Article
|
Open Access
Featured
-
-
Article
| Open Access Exon-junction complex association with stalled ribosomes and slow translation-independent disassembly
Exon-junction complex association with stalled ribosomes and slow translation-independent disassembly
Bensaude et al. use a split luciferase approach to show that exon-junction complex assembly and disassembly occur faster when they are translation-dependent than when they are translation-independent; and they uncover an association with ribosomes.
- Olivier Bensaude
- , Isabelle Barbosa
- & Hervé Le Hir
-
Article
| Open Access Quantifying 3′UTR length from scRNA-seq data reveals changes independent of gene expression
Quantifying 3′UTR length from scRNA-seq data reveals changes independent of gene expression
While gene expression analysis is commonly performed, 3′UTR length analysis is limited due to technical challenges. Here the authors provide an open-access analysis pipeline for scRNA-seq data to simultaneously quantify gene expression and 3′UTR length.
- Mervin M. Fansler
- , Sibylle Mitschka
- & Christine Mayr
-
Article
| Open Access Intramolecular autoinhibition regulates the selectivity of PRPF40A tandem WW domains for proline-rich motifs
Intramolecular autoinhibition regulates the selectivity of PRPF40A tandem WW domains for proline-rich motifs
The specific recognition of a proline-rich motif in the intrinsically disordered region of SF1 by the PRPF40A tandem WW domains is modulated by an intramolecular autoinhibition, suggesting a general mechanism to enhance WW binding selectivity.
- Santiago Martínez-Lumbreras
- , Lena K. Träger
- & Michael Sattler
-
Article
| Open Access CRISPR-dCas13d-based deep screening of proximal and distal splicing-regulatory elements
CRISPR-dCas13d-based deep screening of proximal and distal splicing-regulatory elements
Here the authors develop Splice-RUSH, a high-throughput screening method to map both proximal and distal splicing-regulatory sequences in a native sequence context. These sequences can also be targeted by ASOs to modulate splicing.
- Yocelyn Recinos
- , Dmytro Ustianenko
- & Chaolin Zhang
-
Article
| Open Access Splice modulators target PMS1 to reduce somatic expansion of the Huntington’s disease-associated CAG repeat
Splice modulators target PMS1 to reduce somatic expansion of the Huntington’s disease-associated CAG repeat
Somatic expansion of a CAG repeat in HTT drives onset of Huntington’s disease. Using a human cell line model and splice modulators, here the authors show that PMS1 is an enhancer of CAG repeat expansion, making it a target for therapeutic intervention.
- Zachariah L. McLean
- , Dadi Gao
- & James F. Gusella
-
Article
| Open Access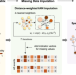 Interrogations of single-cell RNA splicing landscapes with SCASL define new cell identities with physiological relevance
Interrogations of single-cell RNA splicing landscapes with SCASL define new cell identities with physiological relevance
RNA splicing serves as a critical layer of gene expression regulation. Here, authors introduce SCASL for investigating the heterogeneity of RNA splicing landscapes at single-cell resolution, offering a novel scheme for classifying cell identities with physiological relevance.
- Xianke Xiang
- , Yao He
- & Xuerui Yang
-
Article
| Open Access Specificity, synergy, and mechanisms of splice-modifying drugs
Specificity, synergy, and mechanisms of splice-modifying drugs
Two small-molecule drugs, risdiplam and branaplam, have been developed for treating spinal muscular atrophy. Here the authors develop quantitative modeling methods for the sequence-specific and concentration-dependent effects of these and other splice-modifying drugs.
- Yuma Ishigami
- , Mandy S. Wong
- & Justin B. Kinney
-
Article
| Open Access TREX tetramer disruption alters RNA processing necessary for corticogenesis in THOC6 Intellectual Disability Syndrome
TREX tetramer disruption alters RNA processing necessary for corticogenesis in THOC6 Intellectual Disability Syndrome
THOC6 is required for TREX tetramer formation. Analysis of pathogenic THOC6 variants differentiate the conserved mRNA export functions of TREX dimers and RNA processing functions of TREX tetramers underlying THOC6 Intellectual Disability Syndrome.
- Elizabeth A. Werren
- , Geneva R. LaForce
- & Ashleigh E. Schaffer
-
Article
| Open Access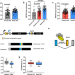 RBFOX2 deregulation promotes pancreatic cancer progression and metastasis through alternative splicing
RBFOX2 deregulation promotes pancreatic cancer progression and metastasis through alternative splicing
The role of alternative splicing in pancreatic cancer (PDAC) development remains to be explored. Here, RBFOX2 is shown to regulate exon splicing events in transcripts encoding proteins involved in cytoskeletal remodelling programs and its downregulation promotes PDAC progression and liver metastasis.
- Michelle Maurin
- , Mohammadreza Ranjouri
- & Karen M. Mann
-
Article
| Open Access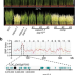 A mitochondrial pentatricopeptide repeat protein enhances cold tolerance by modulating mitochondrial superoxide in rice
A mitochondrial pentatricopeptide repeat protein enhances cold tolerance by modulating mitochondrial superoxide in rice
Cold stress hampers rice growth and yield. This paper demonstrates that mitochondrial superoxide plays a key role in cold responses, and identifies a pentatricopeptide repeat protein which modulates mitochondrial superoxide and rice cold tolerance.
- Xiaofeng Zu
- , Lilan Luo
- & Xiaofeng Cao
-
Article
| Open Access The SMN complex drives structural changes in human snRNAs to enable snRNP assembly
The SMN complex drives structural changes in human snRNAs to enable snRNP assembly
Sm protein binding to pre-snRNA is a key step in snRNP biogenesis catalyzed by the SMN complex. Here, the authors show that pre-snRNAs adopt compact structures incompatible with Sm protein binding and that Gemin3 and 4 are required for pre-snRNA rearrangement to allow Sm protein interaction.
- Josef Pánek
- , Adriana Roithová
- & David Staněk
-
Article
| Open Access Alternative splicing in lung influences COVID-19 severity and respiratory diseases
Alternative splicing in lung influences COVID-19 severity and respiratory diseases
Alternative splicing of transcripts can influence human traits, such as immune response to infection. Here, the authors use mendelian randomization to reveal a role of alternative splicing in lung on COVID-19 severity and susceptibility, offering potential drug discovery avenues.
- Tomoko Nakanishi
- , Julian Willett
- & J. Brent Richards
-
Article
| Open Access Recognition and cleavage mechanism of intron-containing pre-tRNA by human TSEN endonuclease complex
Recognition and cleavage mechanism of intron-containing pre-tRNA by human TSEN endonuclease complex
tRNA splicing is universal. Here the authors report the structure of an active human tRNA splicing endonuclease complex bound to an intron-containing pre-tRNA, which unveils eukaryotic tRNA processing and links archaeal and eukaryotic TSEN evolution.
- Ling Yuan
- , Yaoyao Han
- & Yadong Sun
-
Article
| Open Access Molecular basis of RNA-binding and autoregulation by the cancer-associated splicing factor RBM39
Molecular basis of RNA-binding and autoregulation by the cancer-associated splicing factor RBM39
RBM39 is an essential splicing factor for several cancer cells. Here, the authors described how RBM39 selects RNA at atomic level and autoregulates at the mRNA level by selecting the 3’-splice site of a poison exon.
- Sébastien Campagne
- , Daniel Jutzi
- & Frédéric H-T. Allain
-
Article
| Open Access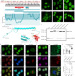 Pharmacological perturbation of the phase-separating protein SMNDC1
Pharmacological perturbation of the phase-separating protein SMNDC1
SMNDC1 is a splicing factor that binds arginine methylation with its Tudor domain. Here, the authors study the protein’s phase-separating behavior and develop small-molecule Tudor domain inhibitors that perturb SMNDC1 function.
- Lennart Enders
- , Marton Siklos
- & Stefan Kubicek
-
Article
| Open Access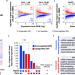 Elevated pre-mRNA 3′ end processing activity in cancer cells renders vulnerability to inhibition of cleavage and polyadenylation
Elevated pre-mRNA 3′ end processing activity in cancer cells renders vulnerability to inhibition of cleavage and polyadenylation
Cancer cells with elevated 3’ end processing activities are vulnerable to CPAi because of transcriptomic disturbances caused by alternative polyadenylation and transcriptional readthrough as well as DNA damages when cells are proliferative.
- Yange Cui
- , Luyang Wang
- & Bin Tian
-
Article
| Open Access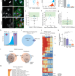 Mislocalization of pathogenic RBM20 variants in dilated cardiomyopathy is caused by loss-of-interaction with Transportin-3
Mislocalization of pathogenic RBM20 variants in dilated cardiomyopathy is caused by loss-of-interaction with Transportin-3
The authors show that loss-of-interaction with the nuclear importer, TNPO3, causes cytoplasmic mislocalization of RBM20 variants linked to severe cases of dilated cardiomyopathy. Restoring their nuclear localization alleviates the disease phenotype.
- Julia Kornienko
- , Marta Rodríguez-Martínez
- & Lars M. Steinmetz
-
Article
| Open Access Structural basis for specific RNA recognition by the alternative splicing factor RBM5
Structural basis for specific RNA recognition by the alternative splicing factor RBM5
The RNA binding protein RBM5 regulates alternative splicing of genes implicated in cancer. Here the authors show structural mechanisms how multiple RNA binding domains of RBM5 cooperate to recognize specific target RNA sequences.
- Komal Soni
- , Pravin Kumar Ankush Jagtap
- & Michael Sattler
-
Article
| Open Access High throughput single cell long-read sequencing analyses of same-cell genotypes and phenotypes in human tumors
High throughput single cell long-read sequencing analyses of same-cell genotypes and phenotypes in human tumors
There is a need for methods that allow the analysis of single-cell long-read sequencing data without depending on known barcode lists or short-read sequencing. Here, the authors develop scNanoGPS, a tool that can independently deconvolute long reads into single cells and single molecules, and apply it on tumour and cell line data.
- Cheng-Kai Shiau
- , Lina Lu
- & Ruli Gao
-
Article
| Open Access Splicing activates transcription from weak promoters upstream of alternative exons
Splicing activates transcription from weak promoters upstream of alternative exons
Few therapeutic strategies are able to upregulate gene expression. Here, the authors developed an approach to activate expression of human genes through small molecules and antisense oligonucleotides that modulate splicing.
- Maritere Uriostegui-Arcos
- , Steven T. Mick
- & Ana Fiszbein
-
Article
| Open Access Mapping PTBP2 binding in human brain identifies SYNGAP1 as a target for therapeutic splice switching
Mapping PTBP2 binding in human brain identifies SYNGAP1 as a target for therapeutic splice switching
Dawicki-McKenna and Felix et al comprehensively map binding and alternative splicing by PTBP2 in human brain and neurons, thus identifying splice switching therapeutic strategies for the neurodevelopmental disorder associated gene SYNGAP1.
- Jennine M. Dawicki-McKenna
- , Alex J. Felix
- & Benjamin L. Prosser
-
Comment
| Open Access The problem of selection bias in studies of pre-mRNA splicing
The problem of selection bias in studies of pre-mRNA splicing
In this comment, the authors discuss the potentially widespread problem of selection bias in drawing biological conclusions from RNA sequencing data.
- Zachary W. Dwyer
- & Jeffrey A. Pleiss
-
Article
| Open Access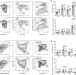 The RNA-binding protein hnRNP F is required for the germinal center B cell response
The RNA-binding protein hnRNP F is required for the germinal center B cell response
The germinal centre (GC) response is characterized by regulated production of high affinity, class-switched antibodies in response to T-cell dependent antigens. Here authors show that the GC response is not only regulated at the transcriptional and protein levels, but also by the RNA-binding protein hnRNP F via alternative splicing of the co-stimulatory molecule CD40.
- Hengjun Huang
- , Yuxing Li
- & Xijun Ou
-
Article
| Open Access Quantitative analysis of C. elegans transcripts by Nanopore direct-cDNA sequencing reveals terminal hairpins in non trans-spliced mRNAs
Quantitative analysis of C. elegans transcripts by Nanopore direct-cDNA sequencing reveals terminal hairpins in non trans-spliced mRNAs
C. elegans long read transcriptomic analysis provides evidence that non-trans-spliced mRNAs display a terminal a hairpin structure mimicking the Spliced Leader. This provides an explanation how the main maturation system might be bypassed.
- Florian Bernard
- , Delphine Dargère
- & Denis Dupuy
-
Article
| Open Access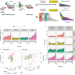 Temporal-iCLIP captures co-transcriptional RNA-protein interactions
Temporal-iCLIP captures co-transcriptional RNA-protein interactions
Dynamic RNA-protein interactions govern the co-transcriptional packaging of RNA polymerase II derived transcripts. Here the authors use temporal-iCLIP which combines transcriptional synchronisation with UV cross-linking of RNA-protein complexes to reveal dynamic RNA-protein interactions during the early phases of transcription and beyond.
- Ross A. Cordiner
- , Yuhui Dou
- & Torben Heick Jensen
-
Article
| Open Access The Heterochromatin protein 1 is a regulator in RNA splicing precision deficient in ulcerative colitis
The Heterochromatin protein 1 is a regulator in RNA splicing precision deficient in ulcerative colitis
This study reports reduced expression of the chromatin and splicing regulator HP1γ in inflammatory bowel disease (IBD) and shows that HP1γ protects against pervasive RNA splicing leading to toxic mRNA products detected in IBD, like progerin.
- Jorge Mata-Garrido
- , Yao Xiang
- & Laurence Arbibe
-
Article
| Open Access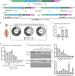 Differential impact of ubiquitous and muscle dynamin 2 isoforms in muscle physiology and centronuclear myopathy
Differential impact of ubiquitous and muscle dynamin 2 isoforms in muscle physiology and centronuclear myopathy
Dynamin 2 is a large GTPase linked to several human diseases. Here, Gómez-Oca et al. investigate the functions of muscle dynamin 2 isoforms and provide insights into their differential implication in centronuclear myopathy pathogenesis and treatment.
- Raquel Gómez-Oca
- , Evelina Edelweiss
- & Jocelyn Laporte
-
Article
| Open Access Splicing factor BUD31 promotes ovarian cancer progression through sustaining the expression of anti-apoptotic BCL2L12
Splicing factor BUD31 promotes ovarian cancer progression through sustaining the expression of anti-apoptotic BCL2L12
Altered expression of splicing factors can contribute to tumour progression. Here the authors show that splicing factor BUD31 enhances ovarian cancer progression by promoting exon inclusion in the anti-apoptotic BCL2 family member, BCL2L12.
- Zixiang Wang
- , Shourong Wang
- & Zhaojian Liu
-
Article
| Open Access The coilin N-terminus mediates multivalent interactions between coilin and Nopp140 to form and maintain Cajal bodies
The coilin N-terminus mediates multivalent interactions between coilin and Nopp140 to form and maintain Cajal bodies
Cajal bodies are membraneless organelles scaffolded by coilin protein. Here, coilin–coilin and coilin–Nopp140 interaction sites are identified and perturbed, revealing coilin’s capacity to form long fibrils and be remodeled into spherical structures.
- Edward Courchaine
- , Sara Gelles-Watnick
- & Karla M. Neugebauer
-
Article
| Open Access Cell-specific regulation of gene expression using splicing-dependent frameshifting
Cell-specific regulation of gene expression using splicing-dependent frameshifting
Precise and reliable gene delivery remains technically challenging. Here, the authors show that rationally designed frameshifting splicing can be used to express genes only in targeted cell types, with the potential to enhance the specificity AAV gene delivery.
- Jonathan P. Ling
- , Alexei M. Bygrave
- & Seth Blackshaw
-
Article
| Open Access Systematic identification of intron retention associated variants from massive publicly available transcriptome sequencing data
Systematic identification of intron retention associated variants from massive publicly available transcriptome sequencing data
This paper proposed a novel in-silico framework for automatically screening disease-related variants and applied it to over 200,000 transcriptomes, providing an example to acquire medically relevant knowledge from publicly available sequence data.
- Yuichi Shiraishi
- , Ai Okada
- & Akihide Yoshimi
-
Article
| Open Access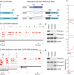 Diverse cell-specific patterns of alternative polyadenylation in Drosophila
Diverse cell-specific patterns of alternative polyadenylation in Drosophila
Single cell data provides cellular resolution on gene expression, but is rarely mined for isoforms. Analysis of 3' isoforms across ~250 Drosophila cell types reveals the cellular bases for numerous tissue-specific 3' programs, identifies new 3' programs, and nominates candidate trans-acting factors
- Seungjae Lee
- , Yen-Chung Chen
- & Eric C. Lai
-
Article
| Open Access Complex regulation of Gephyrin splicing is a determinant of inhibitory postsynaptic diversity
Complex regulation of Gephyrin splicing is a determinant of inhibitory postsynaptic diversity
The protein gephyrin is involved in organizing synapses. Here, the authors show how different transcripts of gephyrin form and regulate inhibitory synapses.
- Raphaël Dos Reis
- , Etienne Kornobis
- & Eric Allemand
-
Article
| Open Access Exploring the cellular landscape of circular RNAs using full-length single-cell RNA sequencing
Exploring the cellular landscape of circular RNAs using full-length single-cell RNA sequencing
Studies of circular RNAs have often been limited to the tissue or organism level. Here, authors investigate the comprehensive expression landscape of circRNAs in human and mouse at single-cell resolution, revealing highly specific and dynamic changes of circRNAs during multiple biological processes.
- Wanying Wu
- , Jinyang Zhang
- & Fangqing Zhao
-
Article
| Open Access Multilayered regulations of alternative splicing, NMD, and protein stability control temporal induction and tissue-specific expression of TRIM46 during axon formation
Multilayered regulations of alternative splicing, NMD, and protein stability control temporal induction and tissue-specific expression of TRIM46 during axon formation
The genetic control underlying axon formation in neurons is unknown. Here, the authors report that neural-specific induction of TRIM46, one of the earliest axonal markers, is regulated by alternative splicing, NMD, and protein stability controls.
- John K. Vuong
- , Volkan Ergin
- & Sika Zheng
-
Article
| Open Access Multilayered control of splicing regulatory networks by DAP3 leads to widespread alternative splicing changes in cancer
Multilayered control of splicing regulatory networks by DAP3 leads to widespread alternative splicing changes in cancer
RNA binding proteins (RBPs) can participate in regulatory networks to control alternative splicing. Here the authors show that DAP3 functions as an RBP splicing modulator via two mechanisms, and that its overexpression leads to mis-splicing events in cancers.
- Jian Han
- , Omer An
- & Leilei Chen
-
Article
| Open Access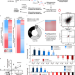 ADARs act as potent regulators of circular transcriptome in cancer
ADARs act as potent regulators of circular transcriptome in cancer
RNA editing and circRNAs are involved in tumorigenesis. Here the authors report that ADARs regulate the circular transcriptome in a bidirectional manner through and beyond their editing function in multiple cancer cells.
- Haoqing Shen
- , Omer An
- & Leilei Chen
-
Article
| Open Access The aberrant upregulation of exon 10-inclusive SREK1 through SRSF10 acts as an oncogenic driver in human hepatocellular carcinoma
The aberrant upregulation of exon 10-inclusive SREK1 through SRSF10 acts as an oncogenic driver in human hepatocellular carcinoma
Alternative splicing is dysregulated in hepatocellular carcinoma. Here, the authors investigate the role of the splice variant of Splicing Regulatory Glutamic Acid and Lysine Rich Protein 1 (SREK1) and its upstream regulator, Serine/arginine-rich splicing factor 10 (SRSF10) in sustaining the oncogenic signal.
- Cunjie Chang
- , Muthukumar Rajasekaran
- & Jianxiang Chen
-
Article
| Open Access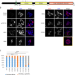 Sirtuin-1 sensitive lysine-136 acetylation drives phase separation and pathological aggregation of TDP-43
Sirtuin-1 sensitive lysine-136 acetylation drives phase separation and pathological aggregation of TDP-43
TDP-43 is a nucleic acid binding protein, whose insoluble aggregates are neuropathological hallmarks of specific subsets of patients with amyotrophic lateral sclerosis and frontotemporal lobar degeneration. Post-translational modifications and acetylation of TDP-43 impact its interaction with RNA, its localization in the cells, and are linked to disease. Using antibodies generated against TDP-43 lysine acetylation sites, sirtuin-1 was found to potently deacetylate amber suppressed [acK136]TDP-43 and reduce its aggregation propensity. Thus, distinct lysine acetylations modulate nuclear import, RNA binding as well as phase separation and aggregation of TDP-43, suggesting regulatory mechanisms for TDP-43 pathogenesis.
- Jorge Garcia Morato
- , Friederike Hans
- & Philipp J. Kahle
-
Article
| Open Access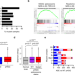 Splicing is an alternate oncogenic pathway activation mechanism in glioma
Splicing is an alternate oncogenic pathway activation mechanism in glioma
Targeting genetic drivers of high grade diffuse glioma (HGG) has not improved patient survival, suggesting the involvement of other mechanisms. Here, across cancer types, the authors identify increased alternative splicing burden in cancer drivers compared to mutation rate as an alternative mechanism for activation of oncogenic pathways such as RAS/MAPK.
- Robert Siddaway
- , Scott Milos
- & Cynthia Hawkins
-
Article
| Open Access Small molecule splicing modifiers with systemic HTT-lowering activity
Small molecule splicing modifiers with systemic HTT-lowering activity
Here the authors describe the discovery of a class of small molecule splicing modifiers which are orally bioavailable, cross the blood-brain barrier, and lower levels of huntingtin in a mouse model of Huntington’s disease (HD).
- Anuradha Bhattacharyya
- , Christopher R. Trotta
- & Stuart W. Peltz
-
Article
| Open Access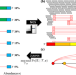 Jumper enables discontinuous transcript assembly in coronaviruses
Jumper enables discontinuous transcript assembly in coronaviruses
@melkebir @psashittal et al. develop a graph-based method for the assembly of discontinuous transcripts produced in Coronaviruses and other Nidovirales, enabling the discovery of transcriptional changes missed by existing methods.
- Palash Sashittal
- , Chuanyi Zhang
- & Mohammed El-Kebir
-
Article
| Open Access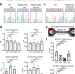 Gain-of-function cardiomyopathic mutations in RBM20 rewire splicing regulation and re-distribute ribonucleoprotein granules within processing bodies
Gain-of-function cardiomyopathic mutations in RBM20 rewire splicing regulation and re-distribute ribonucleoprotein granules within processing bodies
Mutations in the splicing factor RBM20 cause aggressive Dilated Cardiomyopathy. Here the authors generated RBM20 R636S mutants and knockout in human iPSC-derived cardiomyocytes. Mutant RBM20 showed different target RNA binding, altered splicing and localization to cytoplasmic processing bodies.
- Aidan M. Fenix
- , Yuichiro Miyaoka
- & Nathan Salomonis
-
Article
| Open Access SON drives oncogenic RNA splicing in glioblastoma by regulating PTBP1/PTBP2 switching and RBFOX2 activity
SON drives oncogenic RNA splicing in glioblastoma by regulating PTBP1/PTBP2 switching and RBFOX2 activity
Splicing factor PTBP1 is reported to promote oncogenic functions in glioblastoma (GBM). Here the authors show splicing factor SON upregulates PTBP1 expression while supresses its paralog PTBP2 through alternative splicing and the inhibition of SON reduces GBM stemness and growth.
- Jung-Hyun Kim
- , Kyuho Jeong
- & Eun-Young Erin Ahn
-
Article
| Open Access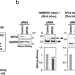 SPF45/RBM17-dependent, but not U2AF-dependent, splicing in a distinct subset of human short introns
SPF45/RBM17-dependent, but not U2AF-dependent, splicing in a distinct subset of human short introns
The length distribution of human pre-mRNA introns is very extensive. The authors demonstrate that splicing in a subset of short introns is dependent on SPF45 (RBM17), which replaces authentic U2AF-heterodimer on the truncated poly-pyrimidine tracts and interacts with the U2 snRNP protein SF3b155.
- Kazuhiro Fukumura
- , Rei Yoshimoto
- & Akila Mayeda
-
Article
| Open Access Nanopore sequencing of brain-derived full-length circRNAs reveals circRNA-specific exon usage, intron retention and microexons
Nanopore sequencing of brain-derived full-length circRNAs reveals circRNA-specific exon usage, intron retention and microexons
Short-read sequencing methods cannot delineate internal exon composition and alternative splicing events of long and multi-exon circular RNAs (circRNAs). Here the authors provide a global map of full-length circRNAs by long-read sequencing in human and mouse brain samples.
- Karim Rahimi
- , Morten T. Venø
- & Jørgen Kjems
-
Article
| Open Access RETRACTED ARTICLE: AKT3-mediated IWS1 phosphorylation promotes the proliferation of EGFR-mutant lung adenocarcinomas through cell cycle-regulated U2AF2 RNA splicing
RETRACTED ARTICLE: AKT3-mediated IWS1 phosphorylation promotes the proliferation of EGFR-mutant lung adenocarcinomas through cell cycle-regulated U2AF2 RNA splicing
IWS1 regulates multiple steps in RNA metabolism, including RNA elongation and alternative RNA splicing. Here the authors show that AKT3 phosphorylates IWS1, which alters U2AF2 RNA splicing and promotes growth of lung adenocarcinomas via a Sororin/ERK-dependent pathway.
- Georgios I. Laliotis
- , Evangelia Chavdoula
- & Philip N. Tsichlis
-
Article
| Open Access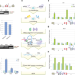 The upstream 5′ splice site remains associated to the transcription machinery during intron synthesis
The upstream 5′ splice site remains associated to the transcription machinery during intron synthesis
We know that most splicing reactions take place co-transcriptionally, but how the transcription machinery facilitate splicing of introns is unknown. Here the authors show that the 5′ splice site remains associated with the transcription machinery during intron synthesis through U1 snRNP, providing a basis for the rapid splicing reaction of introns.
- Yodfat Leader
- , Galit Lev Maor
- & Gil Ast
-
Article
| Open Access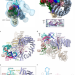 Structural basis of intron selection by U2 snRNP in the presence of covalent inhibitors
Structural basis of intron selection by U2 snRNP in the presence of covalent inhibitors
Chemical modulation of intron selection has emerged as a route for cancer therapy. Here, structures of the U2 snRNP’s SF3B module and of prespliceosome- both in complexes with splicing modulators- provide insight into the mechanisms of intron recognition and branch site inactivation.
- Constantin Cretu
- , Patricia Gee
- & Vladimir Pena

 The debranching enzyme Dbr1 regulates lariat turnover and intron splicing
The debranching enzyme Dbr1 regulates lariat turnover and intron splicing