-21 considered themselves unaffected, although clinical examination revealed minor signs of SPG (fig. 1). We do not have sufficient data to formally conclude that disease anticipation is involved in the disease pathology in family SL-11, however, disease anticipation was also described in other SPG4 families [6, 19]. In families SL-1 and SL-4, that showed suggestive linkage to SPG4, anticipation could be suspected in family SL-1, while in family SL-4 the clinical expression was less variable over successive generations. The presence of anticipation in some SPG4 pedigrees provides a potentially clue towards the underlying molecular genetic mechanism. Several inherited neurodegenerative disorders which show anticipation are now known to be caused by the expansion of an unstable trinucleotide repeat [20]. In some of these disorders, the phenotype in the affected child is influenced by the gender of the transmitting parent, a phenomenon known as genomic imprinting [21]. Interestingly, in families SL-11 and SL-1, all patients with onset ages less than 4 years had inherited the disease from their father, suggesting that there could be a paternal sex effect. However, more SPG4 families will be needed to confirm this observation.
The linkage results in family SL-11 confirm previous data localizing the SPG4 gene between the flanking markers D2S400 and D2S367 [6], separated by a genetic distance of 4 cM [22]. We constructed a YAC contig map of the entire SPG4 candidate region allowing a maximal size estimate of 5 Mb. The YAC contig map will be a useful framework in cloning experiments directed at isolating the SPG4 gene.
References
Harding AE: Classification of the hereditary ataxias and paraplegias. Lancet 1983;i: 1151–1155.
Behan WMH, Maia M: Strümpell’s familial spastic paraplegia: Genetics and neuropathology. J Neurol Neurosurg Psychiatry 1974;37:8–20.
Bruyn RPM: The neuropathology of hereditary spastic paraparesis. Clin Neurol Neurosurg 1992;94(suppl):S16–S18.
Kenwick S, Ionasescu V, Searby C, King A, Dubowitz M, Davies K: Linkage studies of X-linked recessive spastic paraplegia using DNA probes. Hum Genet 1989;73:264–266.
Saugier-Veber P, Munnich A, Bonneau D, Rozet JM, Le Merrer M, Gil R, Boespfug-Tanguy O: X-linked spastic paraplegia and Pelizaeus-Merzbacher disease are allelic disorders at the proteolipid protein locus. Nat Genet 1994;6: 257–262.
Hazan J, Fontaine B, Bruyn RPM, Lamy C, van Deutekom JCT, Rime CS, Dürr A, Melki J, Lyon-Caen O, Agid Y, Munnich A, Padberg GW, De Recondo J, Frants RR, Brice A, Weissenbach J: Linkage of a new locus for autosomal dominant familial spastic paraplegia to chromosome 2p. Hum Mol Genet 1994;3: 1569–1573.
Hentati A, Pericak-Vance MA, Lennon F, Wasserman B, Hentati F, Junega T, Angrist MH, Hung WY, Boustany RM, Bohlega S, Iqbal Z, Huether CH, Ben Hamida M, Siddique T: Linkage of a locus for autosomal dominant familial spastic paraplegia to chromosome 2q markers. Hum Mol Genet 1994;3:1867–1871.
Hazan J, Lamy C, Melki J, Munnich A, De Recondo J, Weissenbach J: Autosomal dominant familial spastic paraplegia is genetically heterogeneous and one locus maps to chromosome 14q. Nat Genet 1993;5:163–167.
Gispert S, Santos N, Damen R, Voit T, Schulz J, Klockgether T, Orozco G, Kreuz F, Weissenbach J, Auburger G: Autosomal dominant familial spastic paraplegia: Reduction of the FSP I candidate region on chromosome 14q to 7cM and locus heterogeneity. Am J Hum Genet 1995;56:183–187.
Fink JK, Wu CB, Jones SM, Sharp GB, Lange BM, Lesicki A, Reinglass T, Varvil T, Otterud B, Leppert M: Autosomal dominant familial spastic paraplegia: Tight linkage to chromosome 15q. Am J Hum Genet 1995;56:188–192.
Cottingham RW, Idury RM, Schäffer AA: Faster sequential genetic linkage computations. Am J Hum Genet 1993;53:252–263.
Lathrop GM, Lalouel JM: Easy calculations of lod scores and genetic risk on small computers. Am J Hum Genet 1984;36:460–465.
Lathrop GM, Lalouel JM, Julier C, Ott J: Multilocus linkage analysis in humans: Detection of linkage and estimation of recombination. Am J Hum Genet 1985;37:482–498.
Schäffer AA, Gupta SK, Shriram K, Cottingham C: Avoiding recomputation in linkage analysis. Hum Hered 1994;44:225–237.
Ott J: Analysis of human genetic linkage. Baltimore, The Johns Hopkins University Press, 1991.
Marcheau MMB, De Keyser G: Familie-onderzoek bij de ziekte van Strümpell-Lorain. Tijdschr Geneesk 1979;6:399–713.
Hudson TJ, Stein LD, Gerety SS, Ma J, Castle AB, Silva J, Slonim DK, Baptista R, Kruglyak L, Xu SH, Hu X, Colbert AME, Rosenberg C, Reeve-Daly MP, Rozen S, Hui L, Wu X, Vestergaard C, Wilson KM, Bae JS, Maitra SH, Ganiatsas S, Evans CA, DeAngelis MM, Ingalls KA, Nahf RW, Horton LT Jr, Anderson MO, Collymore AJ, Ye W, Kouyoumijan V, Zemsteva IS, Tam J, Devine R, Courtney DF, Renaud MT, Nguyen H, O’Connor TJ, Fizames C, Fauré S, Gyapay G, Dib C, Morissette J, Orlin JB, Birren BW, Goodman N, Weissenbach J, Hawkins TL, Foote S, Page DC, Lander ES: An STS-based map of the human genome. Science 1995;270:1945–1954.
Cohen D, Chumakov I, Weissenbach J: A first physical map of the human genome. Nature 1993;366:698–701.
Bruyn RPM, van Deutekom J, Frants RR, Padberg G: Hereditary spastic paraparesis. Clinical and genetic data from a large Dutch family. Clin Neurol Neurosurg 1993;95:125–129.
Willems PJ: Dynamic mutations hit double figures. Nat Genet 1994;8:213–215.
Chatkupt S, Antonowicz M, Johnson WG: Parents do matter: Genomic imprinting and parental sex effects in neurological disorders. J Neurol Sci 1995;130:1–10.
Gyapay G, Morissette J, Vignal A, Dib C, Fizames C, Millasseau P, Marc S, Bernardi G, Lathrop M, Weissenbach J: The 1993–1994 Généthon human genetic linkage map. Nat Genet 1994;7:246–339.
Acknowledgements
The authors are grateful to the patients and their relatives for their cooperation in the research project and to Dr. Marchau for referring family SL-10. This work was in part funded by a concerted action of the Ministry of Education and the National Fund of Scientific Research (NFSR). L.K. holds a grant from the Institute of Science and Technology (IWT).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
De Jonghe, P., Krols, L., Michalik, A. et al. Pure Familial Spastic Paraplegia: Clinical and Genetic Analysis of Nine Belgian Pedigrees. Eur J Hum Genet 4, 260–266 (1996). https://doi.org/10.1159/000472212
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1159/000472212