Article
|
Open Access
Featured
-
-
Article
| Open Access Deciphering bat influenza H18N11 infection dynamics in male Jamaican fruit bats on a single-cell level
Deciphering bat influenza H18N11 infection dynamics in male Jamaican fruit bats on a single-cell level
Here, Kessler et al use single-cell RNA sequencing of the intestine and mesentery from H18N11 influenza-infected bats to show that viral infection is predominant in leukocytes and causes activation of immune cells and antiviral gene signatures.
- Susanne Kessler
- , Bradly Burke
- & Kevin Ciminski
-
Article
| Open Access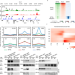 A conserved Pol II elongator SPT6L mediates Pol V transcription to regulate RNA-directed DNA methylation in Arabidopsis
A conserved Pol II elongator SPT6L mediates Pol V transcription to regulate RNA-directed DNA methylation in Arabidopsis
How to facilitate the transcription of plant-specific RNA Pol V is largely unknown. Liu et al. find that a conserved RNA Pol II elongator, SPT6L, mediates DNA methylation by its association with Pol V and promoting the production of scaffold RNA.
- Yujuan Liu
- , Jie Shu
- & Chen Chen
-
Article
| Open Access Multi-omics analysis reveals COVID-19 vaccine induced attenuation of inflammatory responses during breakthrough disease
Multi-omics analysis reveals COVID-19 vaccine induced attenuation of inflammatory responses during breakthrough disease
Here, Drury et al study gene, microRNA and protein expression during COVID-19, in a randomised controlled trial of ChAdOx1 nCoV19 vaccine and find that ChAdOx1 nCoV-19 attenuates the inflammatory response, thought to be the basis for severe COVID-19.
- Ruth E. Drury
- , Susana Camara
- & Daniel O’Connor
-
Article
| Open Access Phylogenomic profiles of whole-genome duplications in Poaceae and landscape of differential duplicate retention and losses among major Poaceae lineages
Phylogenomic profiles of whole-genome duplications in Poaceae and landscape of differential duplicate retention and losses among major Poaceae lineages
Grasses share a whole-genome duplication called rho, but the adaptive implications are unclear. Here, the authors conduct phylogenomic and phylotranscriptomic analyses of 363 grasses, identifying additional whole-genome duplications and finding that duplicates are implicated in environmental adaptations or morphogenesis.
- Taikui Zhang
- , Weichen Huang
- & Hong Ma
-
Article
| Open Access KSNP: a fast de Bruijn graph-based haplotyping tool approaching data-in time cost
KSNP: a fast de Bruijn graph-based haplotyping tool approaching data-in time cost
Haplotyping is the process of distinguishing alleles inherited together on a chromosome, a crucial step in assembling and interpreting genome sequences. Here, the authors present a computationally efficient haplotype assembly tool for long read sequencing data.
- Qian Zhou
- , Fahu Ji
- & Jue Ruan
-
Article
| Open Access Adaptive expansion of ERVK solo-LTRs is associated with Passeriformes speciation events
Adaptive expansion of ERVK solo-LTRs is associated with Passeriformes speciation events
Endogenous retroviruses (ERVs) are remnants of ancient viruses embedded in animal DNA. This study found that the solitary long terminal repeats of ERVs in birds, particularly Passeriformes, have evolved to influence gene expression, potentially contributing to adaptive diversification of species.
- Guangji Chen
- , Dan Yu
- & Shaohong Feng
-
Article
| Open Access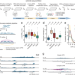 scCircle-seq unveils the diversity and complexity of extrachromosomal circular DNAs in single cells
scCircle-seq unveils the diversity and complexity of extrachromosomal circular DNAs in single cells
Extrachromosomal circular DNAs (eccDNAs) affect gene expression and tumour progression. Here, the authors report a method, scCircle-seq, for eccDNA profiling in single cells, demonstrating the stochasticity, cell type specificity, and dynamics of eccDNAs in cell lines and primary tumour samples.
- Jinxin Phaedo Chen
- , Constantin Diekmann
- & Nicola Crosetto
-
Article
| Open Access Expanded palette of RNA base editors for comprehensive RBP-RNA interactome studies
Expanded palette of RNA base editors for comprehensive RBP-RNA interactome studies
RNA base-editors are often used in methods for RNA binding protein (RBP) target discovery. Here the authors present a new RBP target discovery method, PRINTER, and suggest optimal RNA base-editors for dual-RBP studies, emphasizing the importance of matching rBEs’ editing biases with RBPs’ binding preferences.
- Hugo C. Medina-Munoz
- , Eric Kofman
- & Gene W. Yeo
-
Article
| Open Access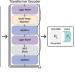 ECOLE: Learning to call copy number variants on whole exome sequencing data
ECOLE: Learning to call copy number variants on whole exome sequencing data
Copy number variants (CNV) are shown to contribute to the etiology of various genetic disorders. Here, authors present ECOLE, a deep learning-based somatic and germline CNV caller for WES data. Utilising a variant of the transformer architecture, the model is trained to call CNVs per exon.
- Berk Mandiracioglu
- , Furkan Ozden
- & A. Ercument Cicek
-
Article
| Open Access Decoding the gene regulatory network of endosperm differentiation in maize
Decoding the gene regulatory network of endosperm differentiation in maize
The cereal endosperm constitutes most of the grain by volume. Here the authors use single-cell analysis of maize developing endosperm to decode gene regulatory networks that likely control endosperm growth and offer a framework for crop improvement.
- Yue Yuan
- , Qiang Huo
- & Zeyang Ma
-
Article
| Open Access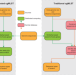 Distributed genotyping and clustering of Neisseria strains reveal continual emergence of epidemic meningococcus over a century
Distributed genotyping and clustering of Neisseria strains reveal continual emergence of epidemic meningococcus over a century
Core genome multilocus sequence typing (cgMLST) is used to classify bacterial strains for epidemiological applications. Here, the authors describe a distributed cgMLST scheme that does not require a central database of allelic sequences, and apply it to study evolutionary patterns of epidemic and endemic strains of the genus Neisseria.
- Ling Zhong
- , Menghan Zhang
- & Zhemin Zhou
-
Article
| Open Access CHEX-seq detects single-cell genomic single-stranded DNA with catalytical potential
CHEX-seq detects single-cell genomic single-stranded DNA with catalytical potential
The in situ single-stranded open chromatin landscape is dynamically regulated in single cells. In their efforts to understand brain cells’ functional dynamics and to complement the other single-cell chromatin approaches, the authors present a method named CHEX-seq (CHromatin EXposed).
- Youtao Lu
- , Jaehee Lee
- & James Eberwine
-
Article
| Open Access PRO-IP-seq tracks molecular modifications of engaged Pol II complexes at nucleotide resolution
PRO-IP-seq tracks molecular modifications of engaged Pol II complexes at nucleotide resolution
Tracking molecular modifications of engaged Pol II complexes is important for studying transcriptional regulation. Here, the authors combine run-on sequencing (PRO-seq) and immunoprecipitation, revealing dynamics of Pol II CTD phosphorylation at nucleotide-resolution.
- Anniina Vihervaara
- , Philip Versluis
- & John T. Lis
-
Article
| Open Access Global mapping of RNA-chromatin contacts reveals a proximity-dominated connectivity model for ncRNA-gene interactions
Global mapping of RNA-chromatin contacts reveals a proximity-dominated connectivity model for ncRNA-gene interactions
Many types of RNAs are associated with chromatin. Here the authors identify chromatin-bound RNAs and their binding sites in human embryonic stem cells suggesting that most chromatin-associated RNAs act proximal to their encoding loci and single RNAs are unlikely to alter gene expression.
- Charles Limouse
- , Owen K. Smith
- & Aaron F. Straight
-
Article
| Open Access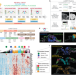 Maintenance of pluripotency-like signature in the entire ectoderm leads to neural crest stem cell potential
Maintenance of pluripotency-like signature in the entire ectoderm leads to neural crest stem cell potential
How the neural crest gains its pluripotency-like stem cell potential is unclear. Here, the authors show that the entire post-gastrula ectoderm maintains expression of pluripotency genes, leading to the high stem cell capacity in the neural crest.
- Ceren Pajanoja
- , Jenny Hsin
- & Laura Kerosuo
-
Article
| Open Access Single cell multiomic analysis reveals diabetes-associated β-cell heterogeneity driven by HNF1A
Single cell multiomic analysis reveals diabetes-associated β-cell heterogeneity driven by HNF1A
The mechanism and disease-relevance of pancreatic b-cell heterogeneity remains elusive. Here the authors show that variable HNF1A-FXYD2 activity drives single b-cell heterogeneity at transcriptomic, epigenomic, and electro-physiological levels, which strongly mark the progression of type 2 diabetes.
- Chen Weng
- , Anniya Gu
- & Yan Li
-
Article
| Open Access Demonstrating paths for unlocking the value of cloud genomics through cross cohort analysis
Demonstrating paths for unlocking the value of cloud genomics through cross cohort analysis
The emergence of large-scale genomics projects has led to genetic studies across cohorts. Here, the authors conduct genome-wide association studies meta-analyzing in trusted research environments or pooling together and find similar, but not identical results.
- Nicole Deflaux
- , Margaret Sunitha Selvaraj
- & Alexander G. Bick
-
Article
| Open Access SEC-seq: association of molecular signatures with antibody secretion in thousands of single human plasma cells
SEC-seq: association of molecular signatures with antibody secretion in thousands of single human plasma cells
Linking proteins secreted from individual cells with other cellular information is challenging. Here, authors report a high-throughput method which uses hydrogel nanovials loaded with single cells to link the secretion profile of individual cells with their surface markers and transcriptomic data.
- Rene Yu-Hong Cheng
- , Joseph de Rutte
- & Richard G. James
-
Article
| Open Access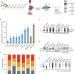 In vivo PAR-CLIP (viP-CLIP) of liver TIAL1 unveils targets regulating cholesterol synthesis and secretion
In vivo PAR-CLIP (viP-CLIP) of liver TIAL1 unveils targets regulating cholesterol synthesis and secretion
Here the authors develop viP-CLIP, a technique allowing the identification of RNA binding proteins (RBP) targets in mouse tissues, and identify the RBP Tial1 as a regulator of cholesterol metabolism in the liver.
- Hasan Vatandaslar
- , Aitor Garzia
- & Markus Stoffel
-
Article
| Open Access High-throughput and high-accuracy single-cell RNA isoform analysis using PacBio circular consensus sequencing
High-throughput and high-accuracy single-cell RNA isoform analysis using PacBio circular consensus sequencing
Long-read single-cell RNA isoform sequencing can elucidate the intricate landscape of alternative RNA splicing in individual cells, but it suffers from a low read throughput. Here, the authors develop circular consensus sequencing methods to allow high-throughput and high-accuracy single-cell RNA isoform sequencing.
- Zhuo-Xing Shi
- , Zhi-Chao Chen
- & Yi-Zhi Liu
-
Article
| Open Access Identifying high-impact variants and genes in exomes of Ashkenazi Jewish inflammatory bowel disease patients
Identifying high-impact variants and genes in exomes of Ashkenazi Jewish inflammatory bowel disease patients
Inflammatory bowel disease (IBD) is highly prevalent among the Ashkenazi Jewish population. Here, the authors identify novel IBD-associated variants and genes, validated by transcriptomic and phenome-wide associations.
- Yiming Wu
- , Kyle Gettler
- & Yuval Itan
-
Article
| Open Access Characterization of genome-wide STR variation in 6487 human genomes
Characterization of genome-wide STR variation in 6487 human genomes
Short tandem repeat studies in humans have often focused on European populations. Here, the authors report a comprehensive map of 366,013 polymorphic short tandem repeats in Chinese individuals and their mutational patterns, functional properties, gene regulatory effects and population characteristics.
- Yirong Shi
- , Yiwei Niu
- & Shunmin He
-
Article
| Open Access The in vivo measurement of replication fork velocity and pausing by lag-time analysis
The in vivo measurement of replication fork velocity and pausing by lag-time analysis
Lag-time analysis was developed to measure in vivo replisome dynamics. Observed dynamics are both locus and cell-cycle dependent: Pauses of seconds are observed at wild-type ribosomal DNA loci, as well as temporal fork velocity oscillations.
- Dean Huang
- , Anna E. Johnson
- & Paul A. Wiggins
-
Article
| Open Access Spatial transcriptomics using multiplexed deterministic barcoding in tissue
Spatial transcriptomics using multiplexed deterministic barcoding in tissue
Examining the spatially resolved transcriptome of tissue sections promises advances in biomedical research. Here, the authors present xDBiT, a versatile, microfluidics-based approach to cost-effectively measure the spatial transcriptome of multiple tissue sections in parallel.
- Johannes Wirth
- , Nina Huber
- & Matthias Meier
-
Article
| Open Access Key innovations and the diversification of Hymenoptera
Key innovations and the diversification of Hymenoptera
Hymenoptera is an incredibly diverse order, with numerous behavioral and morphological innovations. Here, the authors compile a time-calibrated Hymenoptera phylogeny and find that secondary transitions to phytophagy, plant feeding, are associated with significant increases in diversification rate in this group.
- Bonnie B. Blaimer
- , Bernardo F. Santos
- & Matthew L. Buffington
-
Article
| Open Access ChIATAC is an efficient strategy for multi-omics mapping of 3D epigenomes from low-cell inputs
ChIATAC is an efficient strategy for multi-omics mapping of 3D epigenomes from low-cell inputs
Methods connecting genes to their cis-regulatory elements usually requires millions of input cells. Here the authors combine proximity ligation in ChIA-PET and transposase accessibility in ATAC-seq, reporting ChIATAC, to map interactions between open chromatin loci in low numbers of input cells.
- Haoxi Chai
- , Harianto Tjong
- & Yijun Ruan
-
Article
| Open Access A method for multiplexed full-length single-molecule sequencing of the human mitochondrial genome
A method for multiplexed full-length single-molecule sequencing of the human mitochondrial genome
Accurate analysis of mitochondrial DNA is important for mitochondrial disease clinical research and diagnostics. Here, authors present a method using Cas9 cleavage, nanopore sequencing and a custom pipeline to identify pathogenic variants, deletions and accurately quantify heteroplasmy to below 1%.
- Ieva Keraite
- , Philipp Becker
- & Ivo Glynne Gut
-
Article
| Open Access Complex genomic patterns of abasic sites in mammalian DNA revealed by a high-resolution SSiNGLe-AP method
Complex genomic patterns of abasic sites in mammalian DNA revealed by a high-resolution SSiNGLe-AP method
Abasic (AP) sites represent a prominent type of DNA damage, yet the genomics of this lesion remains unexplored. Here, the authors report a method to map such sites at the nucleotide level in complex genomes and use it to extract complex age- and tissue-dependent patterns of AP sites in mammals.
- Ye Cai
- , Huifen Cao
- & Philipp Kapranov
-
Article
| Open Access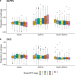 DNA methylation signatures of Alzheimer’s disease neuropathology in the cortex are primarily driven by variation in non-neuronal cell-types
DNA methylation signatures of Alzheimer’s disease neuropathology in the cortex are primarily driven by variation in non-neuronal cell-types
Here the authors identify differences in cortical DNA methylation associated with Alzheimer’s disease pathology, and profiling nuclei from specific cell-types, find that most of these differences reflect variation occurring in non-neuronal cells.
- Gemma Shireby
- , Emma L. Dempster
- & Jonathan Mill
-
Article
| Open Access Combined comparative genomics and clinical modeling reveals plasmid-encoded genes are independently associated with Klebsiella infection
Combined comparative genomics and clinical modeling reveals plasmid-encoded genes are independently associated with Klebsiella infection
Patient variables, such as comorbidities, partially explain which patients will progress to Klebsiella infection, with colonization of the gut acting as a reservoir. Little is known, however, regarding Klebsiella genes that may increase risk of disease in colonized individuals. Here, authors conduct a comparative genomics study to identify genes associated with progression from colonisation to infection.
- Jay Vornhagen
- , Emily K. Roberts
- & Michael A. Bachman
-
Article
| Open Access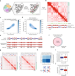 Polycomb-lamina antagonism partitions heterochromatin at the nuclear periphery
Polycomb-lamina antagonism partitions heterochromatin at the nuclear periphery
Here the authors developed ‘Lamina-Inducible Methylation and Hi-C’ (LIMe-Hi-C) to simultaneously measure chromosome conformation, DNA methylation, and nuclear lamina positioning. Application of the method revealed dynamic changes upon PRC2 inhibition and an essential function of H3K27me3 in regulating sub-compartments and lamina association.
- Allison P. Siegenfeld
- , Shelby A. Roseman
- & Brian B. Liau
-
Article
| Open Access RIViT-seq enables systematic identification of regulons of transcriptional machineries
RIViT-seq enables systematic identification of regulons of transcriptional machineries
Here the authors present their method ‘regulon identification by in vitro transcription-sequencing’ (RIViT-seq), which enables systematic identification of target genes of transcription factors of interest. They applied RIViT-seq to 13 sigma factors from Streptomyces coelicolor A3(2) and successfully identified target genes of 11 of these, expanding the regulatory characterisation in this organism.
- Hiroshi Otani
- & Nigel J. Mouncey
-
Article
| Open Access Pervasive misannotation of microexons that are evolutionarily conserved and crucial for gene function in plants
Pervasive misannotation of microexons that are evolutionarily conserved and crucial for gene function in plants
The small size (≤15-nt) of micorexons poses difficulties for genome annotation and identification using standard RNA sequence mapping approaches. Here, the authors develop computational pipelines to discover and predict microexons in plants and reveal diverse evolutionary trajectories via genomewide microexon modeling.
- Huihui Yu
- , Mu Li
- & Chi Zhang
-
Article
| Open Access Biological heterogeneity in idiopathic pulmonary arterial hypertension identified through unsupervised transcriptomic profiling of whole blood
Biological heterogeneity in idiopathic pulmonary arterial hypertension identified through unsupervised transcriptomic profiling of whole blood
Idiopathic pulmonary arterial hypertension is a rare and fatal disease with a heterogeneous treatment response. Here the authors show that unsupervised machine learning of whole blood transcriptomes from 359 patients with idiopathic pulmonary arterial hypertension identifies 3 subgroups (endophenotypes) that improve risk stratification and provide new molecular insights.
- Sokratis Kariotis
- , Emmanuel Jammeh
- & Richard C. Trembath
-
Article
| Open Access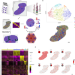 Spatial Transcriptomics to define transcriptional patterns of zonation and structural components in the mouse liver
Spatial Transcriptomics to define transcriptional patterns of zonation and structural components in the mouse liver
Global transcriptional differences across lobular units in the liver remain unknown. Here the authors perform spatial transcriptomics of liver tissue to delineate transcriptional differences in physical space, confirm lobular zonation along transcriptional gradients and suggest the presence of previously uncharacterized structures within liver tissue.
- Franziska Hildebrandt
- , Alma Andersson
- & Johan Ankarklev
-
Article
| Open Access High-throughput RNA sequencing of paraformaldehyde-fixed single cells
High-throughput RNA sequencing of paraformaldehyde-fixed single cells
Current high-throughput single-cell transcriptomic methods are incompatible with paraformaldehyde, a common cell fixation technique. Here the authors present FD-seq, a method for droplet-based RNA sequencing of paraformaldehyde-fixed, stained and sorted single cells.
- Hoang Van Phan
- , Michiel van Gent
- & Savaş Tay
-
Article
| Open Access Development and implementation of a scalable and versatile test for COVID-19 diagnostics in rural communities
Development and implementation of a scalable and versatile test for COVID-19 diagnostics in rural communities
Here, the authors report the development of a versatile academic, SARSCoV-2 RT-qPCR molecular diagnostic test that uses 3D printed technology for sample collection, is implemented in rural setting in the US state of Virginia and validated in its population.
- A. Ceci
- , C. Muñoz-Ballester
- & C. V. Finkielstein
-
Article
| Open Access Nuclear ADP-ribosylation drives IFNγ-dependent STAT1α enhancer formation in macrophages
Nuclear ADP-ribosylation drives IFNγ-dependent STAT1α enhancer formation in macrophages
STAT1a is required for pro-inflammatory responses in macrophages. Here the authors reveal that post-translational modification of STAT1a, ADPribosylation, plays a critical role in enhancer formation and activation, thus modulating IFNγ-stimulated inflammatory responses in macrophages.
- Rebecca Gupte
- , Tulip Nandu
- & W. Lee Kraus
-
Article
| Open Access COVseq is a cost-effective workflow for mass-scale SARS-CoV-2 genomic surveillance
COVseq is a cost-effective workflow for mass-scale SARS-CoV-2 genomic surveillance
Genomic surveillance of SARS-CoV-2 is crucial to monitor the spread of variants of concern. A new sequencing method enables cost-effective SARS-CoV-2 genomic surveillance at scale and is easily adaptable to other viruses.
- Michele Simonetti
- , Ning Zhang
- & Nicola Crosetto
-
Article
| Open Access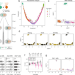 Integrated analysis of Xist upregulation and X-chromosome inactivation with single-cell and single-allele resolution
Integrated analysis of Xist upregulation and X-chromosome inactivation with single-cell and single-allele resolution
X-chromosome inactivation (XCI) ensures dosage compensation between the sexes. Here the authors perform allele-specific single-cell RNA sequencing in differentiating mouse embryonic stem cells to provide a detailed profile of the onset of XCI.
- Guido Pacini
- , Ilona Dunkel
- & Edda G. Schulz
-
Article
| Open Access Spatio-temporal mRNA tracking in the early zebrafish embryo
Spatio-temporal mRNA tracking in the early zebrafish embryo
Early stages of embryogenesis are known to depend on subcellular localization and transport of maternal mRNA, but systematic analyses have been hindered by a lack of methods for tracking of RNA. Here the authors combine spatially-resolved transcriptomics and single-cell RNA labeling to perform a spatio-temporal analysis of the transcriptome during early zebrafish development, revealing insights into this process.
- Karoline Holler
- , Anika Neuschulz
- & Jan Philipp Junker
-
Article
| Open Access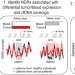 Tissue-specific cell-free DNA degradation quantifies circulating tumor DNA burden
Tissue-specific cell-free DNA degradation quantifies circulating tumor DNA burden
Circulating tumour DNA (ctDNA) represents a non-invasive option to monitor cancer progression. Here, the authors perform deep sequencing of plasma cell-free DNA, and find that nucleosome-dependent cfDNA degradation at 6 specific regulatory regions is predictive of ctDNA burden.
- Guanhua Zhu
- , Yu A. Guo
- & Anders J. Skanderup
-
Article
| Open Access Genome-wide binding potential and regulatory activity of the glucocorticoid receptor’s monomeric and dimeric forms
Genome-wide binding potential and regulatory activity of the glucocorticoid receptor’s monomeric and dimeric forms
Glucocorticoid receptors (GR) are thought to bind DNA as dimers or monomers, to regulate different transcription pathways. Here, the authors perform genome-wide studies on GRs with mutations that impair dimerization and provide evidence that monomeric GRs do not play a significant physiologic role.
- Thomas A. Johnson
- , Ville Paakinaho
- & Diego M. Presman
-
Article
| Open Access Aptardi predicts polyadenylation sites in sample-specific transcriptomes using high-throughput RNA sequencing and DNA sequence
Aptardi predicts polyadenylation sites in sample-specific transcriptomes using high-throughput RNA sequencing and DNA sequence
Short read RNA sequencing and DNA sequence contain useful information for profiling polyadenylation sites, but each also possesses inherent limitations when examined independently. Aptardi combines these data and significantly improves annotation of polyadenylation sites in the expressed transcriptome.
- Ryan Lusk
- , Evan Stene
- & Laura M. Saba
-
Article
| Open Access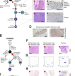 Somatic mutations and single-cell transcriptomes reveal the root of malignant rhabdoid tumours
Somatic mutations and single-cell transcriptomes reveal the root of malignant rhabdoid tumours
Malignant rhabdoid tumours (MRT) have been suggested to originate in the ectoderm-derived neural crest. Here, the authors analyse MRTs using phylogenetics, scRNA-seq, and patient-derived organoids; they find evidence for an MRT origin in the neural crest lineage and suggest differentiation treatment with HDAC/mTOR inhibitors.
- Lars Custers
- , Eleonora Khabirova
- & Jarno Drost
-
Article
| Open Access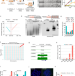 Telomeres reforged with non-telomeric sequences in mouse embryonic stem cells
Telomeres reforged with non-telomeric sequences in mouse embryonic stem cells
Telomeres can be maintained by a telomerase-independent mechanism called an alternative lengthening of telomeres (ALT). Here the authors use mouse Terc (telomerase RNA) knockout embryonic cells and provide longitudinal analysis of ALT telomeres maintained with non-telomeric sequences.
- Chuna Kim
- , Sanghyun Sung
- & Junho Lee
-
Article
| Open Access Radiation-induced DNA damage and repair effects on 3D genome organization
Radiation-induced DNA damage and repair effects on 3D genome organization
Genomic aberrations disrupting chromosome spatial domains can lead to disease. Here, the authors investigate the impact of DNA damage response and repair on 3D genome folding, comparing wild type cells and ataxia telangiectasia mutated patient cells, and characterise both cell type-specific and shared changes to genome organization during the response to damage.
- Jacob T. Sanders
- , Trevor F. Freeman
- & Rachel Patton McCord
-
Article
| Open Access Identification and characterization of constrained non-exonic bases lacking predictive epigenomic and transcription factor binding annotations
Identification and characterization of constrained non-exonic bases lacking predictive epigenomic and transcription factor binding annotations
Genome-wide maps of evolutionary constraint and large-scale compendia of epigenomic and transcription factor data provide complementary information for genome annotation. Here, the authors develop the Constrained Non-Exonic Predictor (CNEP) that enables better understanding of their relationship.
- Olivera Grujic
- , Tanya N. Phung
- & Jason Ernst
-
Article
| Open Access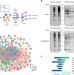 Proteomic profiling and genome-wide mapping of O-GlcNAc chromatin-associated proteins reveal an O-GlcNAc-regulated genotoxic stress response
Proteomic profiling and genome-wide mapping of O-GlcNAc chromatin-associated proteins reveal an O-GlcNAc-regulated genotoxic stress response
Protein O-GlcNAcylation is involved in regulating gene expression and maintaining cellular homeostasis. Here, the authors develop a chemical reporter-based strategy for the proteomic profiling and genome-wide mapping of genotoxic stress-induced O-GlcNAcylated chromatin-associated proteins.
- Yubo Liu
- , Qiushi Chen
- & Jianing Zhang

 Nucleosomal DNA has topological memory
Nucleosomal DNA has topological memory