Featured
-
-
Article
| Open Access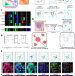 Mapping human tissues with highly multiplexed RNA in situ hybridization
Mapping human tissues with highly multiplexed RNA in situ hybridization
Application of multiplexed RNA in situ mapping techniques to human tissues remains challenging. Here, the authors report DART-FISH, a padlock probe-based technology capable of profiling large numbers of genes in centimetre-sized human tissue sections.
- Kian Kalhor
- , Chien-Ju Chen
- & Kun Zhang
-
Article
| Open Access Highly sensitive spatial transcriptomics using FISHnCHIPs of multiple co-expressed genes
Highly sensitive spatial transcriptomics using FISHnCHIPs of multiple co-expressed genes
Leveraging the fact that eukaryotic genomes are organized into gene modules, FISHnCHIPs images multiple co-expressed genes simultaneously for sensitive and high throughput profiling of gene programs and cell types in tissues.
- Xinrui Zhou
- , Wan Yi Seow
- & Kok Hao Chen
-
Article
| Open Access Kdm1a safeguards the topological boundaries of PRC2-repressed genes and prevents aging-related euchromatinization in neurons
Kdm1a safeguards the topological boundaries of PRC2-repressed genes and prevents aging-related euchromatinization in neurons
Kdm1a is a histone demethylase implicated in intellectual disability. Here, the authors show that removing Kdm1a in neurons of the adult mouse forebrain disrupts silencing of nonneuronal genes and chromatin organization, emphasizing its role in preserving neuronal genome integrity.
- Beatriz del Blanco
- , Sergio Niñerola
- & Ángel Barco
-
Article
| Open Access A single cell atlas of frozen shoulder capsule identifies features associated with inflammatory fibrosis resolution
A single cell atlas of frozen shoulder capsule identifies features associated with inflammatory fibrosis resolution
Unlike most inflammatory fibrotic conditions, frozen shoulder is a spontaneously self-resolving human disease. Here authors study samples from frozen shoulder capsules by single cell RNA sequencing and by microculture modelling of cell-cell interactions to conclude that specific macrophage populations and their interaction with fibroblasts might promote fibrosis resolution.
- Michael T. H. Ng
- , Rowie Borst
- & Stephanie G. Dakin
-
Article
| Open Access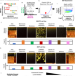 Anti-correlated feature selection prevents false discovery of subpopulations in scRNAseq
Anti-correlated feature selection prevents false discovery of subpopulations in scRNAseq
Typical single-cell RNAseq pipelines will subcluster homogeneous cells. Here, authors present a computational algorithm for accurately identifying cell-type marker genes in single-cell data analysis with a low false discovery rate.
- Scott R. Tyler
- , Daniel Lozano-Ojalvo
- & Eric E. Schadt
-
Article
| Open Access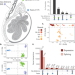 Molecular quantitative trait loci in reproductive tissues impact male fertility in cattle
Molecular quantitative trait loci in reproductive tissues impact male fertility in cattle
Investigating the genetics of male fertility requires comprehensive genotype and phenotype data. Here, the authors characterize the transcriptional complexity of bovine male reproductive tissues to identify loci associated with male fertility.
- Xena Marie Mapel
- , Naveen Kumar Kadri
- & Hubert Pausch
-
Article
| Open Access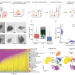 Fragment-sequencing unveils local tissue microenvironments at single-cell resolution
Fragment-sequencing unveils local tissue microenvironments at single-cell resolution
Experimentally preserving tissue microenvironments remains challenging for single-cell sequencing methods. Here, the authors introduce fragment-sequencing, a method that can preserve three-dimensional microenvironments in single-cell RNA-seq, thus allowing the reconstruction of spatial tissue niches.
- Kristina Handler
- , Karsten Bach
- & Andreas E. Moor
-
Article
| Open Access Tracing immune cells around biomaterials with spatial anchors during large-scale wound regeneration
Tracing immune cells around biomaterials with spatial anchors during large-scale wound regeneration
Skin scarring devoid of dermal appendages has unfavorable effects on aesthetic and physiological functions. Here, the authors present a treatment based on extracellular matrix scaffolds and perform multimodal analysis to highlight the role of Tregs recruited by the biomaterial in mitigating tissue fibrous by suppressing excessive inflammation.
- Yang Yang
- , Chenyu Chu
- & Yili Qu
-
Article
| Open Access Zwitterionic microgel preservation platform for circulating tumor cells in whole blood specimen
Zwitterionic microgel preservation platform for circulating tumor cells in whole blood specimen
Blood specimen stabilization for the preservation of circulating tumor cells remains challenging. Here, the authors present a zwitterionic microgel platform for long-term hypothermic preservation of circulating tumor cells in the whole blood of cancer patients for noninvasive diagnostics.
- Yiming Ma
- , Jun Zhang
- & Lei Zhang
-
Article
| Open Access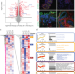 An integrated organoid omics map extends modeling potential of kidney disease
An integrated organoid omics map extends modeling potential of kidney disease
Lassé et al. show that genes involved in kidney organoid proteomic response to TNFα segregate a subset of individuals with poor outcomes in proteinuric kidney disease, demonstrating the relevance of kidney organoid modeling to human kidney disease.
- Moritz Lassé
- , Jamal El Saghir
- & Markus M. Rinschen
-
Article
| Open Access Characterizing cancer metabolism from bulk and single-cell RNA-seq data using METAFlux
Characterizing cancer metabolism from bulk and single-cell RNA-seq data using METAFlux
Metabolic reprogramming is a common indicator of the tumour microenvironment. Here the authors develop the METAflux framework to predict metabolic fluxes from single cell RNA-seq data.
- Yuefan Huang
- , Vakul Mohanty
- & Ken Chen
-
Article
| Open Access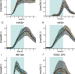 Multidimensional characterization of inducible promoters and a highly light-sensitive LOV-transcription factor
Multidimensional characterization of inducible promoters and a highly light-sensitive LOV-transcription factor
The ability to independently control the expression of different genes is important for quantitative biology. Here, the authors report kinetic parameters, noise scaling, impact on growth, and the fundamental leakiness of a wide range of inducible transcriptional systems, including a new, highly light sensitive LOV-transcription factor.
- Vojislav Gligorovski
- , Ahmad Sadeghi
- & Sahand Jamal Rahi
-
Article
| Open Access Predictive and robust gene selection for spatial transcriptomics
Predictive and robust gene selection for spatial transcriptomics
Gene selection for spatial transcriptomics is currently not optimal. Here the authors report PERSIST, a flexible deep learning framework that uses existing scRNA-seq data to identify gene targets for spatial transcriptomics; they show this allows you to capture more information with fewer genes.
- Ian Covert
- , Rohan Gala
- & Su-In Lee
-
Article
| Open Access A comprehensive benchmarking with practical guidelines for cellular deconvolution of spatial transcriptomics
A comprehensive benchmarking with practical guidelines for cellular deconvolution of spatial transcriptomics
This study comprehensively benchmarks 18 state-of-the-art methods for cellular deconvolution of spatial transcriptomics and provide decision-tree-style guidelines and recommendations for method selection.
- Haoyang Li
- , Juexiao Zhou
- & Xin Gao
-
Article
| Open Access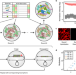 Spatial profiling of microbial communities by sequential FISH with error-robust encoding
Spatial profiling of microbial communities by sequential FISH with error-robust encoding
Spatial analysis of microbiomes at single cell resolution is challenging. Here the authors report a highly multiplexed method for spatial profiling, sequential error-robust fluorescence in situ hybridisation (SEER-FISH), and show that this allows mapping of microbial communities at micron-scale.
- Zhaohui Cao
- , Wenlong Zuo
- & Lei Dai
-
Article
| Open Access Batch alignment of single-cell transcriptomics data using deep metric learning
Batch alignment of single-cell transcriptomics data using deep metric learning
The increasing scale of single-cell RNA-seq studies presents new challenge for integrating datasets from different batches. Here, the authors develop scDML, a tool that simultaneously removes batch effects, improves clustering performance, recovers true cell types, and scales well to large datasets.
- Xiaokang Yu
- , Xinyi Xu
- & Xiangjie Li
-
Article
| Open Access Highly efficient and robust π-FISH rainbow for multiplexed in situ detection of diverse biomolecules
Highly efficient and robust π-FISH rainbow for multiplexed in situ detection of diverse biomolecules
Fluorescence in situ hybridization (FISH) methods with high sensitivity are needed. Here the authors develop multiplex πFISH rainbow to detect a range of biomolecules; they also combine this with the hybridization chain reaction to develop πFISH+ technology for short nucleic acid fragment detection.
- Yingfeng Tao
- , Xiaoliu Zhou
- & Gang Cao
-
Article
| Open Access Harnessing gut microbes for glycan detection and quantification
Harnessing gut microbes for glycan detection and quantification
Detecting distinct glycans within heterogeneous mixtures is hindered by glycan structural complexity and diversity. Here the authors exploit the ability of gut microbes to sense different glycan structures in order to develop quantitative glycan biosensors by coupling bacterial detection machinery to an optimised luciferase reporter.
- Jennifer L. Modesto
- , Victoria H. Pearce
- & Guy E. Townsend II
-
Article
| Open Access Multi-omics identify falling LRRC15 as a COVID-19 severity marker and persistent pro-thrombotic signals in convalescence
Multi-omics identify falling LRRC15 as a COVID-19 severity marker and persistent pro-thrombotic signals in convalescence
End-stage kidney disease confers a high risk for severe COVID-19 infection. Using an at-risk group (end-stage kidney disease patients with COVID-19), authors using RNA-sequencing of immune cells and plasma proteomic profiling to investigate the host response to viral infection.
- Jack S. Gisby
- , Norzawani B. Buang
- & James E. Peters
-
Article
| Open Access Spatial transcriptomics landscape of lesions from non-communicable inflammatory skin diseases
Spatial transcriptomics landscape of lesions from non-communicable inflammatory skin diseases
Inflammatory skin diseases involve various different immune cells in a localised area. Here the authors use spatial transcriptomics to show that disease relevant cytokine transcripts are sparsely expressed in lesional skin, yet are associated with local amplification cascades that promote skin inflammation.
- A. Schäbitz
- , C. Hillig
- & S. Eyerich
-
Article
| Open Access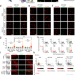 Tagging active neurons by soma-targeted Cal-Light
Tagging active neurons by soma-targeted Cal-Light
Techniques for tagging active neurons with high spatiotemporal precision are limited. Here the authors report soma-targeted CalLight (ST-Cal-Light) which selectively converts somatic calcium rise triggered by action potentials into gene expression, and generate a conditional ST-Cal-Light knock-in mouse.
- Jung Ho Hyun
- , Kenichiro Nagahama
- & Hyung-Bae Kwon
-
Article
| Open Access Graph-based autoencoder integrates spatial transcriptomics with chromatin images and identifies joint biomarkers for Alzheimer’s disease
Graph-based autoencoder integrates spatial transcriptomics with chromatin images and identifies joint biomarkers for Alzheimer’s disease
Methods for jointly analysing the different spatial data modalities in 3D are lacking. Here the authors report the computational framework STACI (Spatial Transcriptomic data using over-parameterized graph-based Autoencoders with Chromatin Imaging data) which they apply to an Alzheimer’s disease mouse model.
- Xinyi Zhang
- , Xiao Wang
- & Caroline Uhler
-
Article
| Open Access Learning the histone codes with large genomic windows and three-dimensional chromatin interactions using transformer
Learning the histone codes with large genomic windows and three-dimensional chromatin interactions using transformer
Existing deep learning-based approaches for the prediction of gene expression by histone modifications (HMs) can only focus on narrow and linear genomic regions around promoters. Here, the authors address these problems by developing a transformer-based deep learning architecture named Chromoformer.
- Dohoon Lee
- , Jeewon Yang
- & Sun Kim
-
Article
| Open Access Absolute protein quantification using fluorescence measurements with FPCountR
Absolute protein quantification using fluorescence measurements with FPCountR
A challenge in synthetic biology is the empirical characterisation of genetic parts. Here the authors present FPCountR, a validated method and accompanying R package that enables the precise quantification of fluorescent protein reporters per bacterial cell to be enumerated in ‘proteins per cell’ or nanomolar units without requiring protein purification.
- Eszter Csibra
- & Guy-Bart Stan
-
Article
| Open Access Decoding the olfactory map through targeted transcriptomics links murine olfactory receptors to glomeruli
Decoding the olfactory map through targeted transcriptomics links murine olfactory receptors to glomeruli
Targeted spatial transcriptomics mapped olfactory receptor mRNAs to sections of the murine olfactory bulb to generate an interactive, statistical, 3D model of glomeruli locations and identify an ultra-sensitive receptor-odorant relationship.
- Kevin W. Zhu
- , Shawn D. Burton
- & Hiroaki Matsunami
-
Article
| Open Access Cell landscape of larval and adult Xenopus laevis at single-cell resolution
Cell landscape of larval and adult Xenopus laevis at single-cell resolution
Single-cell RNA sequencing technology offers a unique opportunity to dissect cell heterogeneity of animals. Here, the authors construct a Xenopus cell landscape including larval and adult organs to dissect cell heterogeneity of the amphibian.
- Yuan Liao
- , Lifeng Ma
- & Xiaoping Han
-
Article
| Open Access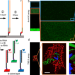 Simple synthesis of massively parallel RNA microarrays via enzymatic conversion from DNA microarrays
Simple synthesis of massively parallel RNA microarrays via enzymatic conversion from DNA microarrays
RNA microarrays have many potential applications, but are difficult to produce. Here, the AUs present a method for converting commercial, customizable DNA microarrays into RNA microarrays using an accessible three-step process involving primer photocrosslinking, extension, and template degradation.
- Erika Schaudy
- , Kathrin Hölz
- & Mark M. Somoza
-
Article
| Open Access RIViT-seq enables systematic identification of regulons of transcriptional machineries
RIViT-seq enables systematic identification of regulons of transcriptional machineries
Here the authors present their method ‘regulon identification by in vitro transcription-sequencing’ (RIViT-seq), which enables systematic identification of target genes of transcription factors of interest. They applied RIViT-seq to 13 sigma factors from Streptomyces coelicolor A3(2) and successfully identified target genes of 11 of these, expanding the regulatory characterisation in this organism.
- Hiroshi Otani
- & Nigel J. Mouncey
-
Article
| Open Access Context-specific effects of sequence elements on subcellular localization of linear and circular RNAs
Context-specific effects of sequence elements on subcellular localization of linear and circular RNAs
Ron and Ulitsky found using massively parallel assays that the effects of short RNA sequences on the subcellular localization of their host RNAs are strongly dependent on the host RNA form, linear or circular, and spliced or unspliced.
- Maya Ron
- & Igor Ulitsky
-
Article
| Open Access The spatial transcriptomic landscape of the healing mouse intestine following damage
The spatial transcriptomic landscape of the healing mouse intestine following damage
The colon is comprised of specialized cells that interact with each other to function, however, the molecular regionalization of the colon is incompletely understood. Here, the authors use spatial transcriptomics to generate a publicly available resource defining the transcriptomic regionalization of the colon during steady state and mucosal healing.
- Sara M. Parigi
- , Ludvig Larsson
- & Eduardo J. Villablanca
-
Article
| Open Access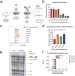 A disease-linked lncRNA mutation in RNase MRP inhibits ribosome synthesis
A disease-linked lncRNA mutation in RNase MRP inhibits ribosome synthesis
Mutations in the non-coding RNA RMRP cause primary immunodeficiency. Robertson et al show that a disease-associated mutation in RMRP impairs pre-ribosomal RNA processing and reduces ribosome abundance, establishing this disorder as a ribosomopathy.
- Nic Robertson
- , Vadim Shchepachev
- & David Tollervey
-
Article
| Open Access Integration of single-cell transcriptomes and chromatin landscapes reveals regulatory programs driving pharyngeal organ development
Integration of single-cell transcriptomes and chromatin landscapes reveals regulatory programs driving pharyngeal organ development
The molecular basis and gene regulatory networks driving pharyngeal endoderm development remain poorly understood. Here the authors report single cell transcriptomic and chromatin landscapes to delineate regulatory programs driving this process and to define the immunodeficiency-associated developmental defects resulting from Foxn1 dysfunction.
- Margaret E. Magaletta
- , Macrina Lobo
- & René Maehr
-
Article
| Open Access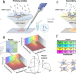 Spatial transcriptomics using combinatorial fluorescence spectral and lifetime encoding, imaging and analysis
Spatial transcriptomics using combinatorial fluorescence spectral and lifetime encoding, imaging and analysis
Spatial-omics methods with ease-of-use and high multiplexing are in demand. Here the authors report Multi Omic Single-scan Assay with Integrated Combinatorial Analysis (MOSAICA) which uses Spectral and Fluorescence Lifetime Imaging and Microscopy; they apply this to co-detection of mRNA and protein.
- Tam Vu
- , Alexander Vallmitjana
- & Weian Zhao
-
Article
| Open Access Cellular variability of nonsense-mediated mRNA decay
Cellular variability of nonsense-mediated mRNA decay
Here the author developed a single-cell reporter system to identify cell-to-cell variability of nonsense-mediated mRNA decay (NMD). This approach provides a sensitive tool to investigate cellular heterogeneity of NMD in various physiological conditions.
- Hanae Sato
- & Robert H. Singer
-
Article
| Open Access The NUCKS1-SKP2-p21/p27 axis controls S phase entry
The NUCKS1-SKP2-p21/p27 axis controls S phase entry
Entry into S phase of the cell cycle is regulated positively by mitogens and negatively by DNA damage; however, how balance of these signals is achieved is not well known. Here the authors show that the NUCKS1-SKP2- p21/p27 axis integrates this information, where the NUCKS1 transcription factor affects levels of p21/p27 to readout the mitogen:DNA damage balance and regulate S phase entry decision.
- Samuel Hume
- , Claudia P. Grou
- & Grigory L. Dianov
-
Article
| Open Access Histone H4 lysine 16 acetylation controls central carbon metabolism and diet-induced obesity in mice
Histone H4 lysine 16 acetylation controls central carbon metabolism and diet-induced obesity in mice
Misregulation of chromatin has been linked to many conditions, including obesity, but the details remain unclear. Here the authors identify the H4 lysine 16 acetyltransferase MOF as a master regulator of glucose metabolism that is required for normal glucose uptake and fat storage.
- Cecilia Pessoa Rodrigues
- , Aindrila Chatterjee
- & Asifa Akhtar
-
Article
| Open Access Standardized preservation, extraction and quantification techniques for detection of fecal SARS-CoV-2 RNA
Standardized preservation, extraction and quantification techniques for detection of fecal SARS-CoV-2 RNA
While the analysis of SARS-CoV-2 RNA in stool samples has led to important insights regarding the disease, quantification is currently challenging. Here the authors use patient samples to benchmark preservation, extraction and quantification methods to optimise detection of viral RNA.
- Aravind Natarajan
- , Alvin Han
- & Ami S. Bhatt
-
Article
| Open Access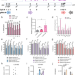 Unraveling the functional role of DNA demethylation at specific promoters by targeted steric blockage of DNA methyltransferase with CRISPR/dCas9
Unraveling the functional role of DNA demethylation at specific promoters by targeted steric blockage of DNA methyltransferase with CRISPR/dCas9
The causal relationship between DNA demethylation and gene expression regulation has not yet been fully resolved. Here the authors develop a nuclease-dead Cas9 (dCas9) and gRNA site-specific targeting approach to physically block DNA methylation at specific promoters to cause DNA demethylation in cells and tackle this question.
- Daniel M. Sapozhnikov
- & Moshe Szyf
-
Article
| Open Access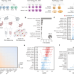 Confronting false discoveries in single-cell differential expression
Confronting false discoveries in single-cell differential expression
Differential expression analysis of single-cell transcriptomics allows scientists to dissect cell-type-specific responses to biological perturbations. Here, the authors show that many commonly used methods are biased and can produce false discoveries.
- Jordan W. Squair
- , Matthieu Gautier
- & Grégoire Courtine
-
Article
| Open Access LineageOT is a unified framework for lineage tracing and trajectory inference
LineageOT is a unified framework for lineage tracing and trajectory inference
Lineage tracing and snapshots of transcriptional state at the single-cell level are powerful, complementary tools for studying development. Here, the authors propose a mathematical method combining lineage tracing with trajectory inference to improve our understanding of development.
- Aden Forrow
- & Geoffrey Schiebinger
-
Article
| Open Access High-depth spatial transcriptome analysis by photo-isolation chemistry
High-depth spatial transcriptome analysis by photo-isolation chemistry
Spatial analysis of RNAseq data is important. Here the authors report a method for transcriptome profiling combined with photo-isolation chemistry to allow determination of expression profiles specifically from photo-irradiated regions of interest which they use in mouse brains and embryonic tissues.
- Mizuki Honda
- , Shinya Oki
- & Yasuyuki Ohkawa
-
Article
| Open Access Single-nucleus RNA-seq2 reveals functional crosstalk between liver zonation and ploidy
Single-nucleus RNA-seq2 reveals functional crosstalk between liver zonation and ploidy
Single-cell RNA-seq reveals the cellular heterogeneity in development and disease. Here the authors present a single-nucleus RNA-seq2 that allows deep characterization of nuclei isolated from frozen archived tissues, apply it for transcriptional profiling of individual hepatocytes, and determine a functional crosstalk between liver zonation and ploidy.
- M. L. Richter
- , I. K. Deligiannis
- & C. P. Martinez-Jimenez
-
Article
| Open Access High-throughput 5′ UTR engineering for enhanced protein production in non-viral gene therapies
High-throughput 5′ UTR engineering for enhanced protein production in non-viral gene therapies
The engineering of 5′ UTRs that modulate protein expression remains a great challenge. Here we leverage synthetic biology and computational design to develop a high-throughput strategy to design, screen, and optimize 5′ UTRs that enhance protein expression for non-viral gene therapies.
- Jicong Cao
- , Eva Maria Novoa
- & Timothy K. Lu
-
Article
| Open Access Engineering an anti-HER2 biparatopic antibody with a multimodal mechanism of action
Engineering an anti-HER2 biparatopic antibody with a multimodal mechanism of action
HER2 acts an oncogenic driver in numerous cancers. Here, the authors design an anti-HER2 biparatopic and tetravalent IgG fusion with inhibitory effects in a xenograft model.
- Florian Kast
- , Martin Schwill
- & Andreas Plückthun
-
Article
| Open Access Cell segmentation-free inference of cell types from in situ transcriptomics data
Cell segmentation-free inference of cell types from in situ transcriptomics data
Inaccurate cell segmentation has been the major problem for cell-type identification and tissue characterization of the in situ spatially resolved transcriptomics data. Here we show a robust cell segmentation-free computational framework (SSAM), for identifying cell types and tissue domains in 2D and 3D.
- Jeongbin Park
- , Wonyl Choi
- & Naveed Ishaque
-
Article
| Open Access Mapping the molecular and structural specialization of the skin basement membrane for inter-tissue interactions
Mapping the molecular and structural specialization of the skin basement membrane for inter-tissue interactions
The basement membrane is located at tissue interfaces, but how it mediates distinct inter-tissue interactions is unclear. Here, the authors systematically define the spatial heterogeneity of skin basement membrane composition and show its functional importance in inter-tissue interactions.
- Ko Tsutsui
- , Hiroki Machida
- & Hironobu Fujiwara
-
Article
| Open Access Osteocyte transcriptome mapping identifies a molecular landscape controlling skeletal homeostasis and susceptibility to skeletal disease
Osteocyte transcriptome mapping identifies a molecular landscape controlling skeletal homeostasis and susceptibility to skeletal disease
Osteocytes are the master regulatory cells within the skeleton. Here, the authors map the transcriptome of osteocytes from diverse skeletal sites, ages and between sexes and identify an osteocyte transcriptome signature associated with rare skeletal disorders and common complex skeletal diseases.
- Scott E. Youlten
- , John P. Kemp
- & Peter I. Croucher
-
Article
| Open Access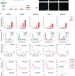 Melanoma subpopulations that rapidly escape MAPK pathway inhibition incur DNA damage and rely on stress signalling
Melanoma subpopulations that rapidly escape MAPK pathway inhibition incur DNA damage and rely on stress signalling
BRAF inhibitors are used to treat late-stage melanoma patients harbouring BRAF mutations. Here the authors track the responses of single melanoma cells to BRAF inhibitors and show that a subset of cells rapidly escapes drug via non-genetic mechanisms and incurs DNA damage.
- Chen Yang
- , Chengzhe Tian
- & Sabrina L. Spencer
-
Article
| Open Access LETR1 is a lymphatic endothelial-specific lncRNA governing cell proliferation and migration through KLF4 and SEMA3C
LETR1 is a lymphatic endothelial-specific lncRNA governing cell proliferation and migration through KLF4 and SEMA3C
Long noncoding RNAs regulate tissue-specific gene expression. Here the authors profile lineage-specific lncRNAs in human dermal lymphatic and blood vascular endothelial cells (LECs and BECs) and show a role of LEC-specific lncRNA, LETR1, in cell proliferation and migration.
- Luca Ducoli
- , Saumya Agrawal
- & Michael Detmar

 TM4SF19-mediated control of lysosomal activity in macrophages contributes to obesity-induced inflammation and metabolic dysfunction
TM4SF19-mediated control of lysosomal activity in macrophages contributes to obesity-induced inflammation and metabolic dysfunction