Article
|
Open Access
Featured
-
-
Article
| Open Access Biochemical-free enrichment or depletion of RNA classes in real-time during direct RNA sequencing with RISER
Biochemical-free enrichment or depletion of RNA classes in real-time during direct RNA sequencing with RISER
It is difficult to detect low abundance RNAs in sequencing experiments, and biochemical methods to enrich or deplete specific RNAs are time-consuming, costly and can damage RNA. Here, authors develop a biochemical-free technology to enrich or deplete RNA classes in real-time during direct RNA sequencing.
- Alexandra Sneddon
- , Agin Ravindran
- & Eduardo Eyras
-
Article
| Open Access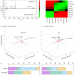 Integrating cryo-OrbiSIMS with computational modelling and metadynamics simulations enhances RNA structure prediction at atomic resolution
Integrating cryo-OrbiSIMS with computational modelling and metadynamics simulations enhances RNA structure prediction at atomic resolution
Conventional structural biology techniques are limited in deciphering complex RNA structures and dynamic interactions. Here the authors show an integrated approach that combines cryogenic OrbiSIMS (cryo-OrbiSIMS) with computational methods for modelling RNA structures at atomic resolution.
- Shannon Ward
- , Alex Childs
- & Aditi N. Borkar
-
Article
| Open Access Mitoribosome structure with cofactors and modifications reveals mechanism of ligand binding and interactions with L1 stalk
Mitoribosome structure with cofactors and modifications reveals mechanism of ligand binding and interactions with L1 stalk
This study uses cryo-EM, biochemical, and computational approaches to shed light on the fundamental mechanisms underlying the human mitoribosome function, including ligand binding, modifications, Fe-S clusters, and aging-related polyamines.
- Vivek Singh
- , Yuzuru Itoh
- & Alexey Amunts
-
Article
| Open Access Emergent ribozyme behaviors in oxychlorine brines indicate a unique niche for molecular evolution on Mars
Emergent ribozyme behaviors in oxychlorine brines indicate a unique niche for molecular evolution on Mars
Mars, an attractive candidate for potential presence of extraterrestrial life, contains oxychlorine species such as perchlorate at its surface. Here, the authors show perchlorate brines support folding and catalysis of functional RNAs, while inactivating representative protein enzymes, and that perchlorate enables new ribozyme functions, including ribozyme catalyzed chlorination of organic molecules.
- Tanner G. Hoog
- , Matthew R. Pawlak
- & Aaron E. Engelhart
-
Article
| Open Access Exon-junction complex association with stalled ribosomes and slow translation-independent disassembly
Exon-junction complex association with stalled ribosomes and slow translation-independent disassembly
Bensaude et al. use a split luciferase approach to show that exon-junction complex assembly and disassembly occur faster when they are translation-dependent than when they are translation-independent; and they uncover an association with ribosomes.
- Olivier Bensaude
- , Isabelle Barbosa
- & Hervé Le Hir
-
Article
| Open Access Structure of HIV-1 RRE stem-loop II identifies two conformational states of the high-affinity Rev binding site
Structure of HIV-1 RRE stem-loop II identifies two conformational states of the high-affinity Rev binding site
HIV relies on the RRE RNA interaction with Rev protein for nuclear export of viral mRNAs. The structure of the high-affinity Rev binding site in RRE in two conformations suggests a mechanism for initial Rev binding and oligomerization onto RRE.
- Jerricho Tipo
- , Keerthi Gottipati
- & Kyung H. Choi
-
Article
| Open Access Ser/Leu-swapped cell-free translation system constructed with natural/in vitro transcribed-hybrid tRNA set
Ser/Leu-swapped cell-free translation system constructed with natural/in vitro transcribed-hybrid tRNA set
The use of orthogonal genetic code can help to prevent the escape of hazardous genes through horizontal gene transfer. Here, the authors develop a cell-free translation system with the Ser/Leu-swapped genetic code using a hybrid tRNA set and show its application in enhancing the production of superfolder GFP.
- Tomoshige Fujino
- , Ryogo Sonoda
- & Hiroshi Murakami
-
Article
| Open Access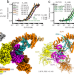 Cryo-EM structures of the human Elongator complex at work
Cryo-EM structures of the human Elongator complex at work
Here the authors determined several cryo-EM structures of the human Elongator complex, which modifies anticodons of tRNAs. The structural work is complemented by functional analyses to understand this clinically relevant cellular machine at the molecular level.
- Nour-el-Hana Abbassi
- , Marcin Jaciuk
- & Sebastian Glatt
-
Article
| Open Access A nascent riboswitch helix orchestrates robust transcriptional regulation through signal integration
A nascent riboswitch helix orchestrates robust transcriptional regulation through signal integration
Here the authors unveil an intermediate state during the folding of the manganese riboswitch from L. lactis. This transient state allows the integration of multiple cellular signals including RNA polymerase pausing and transcription factor NusA.
- Adrien Chauvier
- , Shiba S. Dandpat
- & Nils G. Walter
-
Article
| Open Access Unveiling the A-to-I mRNA editing machinery and its regulation and evolution in fungi
Unveiling the A-to-I mRNA editing machinery and its regulation and evolution in fungi
A-to-I editing in animals is catalyzed by enzymes of the Adenosine Deaminase Acting on RNA family, orthologues of which do not exist in fungi. Here, Feng et al. characterise the enzymes involved in A-to-I mRNA editing in Fusarium graminearum.
- Chanjing Feng
- , Kaiyun Xin
- & Huiquan Liu
-
Article
| Open Access 5′UTR G-quadruplex structure enhances translation in size dependent manner
5′UTR G-quadruplex structure enhances translation in size dependent manner
In eukaryotes, G-quadruplex in mRNA (RG4) 5′ UTR inhibit translation initiation. Here the authors employ single molecule assay to show that RG4 in E. coli reporter increases translation efficiency by preventing ribosome dislodging.
- Chun-Ying Lee
- , Meera Joshi
- & Sua Myong
-
Article
| Open Access Prediction of m6A and m5C at single-molecule resolution reveals a transcriptome-wide co-occurrence of RNA modifications
Prediction of m6A and m5C at single-molecule resolution reveals a transcriptome-wide co-occurrence of RNA modifications
The epitranscriptome holds many unexplored RNA functions, but detecting multiple modifications from one sample remains challenging. Here, authors devise a strategy combining AI and nanopore sequencing to uncover a transcriptome-wide co-occurrence of two modification types in individual RNA molecules.
- P Acera Mateos
- , A J Sethi
- & E Eyras
-
Article
| Open Access Stimulus-responsive assembly of nonviral nucleocapsids
Stimulus-responsive assembly of nonviral nucleocapsids
A stimulus-responsive approach for recapitulating nonviral nucleocapsid assembly on demand under controlled conditions provides a robust platform for applications in synthetic biology and mRNA nanomedicine.
- Mao Hori
- , Angela Steinauer
- & Donald Hilvert
-
Article
| Open Access NMR characterization of RNA binding property of the DEAD-box RNA helicase DDX3X and its implications for helicase activity
NMR characterization of RNA binding property of the DEAD-box RNA helicase DDX3X and its implications for helicase activity
DDX3X is a member of the RNA helicase family, which remodels RNA structures. Using solution NMR, here the authors show that DDX3X preferentially binds to single-stranded RNA, which underlies its unwinding activity toward various structured RNA substrates.
- Yuki Toyama
- & Ichio Shimada
-
Article
| Open Access RNA targeting and cleavage by the type III-Dv CRISPR effector complex
RNA targeting and cleavage by the type III-Dv CRISPR effector complex
Here, Schwartz, Bravo, and Ahsan et al. show how multi-subunit fusion proteins are arranged around a crRNA in a type III CRISPR-Cas effector to cleave target RNA. Structures and molecular dynamics of this complex show three distinct active sites that can be used for programmable RNA cleavage.
- Evan A. Schwartz
- , Jack P. K. Bravo
- & David W. Taylor
-
Article
| Open Access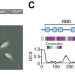 Regulation by the RNA-binding protein Unkempt at its effector interface
Regulation by the RNA-binding protein Unkempt at its effector interface
How RNA-binding proteins (RBPs) regulate gene expression via effectors of RNA processing is unclear. Here, the authors dissect the effector interface of an essential RBP, Unkempt, and investigate its contribution to translational control in cells.
- Kriti Shah
- , Shiyang He
- & Jernej Murn
-
Article
| Open Access Expedient production of site specifically nucleobase-labelled or hypermodified RNA with engineered thermophilic DNA polymerases
Expedient production of site specifically nucleobase-labelled or hypermodified RNA with engineered thermophilic DNA polymerases
A general method for enzymatic synthesis of base-modified RNA was developed using engineered thermostable DNA polymerases enabling introduction of site-specific modifications or synthesis of hypermodified RNA not accessible by in vitro transcription.
- Mária Brunderová
- , Vojtěch Havlíček
- & Michal Hocek
-
Article
| Open Access Predicting nuclear G-quadruplex RNA-binding proteins with roles in transcription and phase separation
Predicting nuclear G-quadruplex RNA-binding proteins with roles in transcription and phase separation
RNA G-quadruplexes are important regulatory elements, yet our knowledge of their structure-based interactions is at present limited. Here the authors combine experimental and computational methods to develop a predictive tool, G4-FUNNIES, to estimate proteins’ RNA G4-binding propensities.
- Johanna Luige
- , Alexandros Armaos
- & Ulf Andersson Vang Ørom
-
Article
| Open Access NAP-seq reveals multiple classes of structured noncoding RNAs with regulatory functions
NAP-seq reveals multiple classes of structured noncoding RNAs with regulatory functions
The genome-wide prevalence, mechanism and function of noncapped RNAs (napRNAs) are currently poorly understood. Here, the authors develop a method called NAP-seq, to globally profile the full-length sequences of napRNAs, revealing several classes of structured noncoding RNAs.
- Shurong Liu
- , Junhong Huang
- & Jianhua Yang
-
Article
| Open Access Structural and mechanistic insights into activation of the human RNA ligase RTCB by Archease
Structural and mechanistic insights into activation of the human RNA ligase RTCB by Archease
RTCB-type RNA ligases play important roles in tRNA splicing, the unfolded protein response and RNA repair. Here, Gerber et al. present structural snapshots of RTCB’s reaction cycle, and show how an activation complex with Archease primes RTCB for ligation.
- Janina Lara Gerber
- , Suria Itzel Morales Guzmán
- & Jirka Peschek
-
Article
| Open Access Toll/interleukin-1 receptor (TIR) domain-containing proteins have NAD-RNA decapping activity
Toll/interleukin-1 receptor (TIR) domain-containing proteins have NAD-RNA decapping activity
Toll/interleukin-1 receptor domain-containing proteins can catabolize NAD+. Here, Wang et al show that these proteins can also function as NAD-RNA decapping enzymes by releasing the NAM moiety from the NAD-RNA, resulting in the regulation of gene expression.
- Xufeng Wang
- , Dongli Yu
- & Xuemei Chen
-
Article
| Open Access Structure-based prediction and characterization of photo-crosslinking in native protein–RNA complexes
Structure-based prediction and characterization of photo-crosslinking in native protein–RNA complexes
Feng et al. developed a computational method PxR3D-map to jointly analyze crosslinked nucleotides and amino acids in protein-RNA complexes, which revealed key structural features underlying photocrosslinking of protein and RNA in cells.
- Huijuan Feng
- , Xiang-Jun Lu
- & Chaolin Zhang
-
Article
| Open Access RNA compaction and iterative scanning for small RNA targets by the Hfq chaperone
RNA compaction and iterative scanning for small RNA targets by the Hfq chaperone
Small RNAs (sRNAs) turn bacterial genes on or off by base pairing with mRNAs. Here the authors employ single molecule fluorescence to show how sRNAs and their chaperone Hfq quickly locate the proper target by repeatedly scanning an mRNA until a stable match is found.
- Ewelina M. Małecka
- & Sarah A. Woodson
-
Article
| Open Access Specificity, synergy, and mechanisms of splice-modifying drugs
Specificity, synergy, and mechanisms of splice-modifying drugs
Two small-molecule drugs, risdiplam and branaplam, have been developed for treating spinal muscular atrophy. Here the authors develop quantitative modeling methods for the sequence-specific and concentration-dependent effects of these and other splice-modifying drugs.
- Yuma Ishigami
- , Mandy S. Wong
- & Justin B. Kinney
-
Article
| Open Access Single-molecule RNA sizing enables quantitative analysis of alternative transcription termination
Single-molecule RNA sizing enables quantitative analysis of alternative transcription termination
The development of RNA technologies demands accurate assessment of transcript size and heterogeneity. Here, authors report a nanopore-based approach to study full-length RNA transcripts at the single-molecule level, identify premature transcription termination and study rolling-circle transcription.
- Gerardo Patiño-Guillén
- , Jovan Pešović
- & Ulrich Felix Keyser
-
Article
| Open Access Fluorogenic CRISPR for genomic DNA imaging
Fluorogenic CRISPR for genomic DNA imaging
Conventional CRISPR-based approaches to monitor genomic loci can be hampered by high background and nonspecific nucleolar signal. Here, the authors propose a fluorogenic CRISPR (fCRISPR) tool that allows for high-contrast and sensitive imaging of genomic DNA.
- Zhongxuan Zhang
- , Xiaoxiao Rong
- & Xing Li
-
Article
| Open Access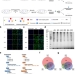 Nuclear and cytoplasmic specific RNA binding proteome enrichment and its changes upon ferroptosis induction
Nuclear and cytoplasmic specific RNA binding proteome enrichment and its changes upon ferroptosis induction
The reported assay shows a subcellular-specific RNA labeling method for efficient enrichment and deep profiling of nuclear and cytoplasmic RBPs, the authors apply this to investigate changes of subcellular-specific RBP-RNA interactions in ferroptosis.
- Haofan Sun
- , Bin Fu
- & Weijie Qin
-
Article
| Open Access 2.7 Å cryo-EM structure of human telomerase H/ACA ribonucleoprotein
2.7 Å cryo-EM structure of human telomerase H/ACA ribonucleoprotein
Here the authors captured the structure of human telomerase H/ACA ribonucleoprotein (RNP) by cryo-EM. The structure rationalizes telomere-disorder disease mutations and reveals insights into the mechanism of pseudouridylation by eukaryotic H/ACA RNPs.
- George E. Ghanim
- , Zala Sekne
- & Thi Hoang Duong Nguyen
-
Article
| Open Access Upregulated hepatic lipogenesis from dietary sugars in response to low palmitate feeding supplies brain palmitate
Upregulated hepatic lipogenesis from dietary sugars in response to low palmitate feeding supplies brain palmitate
The origin of brain palmitic acid (PAM) has been debated. Here, by using natural abundance carbon isotope ratios and RNA sequencing the authors show that the majority of brain PAM is maintained by hepatic PAM synthesis from dietary sugars during development and is upregulated in mice fed low PAM.
- Mackenzie E. Smith
- , Chuck T. Chen
- & Richard P. Bazinet
-
Article
| Open Access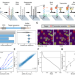 Massively parallel profiling of RNA-targeting CRISPR-Cas13d
Massively parallel profiling of RNA-targeting CRISPR-Cas13d
Systematic understanding of CRISPR enzyme RNA binding specificity and cleavage is lacking. Here the authors report RNA chip-hybridised association-mapping platform (RNA-CHAMP), a workflow that repurposes next generation DNA sequencing chips to measure the binding affinity for RNA targets.
- Hung-Che Kuo
- , Joshua Prupes
- & Ilya J. Finkelstein
-
Article
| Open Access Monovalent metal ion binding promotes the first transesterification reaction in the spliceosome
Monovalent metal ion binding promotes the first transesterification reaction in the spliceosome
Hybrid QM/MM molecular dynamics simulations reveal that the kinetics and thermodynamics of the first splicing step are regulated by a K+ ion that facilitates optimal positioning of reactive moieties.
- Jana Aupič
- , Jure Borišek
- & Alessandra Magistrato
-
Article
| Open Access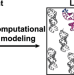 An RNA excited conformational state at atomic resolution
An RNA excited conformational state at atomic resolution
Excited conformation state of biomolecule is transient and high-energy conformation state. Here the authors use NMR spectroscopy and computational modeling to reveal the 3D structure of HIV-1 TAR RNA excited conformational state.
- Ainan Geng
- , Laura Ganser
- & Hashim M. Al-Hashimi
-
Article
| Open Access Extending the toolbox for RNA biology with SegModTeX: a polymerase-driven method for site-specific and segmental labeling of RNA
Extending the toolbox for RNA biology with SegModTeX: a polymerase-driven method for site-specific and segmental labeling of RNA
It has been challenging to make long RNAs with site-specific modifications for NMR study. Here the authors present SegModTeX: a method for site-specific and segmental labeling of RNAs independent of their sequence or segment length, with applications for biological- and artificial NTP analogues at purity and scale sufficient for NMR.
- Raphael Haslecker
- , Vincent V. Pham
- & Victoria M. D’Souza
-
Article
| Open Access Dissecting the basis for differential substrate specificity of ADAR1 and ADAR2
Dissecting the basis for differential substrate specificity of ADAR1 and ADAR2
Human ADAR1 and ADAR2 edit millions of adenosines transcriptome-wide, altering RNA structure. Here the authors show that variations in RNA binding domains influence site-specific editing, enhancing ADAR2-targeted therapeutics.
- Marlon S. Zambrano-Mila
- , Monika Witzenberger
- & Schraga Schwartz
-
Article
| Open Access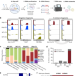 In vivo RNA interactome profiling reveals 3’UTR-processed small RNA targeting a central regulatory hub
In vivo RNA interactome profiling reveals 3’UTR-processed small RNA targeting a central regulatory hub
Here the authors report a new approach to profile RNA-RNA interactions in live bacterial cells. The charted RNA interaction networks unveil a key mRNA regulatory hub targeted by twelve small RNAs, including a novel RNA involved in fatty acid metabolism.
- Fang Liu
- , Ziying Chen
- & Yanjie Chao
-
Article
| Open Access Structural insights into the role of GTPBP10 in the RNA maturation of the mitoribosome
Structural insights into the role of GTPBP10 in the RNA maturation of the mitoribosome
The biogenesis of ribosomes is a highly coordinated process. Here, Nguyen et al. uncover how the mitochondria-specific interplay of the GTPases GTPBP10 and GTPBP7 ensures proper maturation of the catalytic RNA center of the human mitoribosome.
- Thu Giang Nguyen
- , Christina Ritter
- & Eva Kummer
-
Article
| Open Access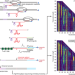 Observation of coordinated RNA folding events by systematic cotranscriptional RNA structure probing
Observation of coordinated RNA folding events by systematic cotranscriptional RNA structure probing
RNA begins to fold as the nascent transcript emerges from a transcribing RNA polymerase. Here, the authors develop a concise method for mapping RNA folding pathways that couples in vitro transcription with high-throughput RNA chemical probing.
- Courtney E. Szyjka
- & Eric J. Strobel
-
Article
| Open Access A single pseudouridine on rRNA regulates ribosome structure and function in the mammalian parasite Trypanosoma brucei
A single pseudouridine on rRNA regulates ribosome structure and function in the mammalian parasite Trypanosoma brucei
Trypanosomes are the causative agent of sleeping sickness. Here the authors demonstrate that the loss of single pseudouridine in ribosomal RNA affects the stoichiometry of ribosomal proteins and translation of a subset of proteins.
- K. Shanmugha Rajan
- , Hava Madmoni
- & Shulamit Michaeli
-
Article
| Open Access Structural and dynamic mechanisms for coupled folding and tRNA recognition of a translational T-box riboswitch
Structural and dynamic mechanisms for coupled folding and tRNA recognition of a translational T-box riboswitch
T-box riboswitches are RNA-based gene regulators, composed of highly structured noncoding RNAs: the T-box and a tRNA ligand. Here, the authors assess the folding of a translational T-box aptamer and dissect the role of Mg2+, intra- and intermolecular RNA-RNA interactions in modulating its folding and function.
- Xiaolin Niu
- , Zhonghe Xu
- & Xianyang Fang
-
Article
| Open Access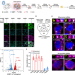 Profiling stress-triggered RNA condensation with photocatalytic proximity labeling
Profiling stress-triggered RNA condensation with photocatalytic proximity labeling
Stress granules (SGs) are highly dynamic cytoplasmic membraneless organelles that assemble when cells are challenged by stress. Herein, the authors apply a proximity-dependent RNA labeling method, CAP-seq, to comprehensively investigate the content of SG-proximal transcriptome and the dynamic change in SG-proximal transcriptome along the time course of granule assembly and disassembly processes in live mammalian cells.
- Ziqi Ren
- , Wei Tang
- & Peng Zou
-
Article
| Open Access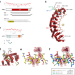 Intra- and inter-molecular regulation by intrinsically-disordered regions governs PUF protein RNA binding
Intra- and inter-molecular regulation by intrinsically-disordered regions governs PUF protein RNA binding
FBF-2 and LST-1 repress gld-1 mRNA expression to maintain C. elegans germline stem cells. The authors show that an intrinsically-disordered region of FBF-2 autoinhibits its RNA binding. LST-1 antagonizes this interaction to promote RNA binding.
- Chen Qiu
- , Zihan Zhang
- & Traci M. Tanaka Hall
-
Article
| Open Access The E3 ligase Riplet promotes RIG-I signaling independent of RIG-I oligomerization
The E3 ligase Riplet promotes RIG-I signaling independent of RIG-I oligomerization
Riplet conjugates K63-Ub chain to RIG-I in order to induce a robust antiviral response, but the mechanism of action remains unclear. Here, the authors show that Riplet recognizes RIG-I regardless of its RNA-bound status and promotes RIG-I signaling independent of RIG-I oligomerization.
- Wenshuai Wang
- , Benjamin Götte
- & Anna Marie Pyle
-
Article
| Open Access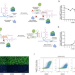 Rational design of microRNA-responsive switch for programmable translational control in mammalian cells
Rational design of microRNA-responsive switch for programmable translational control in mammalian cells
Artificial regulation of translation by intracellular RNAs has many potential applications. Here, authors design a platform capable of miRNA-triggered upregulation or downregulation using a single RNA construct, and demonstrate its use in constructing logic gates and cell-type classifiers.
- Hui Ning
- , Gan Liu
- & Zhen Xie
-
Article
| Open Access MePMe-seq: antibody-free simultaneous m6A and m5C mapping in mRNA by metabolic propargyl labeling and sequencing
MePMe-seq: antibody-free simultaneous m6A and m5C mapping in mRNA by metabolic propargyl labeling and sequencing
Methylation is the dominant modification in mRNA and occurs at a variety of sites. Here, Hartstock et al. show that a clickable analogue of the key cosubstrate S-adenosyl-L-methionine (SAM) can be produced in cells, allowing for identification and mapping of different methylated nucleosides in mRNA.
- Katja Hartstock
- , Nadine A. Kueck
- & Andrea Rentmeister
-
Article
| Open Access RNA-based translation activators for targeted gene upregulation
RNA-based translation activators for targeted gene upregulation
Many diseases are driven by the insufficient expression of critical genes, but few technologies are capable of rescuing these endogenous protein levels. Here, Cao et al. present an RNA-based technology that boosts protein production from endogenous mRNAs by upregulating their translation.
- Yang Cao
- , Huachun Liu
- & Bryan C. Dickinson
-
Article
| Open Access PAPγ associates with PAXT nuclear exosome to control the abundance of PROMPT ncRNAs
PAPγ associates with PAXT nuclear exosome to control the abundance of PROMPT ncRNAs
Pervasive transcription of the human genome generates an abundance of RNAs that must be processed and degraded by the nuclear RNA exosome. Here the authors show that polyA polymerase gamma (PAPγ) associates with PAXT providing key insights into the direct targeting of PROMPT ncRNAs by PAXT at their genomic sites.
- Xavier Contreras
- , David Depierre
- & Rosemary Kiernan
-
Article
| Open Access The SMN complex drives structural changes in human snRNAs to enable snRNP assembly
The SMN complex drives structural changes in human snRNAs to enable snRNP assembly
Sm protein binding to pre-snRNA is a key step in snRNP biogenesis catalyzed by the SMN complex. Here, the authors show that pre-snRNAs adopt compact structures incompatible with Sm protein binding and that Gemin3 and 4 are required for pre-snRNA rearrangement to allow Sm protein interaction.
- Josef Pánek
- , Adriana Roithová
- & David Staněk
-
Article
| Open Access Chemoproteomic capture of RNA binding activity in living cells
Chemoproteomic capture of RNA binding activity in living cells
Here the authors introduce a photo-activatable-competition and chemoproteomic enrichment (PACCE) method to localize protein-RNA interfaces using photoactivatable cellular RNA to protect RNA binding regions on proteins from electrophilic purine probe labeling.
- Andrew J. Heindel
- , Jeffrey W. Brulet
- & Ku-Lung Hsu
-
Article
| Open Access Secondary structures that regulate mRNA translation provide insights for ASO-mediated modulation of cardiac hypertrophy
Secondary structures that regulate mRNA translation provide insights for ASO-mediated modulation of cardiac hypertrophy
The GAT4A transcription factor mediates cardiac development. Here the authors identify that the 5′ UTR of GATA4 mRNA contains a double stranded structure downstream of an upstream open reading frame (uORF) that promotes uORF-mediated suppression of the main ORF.
- Omar M. Hedaya
- , Kadiam C. Venkata Subbaiah
- & Peng Yao

 Structures of co-transcriptional RNA capping enzymes on paused transcription complex
Structures of co-transcriptional RNA capping enzymes on paused transcription complex