Article
|
Open Access
Featured
-
-
Article
| Open Access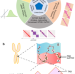 A comprehensive benchmarking with interpretation and operational guidance for the hierarchy of topologically associating domains
A comprehensive benchmarking with interpretation and operational guidance for the hierarchy of topologically associating domains
TAD hierarchy demonstrates cell-to-cell variability, leading to the development of numerous callers. Here, authors present a comprehensive benchmark of TAD hierarchy callers and introduce the ‘air conditioner’ model to illustrate TAD hierarchy’s role in transcription.
- Jingxuan Xu
- , Xiang Xu
- & Hebing Chen
-
Article
| Open Access Benchmarking of methods for DNA methylome deconvolution
Benchmarking of methods for DNA methylome deconvolution
Determining the different cell types that contribute to a mixture of DNA is key for research and diagnostic applications. Here, authors comprehensively benchmark DNA methylation-based deconvolution methods, evaluating their performance and robustness to technical bias.
- Kobe De Ridder
- , Huiwen Che
- & Bernard Thienpont
-
Article
| Open Access LoCoHD: a metric for comparing local environments of proteins
LoCoHD: a metric for comparing local environments of proteins
The techniques available for comparing protein structures do not focus directly on the chemical nature of residue environments. Here, authors describe a computational method that can capture both the spatial and chemical dissimilarities of residue surroundings.
- Zsolt Fazekas
- , Dóra K. Menyhárd
- & András Perczel
-
Article
| Open Access BERNN: Enhancing classification of Liquid Chromatography Mass Spectrometry data with batch effect removal neural networks
BERNN: Enhancing classification of Liquid Chromatography Mass Spectrometry data with batch effect removal neural networks
Liquid Chromatography Mass Spectrometry (LC-MS) is a powerful method for profiling biological samples. Here, the authors have developed a suit of Batch Effect Removal Neural Networks (BERNN) to remove batch effects in large LC-MS experiments to maximize sample classification between conditions.
- Simon J. Pelletier
- , Mickaël Leclercq
- & Arnaud Droit
-
Article
| Open Access MetaboAnalystR 4.0: a unified LC-MS workflow for global metabolomics
MetaboAnalystR 4.0: a unified LC-MS workflow for global metabolomics
Several bottlenecks exist in metabolomics data analysis. Here, the authors present MetaboAnalystR 4.0 as a unified workflow for LC-MS untargeted metabolomics. It highlights significant improvements in LC-MS2 spectral processing and functional analysis, providing an end-to-end computational pipeline.
- Zhiqiang Pang
- , Lei Xu
- & Jianguo Xia
-
Article
| Open Access scLENS: data-driven signal detection for unbiased scRNA-seq data analysis
scLENS: data-driven signal detection for unbiased scRNA-seq data analysis
Single-cell RNA sequencing data analysis is limited by noise and high dimensionality. Here, authors present scLENS, a tool that automates accurate signal detection without manual input, particularly in complex datasets.
- Hyun Kim
- , Won Chang
- & Jae Kyoung Kim
-
Article
| Open Access GENESIS CGDYN: large-scale coarse-grained MD simulation with dynamic load balancing for heterogeneous biomolecular systems
GENESIS CGDYN: large-scale coarse-grained MD simulation with dynamic load balancing for heterogeneous biomolecular systems
Here, the authors report the development of heterogeneous domain decomposition with load balancing for large biological molecular dynamics simulations using residue-level coarse-grained models.
- Jaewoon Jung
- , Cheng Tan
- & Yuji Sugita
-
Article
| Open Access Data-driven recombination detection in viral genomes
Data-driven recombination detection in viral genomes
Here, the authors present RecombinHunt, a computational method based on big data analysis, that enhances community-based detection of recombinant viral lineages.
- Tommaso Alfonsi
- , Anna Bernasconi
- & Stefano Ceri
-
Article
| Open Access Accurately clustering biological sequences in linear time by relatedness sorting
Accurately clustering biological sequences in linear time by relatedness sorting
Accurately clustering biological sequences is an increasingly important task but is challenging for large datasets. This study introduces a new approach called ‘relatedness sorting’ to accurately cluster sequences with linear-time scalability.
- Erik Wright
-
Article
| Open Access Pianno: a probabilistic framework automating semantic annotation for spatial transcriptomics
Pianno: a probabilistic framework automating semantic annotation for spatial transcriptomics
Recognising spatial spots’ biological identity in spatial transcriptomics remains a challenge. Here, authors introduce Pianno, a tool that helps annotate the biological structures or cell-type constructions across diverse tissues, offering new perspectives on understanding spatial transcriptomics.
- Yuqiu Zhou
- , Wei He
- & Ying Zhu
-
Article
| Open Access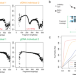 FinaleMe: Predicting DNA methylation by the fragmentation patterns of plasma cell-free DNA
FinaleMe: Predicting DNA methylation by the fragmentation patterns of plasma cell-free DNA
DNA methylation from cell-free DNA (cfDNA) can be profiled using whole genome bisulfite sequencing (WGBS). Here, the authors develop a computational method, FinaleMe, that predicts DNA methylation and tissues of-origin in cfDNA and validate its performance using paired deep and shallow-coverage whole-genome sequencing (WGS) and WGBS data.
- Yaping Liu
- , Sarah C. Reed
- & Manolis Kellis
-
Article
| Open Access PLMSearch: Protein language model powers accurate and fast sequence search for remote homology
PLMSearch: Protein language model powers accurate and fast sequence search for remote homology
Homologous protein search is one of the most commonly used methods for protein analysis. Here, authors propose PLMSearch, a search method that takes only sequences as input and can search millions of protein pairs in seconds while maintaining sensitivity comparable to SOTA structure search methods.
- Wei Liu
- , Ziye Wang
- & Shanfeng Zhu
-
Article
| Open Access Mapping cell-to-tissue graphs across human placenta histology whole slide images using deep learning with HAPPY
Mapping cell-to-tissue graphs across human placenta histology whole slide images using deep learning with HAPPY
Placenta histopathology for maternal and newborn health is highly specialised and time consuming. Here, authors present a deep learning pipeline for quantifying cells and tissues in placenta whole slide images, revealing biological and clinical insights.
- Claudia Vanea
- , Jelisaveta Džigurski
- & Christoffer Nellåker
-
Article
| Open Access Tradeoffs in alignment and assembly-based methods for structural variant detection with long-read sequencing data
Tradeoffs in alignment and assembly-based methods for structural variant detection with long-read sequencing data
Long-read sequencing can greatly improve detection of genomic structural variants (SVs), and numerous methods have been developed to identify SVs using long-read data. Here the authors compare the performance of these methods and provide guidelines to aid users in selecting the most suitable tools for various scenarios.
- Yichen Henry Liu
- , Can Luo
- & Xin Maizie Zhou
-
Article
| Open Access BASALT refines binning from metagenomic data and increases resolution of genome-resolved metagenomic analysis
BASALT refines binning from metagenomic data and increases resolution of genome-resolved metagenomic analysis
Binning is an essential step in genome-resolved metagenomic analysis in which assembled contigs originating from the same source population are clustered. However it is challenging, especially for low abundance microbial species. Here the authors introduce a toolkit that integrates multiple prominent binning tools and AI for efficient and high-resolution recovery of non-redundant bins from short- and long-read metagenomic sequencing datasets.
- Zhiguang Qiu
- , Li Yuan
- & Ke Yu
-
Article
| Open Access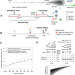 Evolving copy number gains promote tumor expansion and bolster mutational diversification
Evolving copy number gains promote tumor expansion and bolster mutational diversification
Understanding the timing and fitness of somatic copy number alterations (SCNAs) in cancer would shed light on cancer progression and evolution. Here, the authors develop Butte, a computational framework to estimate the timing of clonal SCNAs that encompass multiple gains, and apply it on whole-genome sequencing data from 184 samples.
- Zicheng Wang
- , Yunong Xia
- & Ruping Sun
-
Article
| Open Access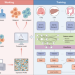 Domain generalization enables general cancer cell annotation in single-cell and spatial transcriptomics
Domain generalization enables general cancer cell annotation in single-cell and spatial transcriptomics
Efficient and accurate annotation of malignant cells is crucial for single-cell and spatial transcriptomics in cancer. Here, the authors develop Cancer-Finder, a deep-learning algorithm that can identify malignant cells in cancer single-cell and spatial transcriptomics data with speed and precision.
- Zhixing Zhong
- , Junchen Hou
- & Jia Song
-
Article
| Open Access Automatic data-driven design and 3D printing of custom ocular prostheses
Automatic data-driven design and 3D printing of custom ocular prostheses
Manual processes to produce ocular prostheses are time-consuming and yield varying quality. Here, authors present an automatic digital end-to-end process for custom ocular prostheses. It creates shape and appearance from image data of an OCT device and produces them using a full-colour 3D printer.
- Johann Reinhard
- , Philipp Urban
- & Mandeep S. Sagoo
-
Article
| Open Access Statistical method scDEED for detecting dubious 2D single-cell embeddings and optimizing t-SNE and UMAP hyperparameters
Statistical method scDEED for detecting dubious 2D single-cell embeddings and optimizing t-SNE and UMAP hyperparameters
2D visualisation of single-cell data is highly impacted by the hyperparameter setting of the 2D embedding method, such as t-SNE and UMAP. Here, authors develop a statistical method scDEED to detect dubious cell embeddings and optimise the hyperparameter setting for trustworthy visualisation.
- Lucy Xia
- , Christy Lee
- & Jingyi Jessica Li
-
Article
| Open Access Machine learning-based extrachromosomal DNA identification in large-scale cohorts reveals its clinical implications in cancer
Machine learning-based extrachromosomal DNA identification in large-scale cohorts reveals its clinical implications in cancer
‘Extrachromosomal DNA has been previously linked to tumour progression and heterogeneity, but its potential as a cancer biomarker has not been fully explored. Here, the authors develop a computational framework to refine genomic subtypes and predict response to immunotherapy in gastrointestinal cancer.
- Shixiang Wang
- , Chen-Yi Wu
- & Qi Zhao
-
Article
| Open Access High resolution spatial profiling of kidney injury and repair using RNA hybridization-based in situ sequencing
High resolution spatial profiling of kidney injury and repair using RNA hybridization-based in situ sequencing
Advancements in spatial transcriptomics technologies have enabled the analysis of gene expression at cellular resolution in situ. The authors applied direct RNA hybridization-based in situ sequencing (dRNA HybISS) and developed a computational tool, CellScopes, to study gene expression in mouse kidneys, identifying cellular changes and interactions during injury and repair.
- Haojia Wu
- , Eryn E. Dixon
- & Benjamin D. Humphreys
-
Article
| Open Access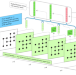 Sequential stacking link prediction algorithms for temporal networks
Sequential stacking link prediction algorithms for temporal networks
Link prediction in temporal networks is relevant for many real-world systems, however, current approaches are usually characterized by high computational costs. The authors propose a temporal link prediction framework based on the sequential stacking of static network features, for improved computational speed, appropriate for temporal networks with completely unobserved or partially observed target layers.
- Xie He
- , Amir Ghasemian
- & Peter J. Mucha
-
Article
| Open Access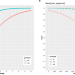 Utility of long-read sequencing for All of Us
Utility of long-read sequencing for All of Us
Using All of Us pilot data, the authors compared short- and long-read performance across medically relevant genes and showcased the utility of long reads to improve variant detection and phasing in easy and hard to resolve medically relevant genes.
- M. Mahmoud
- , Y. Huang
- & F. J. Sedlazeck
-
Article
| Open Access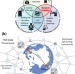 A dynamic knowledge graph approach to distributed self-driving laboratories
A dynamic knowledge graph approach to distributed self-driving laboratories
Global challenges demand global solutions. Here, the authors show a distributed self-driving lab architecture in The World Avatar, linking robots in Cambridge and Singapore for asynchronous multi-objective reaction optimisation.
- Jiaru Bai
- , Sebastian Mosbach
- & Markus Kraft
-
Article
| Open Access Clinical application of tumour-in-normal contamination assessment from whole genome sequencing
Clinical application of tumour-in-normal contamination assessment from whole genome sequencing
Assessing tumour contamination in normal samples is critical for accurate variant calling in cancer samples. Here, the authors develop TINC, a computational method to determine the level of tumour in normal contamination, and demonstrate its application in the Genomics England 100,000 Genomes Project dataset.
- Jonathan Mitchell
- , Salvatore Milite
- & Giulio Caravagna
-
Article
| Open Access PROST: quantitative identification of spatially variable genes and domain detection in spatial transcriptomics
PROST: quantitative identification of spatially variable genes and domain detection in spatial transcriptomics
Understanding biological mechanisms requires a thorough exploration of spatiotemporal transcriptional patterns in complex tissues. Here, authors present PROST to quantify spatial gene expression patterns and detect spatial domains using spatial transcriptomics data of varying resolutions.
- Yuchen Liang
- , Guowei Shi
- & Zhonghui Tang
-
Article
| Open Access Mesoscale simulation of biomembranes with FreeDTS
Mesoscale simulation of biomembranes with FreeDTS
In this work, the authors report the FreeDTS software to simulate biomembranes at the mesoscale. The software provides various membrane simulations, focusing on protein organization and shape remodeling. A versatile tool propelling realistic membrane studies and diverse applications.
- Weria Pezeshkian
- & John H. Ipsen
-
Article
| Open Access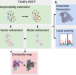 Cryo-EM structure and B-factor refinement with ensemble representation
Cryo-EM structure and B-factor refinement with ensemble representation
Cryo-EM is the go-to method for visualizing large, flexible biomolecules. Here, authors introduce a new Gaussian mixture modelling method for cryo-EM modelling tasks, including refinement, composite map generation and ensemble representation.
- Joseph G. Beton
- , Thomas Mulvaney
- & Maya Topf
-
Article
| Open Access MENDER: fast and scalable tissue structure identification in spatial omics data
MENDER: fast and scalable tissue structure identification in spatial omics data
Identifying tissue structure in large-scale spatial omics datasets from multiple slices is challenging. Here, authors present MENDER, an optimisation-free spatial clustering method that can scale to million-level spatial data, enabling efficient analysis of spatial cell atlases.
- Zhiyuan Yuan
-
Article
| Open Access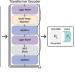 ECOLE: Learning to call copy number variants on whole exome sequencing data
ECOLE: Learning to call copy number variants on whole exome sequencing data
Copy number variants (CNV) are shown to contribute to the etiology of various genetic disorders. Here, authors present ECOLE, a deep learning-based somatic and germline CNV caller for WES data. Utilising a variant of the transformer architecture, the model is trained to call CNVs per exon.
- Berk Mandiracioglu
- , Furkan Ozden
- & A. Ercument Cicek
-
Article
| Open Access rworkflows: automating reproducible practices for the R community
rworkflows: automating reproducible practices for the R community
Reproducibility is essential for the progress of research, yet achieving it remains elusive even in computational fields. Here, authors develop the rworkflows suite, making robust CI/CD workflows easy and freely accessible to all R package developers.
- Brian M. Schilder
- , Alan E. Murphy
- & Nathan G. Skene
-
Article
| Open Access Design automation of microfluidic single and double emulsion droplets with machine learning
Design automation of microfluidic single and double emulsion droplets with machine learning
Generating microfluidic droplets with application-specific desired characteristics is hard. Here the authors report fluid-agnostic machine learning models capable of accurately predicting device geometries and flow conditions required to generate stable single and double emulsions.
- Ali Lashkaripour
- , David P. McIntyre
- & Polly M. Fordyce
-
Article
| Open Access ACIDES: on-line monitoring of forward genetic screens for protein engineering
ACIDES: on-line monitoring of forward genetic screens for protein engineering
Screening mutated proteins is a versatile strategy in protein research, producing massive datasets when combined with NGS. Here, authors present ACIDES to estimate mutated protein fitness and aid protein engineering pipelines in a range of applications, including gene therapy.
- Takahiro Nemoto
- , Tommaso Ocari
- & Ulisse Ferrari
-
Article
| Open Access JOINTLY: interpretable joint clustering of single-cell transcriptomes
JOINTLY: interpretable joint clustering of single-cell transcriptomes
Batch integration is a critical yet challenging step in many single-cell RNA-seq analysis workflows. Here, authors present JOINTLY, a hybrid linear and non-linear NMF-based algorithm, providing interpretable and robust cell clustering against over-integration.
- Andreas Fønss Møller
- & Jesper Grud Skat Madsen
-
Article
| Open Access Pathway centric analysis for single-cell RNA-seq and spatial transcriptomics data with GSDensity
Pathway centric analysis for single-cell RNA-seq and spatial transcriptomics data with GSDensity
Clustering-based analysis has limited power in highly dynamic single-cell data, which is a common situation in tumour samples. Here, authors introduce GSDensity, enabling pathway-centric analysis for the direct integration of data with their domain knowledge.
- Qingnan Liang
- , Yuefan Huang
- & Ken Chen
-
Article
| Open Access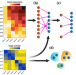 Accurate integration of single-cell DNA and RNA for analyzing intratumor heterogeneity using MaCroDNA
Accurate integration of single-cell DNA and RNA for analyzing intratumor heterogeneity using MaCroDNA
Here, the authors develop MaCroDNA, an algorithm to integrate single-cell DNA and RNA sequencing data from the same tissue. They use MaCroDNA to show—in agreement with previous studies—that copy number changes can predict progression from Barrett’s esophagus to esophageal adenocarcinoma.
- Mohammadamin Edrisi
- , Xiru Huang
- & Luay Nakhleh
-
Article
| Open Access DeepRTAlign: toward accurate retention time alignment for large cohort mass spectrometry data analysis
DeepRTAlign: toward accurate retention time alignment for large cohort mass spectrometry data analysis
Retention time (RT) alignment is a crucial step in large cohort proteomics and metabolomics studies. Here, the authors introduce DeepRTAlign, a deep learning tool for RT alignment that shows high identification sensitivity and quantitative accuracy.
- Yi Liu
- , Yun Yang
- & Cheng Chang
-
Article
| Open Access vcfdist: accurately benchmarking phased small variant calls in human genomes
vcfdist: accurately benchmarking phased small variant calls in human genomes
Accurately benchmarking small variant calling accuracy is critical for the continued improvement of human genome sequencing. Here, the authors show that current approaches are biased towards certain variant representations and develop a new approach to ensure consistent and accurate benchmarking, regardless of the original variant representations.
- Tim Dunn
- & Satish Narayanasamy
-
Article
| Open Access Spatial transcriptomics deconvolution at single-cell resolution using Redeconve
Spatial transcriptomics deconvolution at single-cell resolution using Redeconve
Computational deconvolution with single-cell RNA sequencing data as a reference is pivotal for interpreting spatial transcriptomics data. Here, authors present Redeconve, which improves the resolution by more than 100-fold with higher accuracy and speed.
- Zixiang Zhou
- , Yunshan Zhong
- & Xianwen Ren
-
Article
| Open Access Integrating spatial and single-cell transcriptomics data using deep generative models with SpatialScope
Integrating spatial and single-cell transcriptomics data using deep generative models with SpatialScope
Spatial transcriptomics (ST) is transforming tissue analysis but has limitations. Here, authors introduce SpatialScope, an integrated approach combining scRNA-seq and ST data using deep generative models, enabling comprehensive spatial characterisation at transcriptome-wide single-cell resolution.
- Xiaomeng Wan
- , Jiashun Xiao
- & Can Yang
-
Article
| Open Access On-tissue dataset-dependent MALDI-TIMS-MS2 bioimaging
On-tissue dataset-dependent MALDI-TIMS-MS2 bioimaging
There is a need for dataset-dependent MS2 acquisition in trapped ion mobility spectrometry imaging. Here the authors report spatial ion mobility-scheduled exhaustive fragmentation (SIMSEF) which enables on-tissue metabolite and lipid annotation in mass spectrometry bioimaging studies, and use this to visualise the chemical space in rat brains.
- Steffen Heuckeroth
- , Arne Behrens
- & Robin Schmid
-
Article
| Open Access scReadSim: a single-cell RNA-seq and ATAC-seq read simulator
scReadSim: a single-cell RNA-seq and ATAC-seq read simulator
Benchmarking computational tools for analysis of single-cell sequencing data demands simulation of realistic sequencing reads. However, none of the few existing read simulators aim to mimic real data. Here, the authors introduce scReadSim, a single-cell RNA-seq and ATAC-seq read simulator that works by mimicking real data.
- Guanao Yan
- , Dongyuan Song
- & Jingyi Jessica Li
-
Article
| Open Access Sequence-based prediction of the intrinsic solubility of peptides containing non-natural amino acids
Sequence-based prediction of the intrinsic solubility of peptides containing non-natural amino acids
Posttranslationally modified amino acids are crucial in physiology and drug development as they alter physicochemical properties such as the solubility of proteins. Here the authors describe CamSolPTM, a software that accurately predicts the solubility of proteins containing these residues.
- Marc Oeller
- , Ryan J. D. Kang
- & Michele Vendruscolo
-
Article
| Open Access ProRefiner: an entropy-based refining strategy for inverse protein folding with global graph attention
ProRefiner: an entropy-based refining strategy for inverse protein folding with global graph attention
Inverse Protein Folding is a critical component of protein design. Here, authors introduce ProRefiner, a deep-learning model for IPF that exhibits both high performance and memory efficiency, thereby contributing to advancements in protein design.
- Xinyi Zhou
- , Guangyong Chen
- & Pheng Ann Heng
-
Article
| Open Access A statistical framework for differential pseudotime analysis with multiple single-cell RNA-seq samples
A statistical framework for differential pseudotime analysis with multiple single-cell RNA-seq samples
Pseudotime analysis is prevalent in single-cell RNA-seq, but it remains challenging to perform it across multiple samples and experimental conditions. Here, the authors develop Lamian, a computational framework for multi-sample pseudotime analysis that adjusts for biological and technical variation to detect gene program changes along cell trajectories and across conditions.
- Wenpin Hou
- , Zhicheng Ji
- & Hongkai Ji
-
Article
| Open Access Spatial-linked alignment tool (SLAT) for aligning heterogenous slices
Spatial-linked alignment tool (SLAT) for aligning heterogenous slices
Spatial omics technologies reveal the organisation of cells in various biological systems. Here, authors propose SLAT, a graph-based algorithm for aligning heterogenous data across technologies, modalities and timepoints, enabling spatiotemporal reconstruction of complex developmental processes.
- Chen-Rui Xia
- , Zhi-Jie Cao
- & Ge Gao
-
Article
| Open Access CamoTSS: analysis of alternative transcription start sites for cellular phenotypes and regulatory patterns from 5' scRNA-seq data
CamoTSS: analysis of alternative transcription start sites for cellular phenotypes and regulatory patterns from 5' scRNA-seq data
Five-prime single-cell RNA-seq, especially the read 1, has precise capture of transcription start sites (TSS), but such information is often overlooked. Here, authors present a computational method suite, CamoTSS, to precisely identify TSS and quantify its expression, enabling effective detection of alternative TSS usage in different biological processes.
- Ruiyan Hou
- , Chung-Chau Hon
- & Yuanhua Huang
-
Article
| Open Access trRosettaRNA: automated prediction of RNA 3D structure with transformer network
trRosettaRNA: automated prediction of RNA 3D structure with transformer network
Here, authors develop trRosettaRNA, a deep learning-based approach for predicting RNA 3D structures. Blind tests demonstrate that the automated predictions compete effectively with top human predictions on natural RNAs.
- Wenkai Wang
- , Chenjie Feng
- & Jianyi Yang
-
Article
| Open Access EASTR: Identifying and eliminating systematic alignment errors in multi-exon genes
EASTR: Identifying and eliminating systematic alignment errors in multi-exon genes
The study reveals limitations in widely used RNA-seq aligners, which create 'phantom' introns in reference databases. The authors introduce EASTR, a computational tool that not only enhances alignment accuracy but also uncovers existing annotation errors. This improvement bolsters the dependability of subsequent RNA-seq analyses.
- Ida Shinder
- , Richard Hu
- & Mihaela Pertea

 Crykey: Rapid identification of SARS-CoV-2 cryptic mutations in wastewater
Crykey: Rapid identification of SARS-CoV-2 cryptic mutations in wastewater