Article
|
Open Access
Featured
-
-
Article
| Open Access Proteome partitioning constraints in long-term laboratory evolution
Proteome partitioning constraints in long-term laboratory evolution
Adaptive laboratory evolution provides a real-time record of physiological change. In bacteria adapted to glucose over 40 000 generations, this study finds an apparent increase in enzyme efficiency consistent with increased substrate saturation due to loss of a flux sensing mechanism early in adaptation.
- Matteo Mori
- , Vadim Patsalo
- & Matthew Scott
-
Article
| Open Access The Shigella kinase effector OspG modulates host ubiquitin signaling to escape septin-cage entrapment
The Shigella kinase effector OspG modulates host ubiquitin signaling to escape septin-cage entrapment
Here, Xian et al. use phosphoproteomics to identify that the Shigella effector OspG interacts with a regulator of Cullin-RING ubiquitin ligases to promote the ubiquitination of septins and consequent inhibition of septin cage formation.
- Wei Xian
- , Jiaqi Fu
- & Xiaoyun Liu
-
Article
| Open Access Cerebrospinal fluid reference proteins increase accuracy and interpretability of biomarkers for brain diseases
Cerebrospinal fluid reference proteins increase accuracy and interpretability of biomarkers for brain diseases
CSF biomarker concentrations may be influenced by non-disease related interindividual variability. Here, the authors show that reference proteins can capture this variability and enhance the accuracy of Alzheimer’s disease biomarkers.
- Linda Karlsson
- , Jacob Vogel
- & Oskar Hansson
-
Article
| Open Access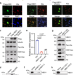 HBO1 catalyzes lysine lactylation and mediates histone H3K9la to regulate gene transcription
HBO1 catalyzes lysine lactylation and mediates histone H3K9la to regulate gene transcription
The regulatory mechanism and functional consequence of lysine lactylation remain to be explored. Here, the authors identify HBO1 as a lysine lactyltransferase and suggest a potential role of HBO1 in tumorigenesis through H3K9la-mediated transcription regulation.
- Ziping Niu
- , Chen Chen
- & Kai Zhang
-
Article
| Open Access An individualized protein-based prognostic model to stratify pediatric patients with papillary thyroid carcinoma
An individualized protein-based prognostic model to stratify pediatric patients with papillary thyroid carcinoma
Papillary thyroid carcinoma has a heterogenous outcome, particularly in paediatric patients. Here, the authors utilise machine learning to create a protein-based prognostic model to predict recurrence risk.
- Zhihong Wang
- , He Wang
- & Yaoting Sun
-
Article
| Open Access Archaeological and molecular evidence for ancient chickens in Central Asia
Archaeological and molecular evidence for ancient chickens in Central Asia
The origin and dispersal of the chicken across Eurasia is unclear. Here, the authors examine eggshell fragments from southern Central Asia with paleoproteomics to identify chicken eggshells, suggesting that chickens may have been an important dietary component as early as 400BCE.
- Carli Peters
- , Kristine K. Richter
- & Robert N. Spengler III
-
Article
| Open Access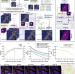 SpiDe-Sr: blind super-resolution network for precise cell segmentation and clustering in spatial proteomics imaging
SpiDe-Sr: blind super-resolution network for precise cell segmentation and clustering in spatial proteomics imaging
Imaging mass cytometry (IMC) is a powerful single-cell resolution platform for targeted spatial proteomics, but it can be constrained by imaging noise and resolution. Here, the authors propose SpiDe-Sr, a super-resolution network embedded with a denoising module for IMC spatial resolution enhancement.
- Rui Chen
- , Jiasu Xu
- & Xianting Ding
-
Article
| Open Access One-Tip enables comprehensive proteome coverage in minimal cells and single zygotes
One-Tip enables comprehensive proteome coverage in minimal cells and single zygotes
Traditional proteomics methods are complex and resource-intensive. Here, the authors develop One-Tip, a highly simplified approach that enables efficient, sensitive, and comprehensive analysis across various sample types, from blood plasma to single cells.
- Zilu Ye
- , Pierre Sabatier
- & Jesper V. Olsen
-
Article
| Open Access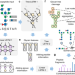 Prediction of glycopeptide fragment mass spectra by deep learning
Prediction of glycopeptide fragment mass spectra by deep learning
Deep learning has achieved a notable success in proteomics and is now emerging in glycoproteomics. Here, the authors develop a neural network-based method to predict mass spectra of intact glycopeptides and demonstrate its potential in data-dependent and data-independent acquisition glycoproteomics.
- Yi Yang
- & Qun Fang
-
Article
| Open Access Thunder-DDA-PASEF enables high-coverage immunopeptidomics and is boosted by MS2Rescore with MS2PIP timsTOF fragmentation prediction model
Thunder-DDA-PASEF enables high-coverage immunopeptidomics and is boosted by MS2Rescore with MS2PIP timsTOF fragmentation prediction model
Human leukocyte antigen (HLA) class I peptide ligands (HLAIps) are targets for developing vaccines and immunotherapies. Here the authors report Thunder-DDA-PASEF, an immunopeptidomics method which enhances the identification of vital HLAIps crucial for vaccine and immunotherapy development.
- David Gomez-Zepeda
- , Danielle Arnold-Schild
- & Stefan Tenzer
-
Article
| Open Access Simultaneous proteome localization and turnover analysis reveals spatiotemporal features of protein homeostasis disruptions
Simultaneous proteome localization and turnover analysis reveals spatiotemporal features of protein homeostasis disruptions
Protein function depends on their subcellular location and turnover rate. Here, the authors report a method to measure spatial and temporal proteome dynamics simultaneously, revealing compartment-specific protein turnover and translocation in cardiac cells under ER stress and carfilzomib treatment.
- Jordan Currie
- , Vyshnavi Manda
- & Edward Lau
-
Article
| Open Access Pick-up single-cell proteomic analysis for quantifying up to 3000 proteins in a Mammalian cell
Pick-up single-cell proteomic analysis for quantifying up to 3000 proteins in a Mammalian cell
Single-cell proteomics is of fundamental importance to capture biological heterogeneity, while limited in proteome depth. Here, the authors develop a pick-up single-cell proteomic analysis (PiSPA) workflow to achieve a deep coverage of quantifying up to 3000 protein groups in a mammalian cell.
- Yu Wang
- , Zhi-Ying Guan
- & Qun Fang
-
Article
| Open Access Nanoparticle enrichment mass-spectrometry proteomics identifies protein-altering variants for precise pQTL mapping
Nanoparticle enrichment mass-spectrometry proteomics identifies protein-altering variants for precise pQTL mapping
Genetic association studies with affinity proteomics face challenges when dealing with protein altering variants. Suhre et al. show that nanoparticle enrichment mass-spectrometry can distinguish between epitope effects and bona fide protein quantitative traits.
- Karsten Suhre
- , Guhan Ram Venkataraman
- & Frank Schmidt
-
Article
| Open Access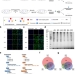 Nuclear and cytoplasmic specific RNA binding proteome enrichment and its changes upon ferroptosis induction
Nuclear and cytoplasmic specific RNA binding proteome enrichment and its changes upon ferroptosis induction
The reported assay shows a subcellular-specific RNA labeling method for efficient enrichment and deep profiling of nuclear and cytoplasmic RBPs, the authors apply this to investigate changes of subcellular-specific RBP-RNA interactions in ferroptosis.
- Haofan Sun
- , Bin Fu
- & Weijie Qin
-
Article
| Open Access Mapping protein states and interactions across the tree of life with co-fractionation mass spectrometry
Mapping protein states and interactions across the tree of life with co-fractionation mass spectrometry
Co-fractionation mass spectrometry (CF-MS) is a powerful technique for mapping protein interactions under physiological conditions. Here, the authors uniformly re-process 411 CF-MS experiments and carry out meta-analyses of protein abundance, protein-protein interactions, and phosphorylation sites in the resulting resource.
- Michael A. Skinnider
- , Mopelola O. Akinlaja
- & Leonard J. Foster
-
Article
| Open Access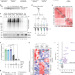 Profiling ubiquitin signalling with UBIMAX reveals DNA damage- and SCFβ-Trcp1-dependent ubiquitylation of the actin-organizing protein Dbn1
Profiling ubiquitin signalling with UBIMAX reveals DNA damage- and SCFβ-Trcp1-dependent ubiquitylation of the actin-organizing protein Dbn1
Using Xenopus egg extracts, the authors developed a mass spectrometry method (UBIMAX) to identify proteins ubiquitylated in response to defined DNA lesions. Highlighting UBIMAX’s versatility, they describe the ubiquitylation of the actin regulator Dbn1 in response to DNA double-strand breaks.
- Camilla S. Colding-Christensen
- , Ellen S. Kakulidis
- & Michael L. Nielsen
-
Article
| Open Access Proximity extracellular protein-protein interaction analysis of EGFR using AirID-conjugated fragment of antigen binding
Proximity extracellular protein-protein interaction analysis of EGFR using AirID-conjugated fragment of antigen binding
Extracellular protein-protein interactions (exPPIs) are essential for understanding the biological function of membrane receptor proteins. Here, the authors demonstrate the FabID technology as a new proximity biotinylation approach that can analyse exPPIs dynamically modulated by drugs and ligands.
- Kohdai Yamada
- , Ryouhei Shioya
- & Tatsuya Sawasaki
-
Article
| Open Access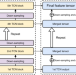 Accurate de novo peptide sequencing using fully convolutional neural networks
Accurate de novo peptide sequencing using fully convolutional neural networks
De novo peptide sequencing allows the identification of peptides without requiring target databases. Here, the authors present PepNet, a convolutional neural network model for accurate de novo peptide sequencing that is capable of analysing large-scale proteomics data.
- Kaiyuan Liu
- , Yuzhen Ye
- & Haixu Tang
-
Article
| Open Access Deep topographic proteomics of a human brain tumour
Deep topographic proteomics of a human brain tumour
Ultrasensitive, spatially-resolved proteomics techniques allow mapping the organisation of healthy and diseased tissues. Here, the authors develop a workflow for spatially-resolved, quantitative tissue proteomics with spatially aware statistics and clustering, with which they characterise a human atypical teratoid-rhabdoid tumour at different spatial resolutions.
- Simon Davis
- , Connor Scott
- & Roman Fischer
-
Article
| Open Access Improved in situ characterization of protein complex dynamics at scale with thermal proximity co-aggregation
Improved in situ characterization of protein complex dynamics at scale with thermal proximity co-aggregation
Vast majority of cellular activities are carried out by protein complexes that assembled dynamically in response to cellular needs and environmental cues. Here, the authors present Slim-TPCA, an effective and readily deployable strategy to unravel the functional roles of protein complexes en masse across various cellular processes.
- Siyuan Sun
- , Zhenxiang Zheng
- & Chris Soon Heng Tan
-
Article
| Open Access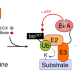 BioE3 identifies specific substrates of ubiquitin E3 ligases
BioE3 identifies specific substrates of ubiquitin E3 ligases
Here, the authors describe BioE3, a biotin-based method to discriminate direct substrates of ubiquitin E3 ligases of interest from mere interactors using proximity proteomics. BioE3 responds to chemical treatments, and works with RING- and HECT-type E3s, as well as ubiquitin-likes (e.g., SUMO).
- Orhi Barroso-Gomila
- , Laura Merino-Cacho
- & James D. Sutherland
-
Article
| Open Access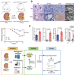 SETD2 deficiency accelerates sphingomyelin accumulation and promotes the development of renal cancer
SETD2 deficiency accelerates sphingomyelin accumulation and promotes the development of renal cancer
SET domain–containing 2 (SETD2) is reported as an immunosuppressor in clear cell renal cell carcinoma (ccRCC). Here the authors show that SETD2 loss enhances de novo sphingomyelin biosynthesis during the transition from polycystic kidney disease to ccRCC.
- Hanyu Rao
- , Changwei Liu
- & Xianting Ding
-
Article
| Open Access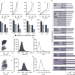 TOFIMS mass spectrometry-based immunopeptidomics refines tumor antigen identification
TOFIMS mass spectrometry-based immunopeptidomics refines tumor antigen identification
MS-based immunopeptidomics provides direct evidence for HLA peptide-antigen presentation, which is indispensable for therapeutic use. Here the authors present an ion mobility MS-based immunopeptidome workflow, largely expand benign reference databases and enables next generation tumor antigen discovery.
- Naomi Hoenisch Gravel
- , Annika Nelde
- & Juliane S. Walz
-
Article
| Open Access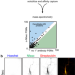 Antibody-directed extracellular proximity biotinylation reveals that Contactin-1 regulates axo-axonic innervation of axon initial segments
Antibody-directed extracellular proximity biotinylation reveals that Contactin-1 regulates axo-axonic innervation of axon initial segments
Few resident cell surface proteins have been identified at the axon initial segment. Here, Ogawa and colleagues use proximity labeling and proteomics to identify Contactin-1 as a transmembrane axon initial segment protein that regulates brain wiring.
- Yuki Ogawa
- , Brian C. Lim
- & Matthew N. Rasband
-
Article
| Open Access Automated imaging and identification of proteoforms directly from ovarian cancer tissue
Automated imaging and identification of proteoforms directly from ovarian cancer tissue
Identification of tissue proteoforms by top-down mass spectrometry remains challenging. Here, the authors present AutoPiMS, a semi-automated multiplexed tandem mass spectrometry workflow for proteoform identification directly from tissue contexts.
- John P. McGee
- , Pei Su
- & Neil L. Kelleher
-
Article
| Open Access SEPepQuant enhances the detection of possible isoform regulations in shotgun proteomics
SEPepQuant enhances the detection of possible isoform regulations in shotgun proteomics
Protein isoform quantification in shotgun proteomics is challenging due to the mapping of many peptides to multiple protein isoforms. Here, the authors present a computational method SEPepQuant and demonstrate its utility in revealing protein isoform level regulation in shotgun proteomics.
- Yongchao Dou
- , Yuejia Liu
- & Bing Zhang
-
Article
| Open Access Next-generation proteomics for quantitative Jumbophage-bacteria interaction mapping
Next-generation proteomics for quantitative Jumbophage-bacteria interaction mapping
In this study Fossati et al. demonstrate how co-fractionation mass spectrometry can be applied to systematically investigate pathogen proteome organization and host interactome plasticity upon Jumbophage infection of Pseudomonas aeruginosa.
- Andrea Fossati
- , Deepto Mozumdar
- & Danielle L. Swaney
-
Article
| Open Access Loss of N-terminal acetyltransferase A activity induces thermally unstable ribosomal proteins and increases their turnover in Saccharomyces cerevisiae
Loss of N-terminal acetyltransferase A activity induces thermally unstable ribosomal proteins and increases their turnover in Saccharomyces cerevisiae
N-terminal acetylation is a common modification with unclear function. Here, using multidimensional proteomics, the authors found that NatA-deficient yeast show increased ribosomal protein degradation and decreased ribosome thermostability, suggesting that N-terminal acetylation enhances proteome stability.
- Ulises H. Guzman
- , Henriette Aksnes
- & Jesper V. Olsen
-
Article
| Open Access Next generation pan-cancer blood proteome profiling using proximity extension assay
Next generation pan-cancer blood proteome profiling using proximity extension assay
Comprehensive and scalable proteomic profiling of plasma samples can improve the screening and diagnosis of cancer patients. Here, the authors use the Olink Proximity Extension Assay technology to characterise the plasma proteomes of 1477 patients across twelve cancer types, and use machine learning to obtain a protein panel for cancer classification.
- María Bueno Álvez
- , Fredrik Edfors
- & Mathias Uhlén
-
Article
| Open Access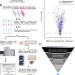 A seven-transmembrane methyltransferase catalysing N-terminal histidine methylation of lytic polysaccharide monooxygenases
A seven-transmembrane methyltransferase catalysing N-terminal histidine methylation of lytic polysaccharide monooxygenases
N-terminal histidine methylation modification has only been observed on certain fungal proteins. Here, the authors identify and validate the methyltransferase responsible for this modification through the combination of mass spectrometry-based proteomics and CRISPR/Cas9.
- Tanveer S. Batth
- , Jonas L. Simonsen
- & Jesper V. Olsen
-
Article
| Open Access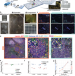 Expanded vacuum-stable gels for multiplexed high-resolution spatial histopathology
Expanded vacuum-stable gels for multiplexed high-resolution spatial histopathology
Emerging high-plex imaging technologies are limited in resolving subcellular biomolecular features. Here, the authors propose a spatial histopathology tool that allows for high-plex protein staining and physical expansion, while retaining the lateral tissue expansion.
- Yunhao Bai
- , Bokai Zhu
- & Sizun Jiang
-
Article
| Open Access The proteomic landscape of soft tissue sarcomas
The proteomic landscape of soft tissue sarcomas
Characterising the molecular profile of soft tissue sarcomas (STS) remains critical. Here, the authors analyse samples from 321 STS patients across 11 histological subtypes using proteomics and identify prognostic signatures that can be applied to multiple subtypes.
- Jessica Burns
- , Christopher P. Wilding
- & Paul H. Huang
-
Article
| Open Access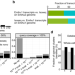 A joint proteomic and genomic investigation provides insights into the mechanism of calcification in coccolithophores
A joint proteomic and genomic investigation provides insights into the mechanism of calcification in coccolithophores
Coccolithophorid algae are globally important for marine biogeochemical cycles, but the molecular basis of their biology is poorly understood. Using proteomics and a new genome, Skeffington et al. identify candidate proteins involved in calcification in Emiliania huxleyi.
- Alastair Skeffington
- , Axel Fischer
- & André Scheffel
-
Article
| Open Access Hybrid-DIA: intelligent data acquisition integrates targeted and discovery proteomics to analyze phospho-signaling in single spheroids
Hybrid-DIA: intelligent data acquisition integrates targeted and discovery proteomics to analyze phospho-signaling in single spheroids
Standard mass spectrometry analyses often miss key targets required for phospho-signalling reconstruction. Here, authors present an intelligent data acquisition strategy that combines discovery and targeted analysis in one run and apply it to maximize the information from single spheroids drug screenings.
- Ana Martínez-Val
- , Kyle Fort
- & Jesper V. Olsen
-
Article
| Open Access Machine learning optimization of candidate antibody yields highly diverse sub-nanomolar affinity antibody libraries
Machine learning optimization of candidate antibody yields highly diverse sub-nanomolar affinity antibody libraries
Therapeutic antibody discovery is time and cost-intensive. Here, the authors develop a machine learning-driven method enabling accelerated design of large and diverse single-chain variable fragments with high binding efficiency, especially at high levels of diversity.
- Lin Li
- , Esther Gupta
- & Matthew E. Walsh
-
Article
| Open Access Unraveling the glycosylated immunopeptidome with HLA-Glyco
Unraveling the glycosylated immunopeptidome with HLA-Glyco
Protein glycosylation plays a vital role in antigen recognition and immune modulation. Here, the authors present a computational workflow for identifying glycosylated peptides from mass spectrometry-based immunopeptidome data and investigate the properties of glycosylated MHC associated peptides.
- Georges Bedran
- , Daniel A. Polasky
- & Alexey I. Nesvizhskii
-
Article
| Open Access FAT-switch-based quantitative S-nitrosoproteomics reveals a key role of GSNOR1 in regulating ER functions
FAT-switch-based quantitative S-nitrosoproteomics reveals a key role of GSNOR1 in regulating ER functions
This study developed a highly sensitive method for detecting S-nitrosylation peptides, which allows quantitative identification of S-nitrosylated proteins and reveals a key role of GSNOR1 in regulating endoplasmic reticulum functions in Arabidopsis.
- Guochen Qin
- , Menghuan Qu
- & Pengcheng Wang
-
Article
| Open Access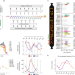 Cellular population dynamics shape the route to human pluripotency
Cellular population dynamics shape the route to human pluripotency
The contribution of cell-extrinsic factors during cellular reprogramming to human induced pluripotent stem cells has long been overlooked. Here, the authors show functional protein communication between reprogramming intermediates and the re-shaping of a permissive extracellular environment.
- Francesco Panariello
- , Onelia Gagliano
- & Nicola Elvassore
-
Article
| Open Access Brain proteomic analysis implicates actin filament processes and injury response in resilience to Alzheimer’s disease
Brain proteomic analysis implicates actin filament processes and injury response in resilience to Alzheimer’s disease
Resilience to Alzheimer’s disease (RAD) is an uncommon combination of high disease burden without dementia. The authors perform proteomic analysis of RAD brains and show lower isocortical and hippocampal soluble Aβ levels, actin filament-based processes, cellular detoxification, and wound healing are significant features.
- Zhi Huang
- , Gennifer E. Merrihew
- & Thomas J. Montine
-
Article
| Open Access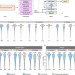 DeepFLR facilitates false localization rate control in phosphoproteomics
DeepFLR facilitates false localization rate control in phosphoproteomics
Protein phosphorylation is a critical modification in many cellular processes. Here, the authors present DeepFLR, a deep learning-based framework to accurately predict phosphopeptide tandem mass spectra and effectively control false localization rates in phosphoproteomics.
- Yu Zong
- , Yuxin Wang
- & Liang Qiao
-
Article
| Open Access Workflow enabling deepscale immunopeptidome, proteome, ubiquitylome, phosphoproteome, and acetylome analyses of sample-limited tissues
Workflow enabling deepscale immunopeptidome, proteome, ubiquitylome, phosphoproteome, and acetylome analyses of sample-limited tissues
Patient samples are often available in limited amounts, restricting the number of possible omics analyses. Here the authors present MONTE, a workflow that enables serial HLA-I and HLA-II immunopeptidome, ubiquitylome, proteome, phosphoproteome, and acetylome data collection from patient samples.
- Jennifer G. Abelin
- , Erik J. Bergstrom
- & Steven A. Carr
-
Article
| Open Access Integrative omics identifies conserved and pathogen-specific responses of sepsis-causing bacteria
Integrative omics identifies conserved and pathogen-specific responses of sepsis-causing bacteria
Severe sepsis has a high mortality rate. Here, the authors provide genomic, transcriptomic, proteomic and metabolomic data across four sepsis-causing pathogens and identify a signature of global increase in fatty acid and lipid biosynthesis as well as cholesterol acquisition.
- Andre Mu
- , William P. Klare
- & Mark J. Walker
-
Article
| Open Access Annexin A1 is a polarity cue that directs mitotic spindle orientation during mammalian epithelial morphogenesis
Annexin A1 is a polarity cue that directs mitotic spindle orientation during mammalian epithelial morphogenesis
Regulation of oriented cell divisions during development is important to position daughter cells and build a structured and functional tissue. Here the authors show that Annexin A1 is a key polarity protein that regulates planar orientation of the cell division axis to guide mammary epithelial morphogenesis.
- Maria Fankhaenel
- , Farahnaz S. Golestan Hashemi
- & Salah Elias
-
Article
| Open Access Benchmarking commonly used software suites and analysis workflows for DIA proteomics and phosphoproteomics
Benchmarking commonly used software suites and analysis workflows for DIA proteomics and phosphoproteomics
Many software suites and spectral libraries have been developed for DIA proteomics data analysis. Here, the authors create benchmark data sets to evaluate four commonly used software tools combined with seven spectral libraries in both global proteomics and phosphoproteomics analysis.
- Ronghui Lou
- , Ye Cao
- & Wenqing Shui
-
Article
| Open Access Multi-omics identify falling LRRC15 as a COVID-19 severity marker and persistent pro-thrombotic signals in convalescence
Multi-omics identify falling LRRC15 as a COVID-19 severity marker and persistent pro-thrombotic signals in convalescence
End-stage kidney disease confers a high risk for severe COVID-19 infection. Using an at-risk group (end-stage kidney disease patients with COVID-19), authors using RNA-sequencing of immune cells and plasma proteomic profiling to investigate the host response to viral infection.
- Jack S. Gisby
- , Norzawani B. Buang
- & James E. Peters
-
Article
| Open Access Phosphoproteomic analysis of neoadjuvant breast cancer suggests that increased sensitivity to paclitaxel is driven by CDK4 and filamin A
Phosphoproteomic analysis of neoadjuvant breast cancer suggests that increased sensitivity to paclitaxel is driven by CDK4 and filamin A
Phosphoproteomics is a promising tool for identifying biomarkers of treatment response in cancer. Here, the authors analyse proteomics profiling of HER2-negative female breast cancer patients and identify potential predictors of paclitaxel response.
- S. Mouron
- , M. J. Bueno
- & M. Quintela-Fandino
-
Article
| Open Access Protein-Peptide Turnover Profiling reveals the order of PTM addition and removal during protein maturation
Protein-Peptide Turnover Profiling reveals the order of PTM addition and removal during protein maturation
Metabolic labeling is often used to measure protein turnover. Here the authors show that for interconvertible protein species like phosphoforms metabolic labeling does not provide information on turnover differences, but that the relative order of modification can determine the observed dynamics.
- Henrik M. Hammarén
- , Eva-Maria Geissen
- & Mikhail M. Savitski
-
Article
| Open Access Integrated proteomic and transcriptomic landscape of macrophages in mouse tissues
Integrated proteomic and transcriptomic landscape of macrophages in mouse tissues
Macrophage is located in different tissue to serve diverse functions. Here the authors use mass spectrometry and bulk RNA-sequencing to profile 11 mouse macrophage populations from 8 tissues, and combine their de novo data with public datasets to report an integrated proteomic and transcriptomic landscape of mouse macrophage as a valuable resource.
- Jingbo Qie
- , Yang Liu
- & Chen Ding
-
Article
| Open Access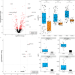 CXCL13 is a predictive biomarker in idiopathic multicentric Castleman disease
CXCL13 is a predictive biomarker in idiopathic multicentric Castleman disease
Idiopathic multicentric Castleman disease (iMCD) is a life-threatening inflammatory disease requiring immediate intervention, for which the recommended first-line therapy is the Interleukin-6 pathway inhibitor siltuximab. Authors here show that the change in levels of the chemokine CXCL13 shortly following the start of siltuximab treatment is predictive of response.
- Sheila K. Pierson
- , Laura Katz
- & David C. Fajgenbaum

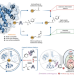 Copper(I)-nitrene platform for chemoproteomic profiling of methionine
Copper(I)-nitrene platform for chemoproteomic profiling of methionine