Article
|
Open Access
Featured
-
-
Article
| Open Access Angle between DNA linker and nucleosome core particle regulates array compaction revealed by individual-particle cryo-electron tomography
Angle between DNA linker and nucleosome core particle regulates array compaction revealed by individual-particle cryo-electron tomography
Here, using cryo-ET, the 3D structures of individual nucleosome particles were characterized to observe changes under varying ionic strengths and in the presence of protein H1, revealing key regulatory roles in chromatin organization dynamics.
- Meng Zhang
- , César Díaz-Celis
- & Gang Ren
-
Article
| Open Access Enzyme-assisted high throughput sequencing of an expanded genetic alphabet at single base resolution
Enzyme-assisted high throughput sequencing of an expanded genetic alphabet at single base resolution
The expansion of the genetic code with synthetic nucleotides has broadened our ability to evolve DNA as a functional material, but we lack analytical tools for the expanded alphabet. Here the authors demonstrate an enzyme-assisted method for the sequencing of six-letter DNA.
- Bang Wang
- , Kevin M. Bradley
- & Steven A. Benner
-
Article
| Open Access Development of Aptamer-DNAzyme based metal-nucleic acid frameworks for gastric cancer therapy
Development of Aptamer-DNAzyme based metal-nucleic acid frameworks for gastric cancer therapy
The development of metal-nucleic acid nanocomposites for therapeutic is limited by poor stability and synthesis efficiency. Here, the authors develop a multi-fragmented aptamer DNAzyme metal-nucleic acid frameworks (MNFs) under milder conditions and demonstrate its preclinical efficacy in gastric cancer.
- Jiaqi Yan
- , Rajendra Bhadane
- & Hongbo Zhang
-
Article
| Open Access Nucleoside Phosphorylases make N7-xanthosine
Nucleoside Phosphorylases make N7-xanthosine
Nucleoside-processing enzymes exhibit strict regioselectivity for glycosylation of purine nucleobases at N9. Here, the authors report an exception and show that wild type nucleoside phosphorylases also furnish N7-xanthosine, a non-native ribosylation regioisomer of xanthosine.
- Sarah Westarp
- , Felix Brandt
- & Felix Kaspar
-
Article
| Open Access Mechanism of DNA unwinding by MCM8-9 in complex with HROB
Mechanism of DNA unwinding by MCM8-9 in complex with HROB
MCM8-9 and HROB function together in DNA damage response. Here, the authors describe the mechanism of DNA unwinding by MCM8-9 and its activation by HROB. HROB makes direct contacts with both MCM8 and MCM9 and promotes DNA unwinding downstream of MCM8-9 loading and hexameric ring formation on DNA.
- Ananya Acharya
- , Hélène Bret
- & Petr Cejka
-
Article
| Open Access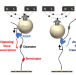 Reciprocating RNA Polymerase batters through roadblocks
Reciprocating RNA Polymerase batters through roadblocks
During transcription, RNA polymerases may encounter protein roadblocks along template DNA. Here, Qian et al. use magnetic tweezers to show that RNA polymerases can backtrack and ram into longer lived roadblocks to transit through them.
- Jin Qian
- , Allison Cartee
- & Laura Finzi
-
Article
| Open Access Chemical unclonable functions based on operable random DNA pools
Chemical unclonable functions based on operable random DNA pools
Physical unclonable functions provide algorithm-independent cryptography based on non-distributable unique tokens. Here, the authors introduce unclonable functions based on random DNA pools, enabling secure decentralized authentication.
- Anne M. Luescher
- , Andreas L. Gimpel
- & Robert N. Grass
-
Article
| Open Access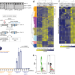 Dbf4-dependent kinase promotes cell cycle controlled resection of DNA double-strand breaks and repair by homologous recombination
Dbf4-dependent kinase promotes cell cycle controlled resection of DNA double-strand breaks and repair by homologous recombination
The repair of DNA double strand breaks is strictly controlled during the cell cycle by the CDK kinase. Here the authors identify the DDK kinase as a second major regulator for this cell cycle regulation and elucidate its functional targets.
- Lorenzo Galanti
- , Martina Peritore
- & Boris Pfander
-
Article
| Open Access Correlating fluorescence microscopy, optical and magnetic tweezers to study single chiral biopolymers such as DNA
Correlating fluorescence microscopy, optical and magnetic tweezers to study single chiral biopolymers such as DNA
It is hard to correlate force, torque and localization information. The authors report Combined Optical and Magnetic BIomolecule TWEEZers, COMBI-Tweez, that integrates optical trapping, time-resolved electromagnetic tweezers, and fluorescence microscopy: they demonstrate visualisation of higher order structural motifs in DNA.
- Jack W. Shepherd
- , Sebastien Guilbaud
- & Mark C. Leake
-
Article
| Open Access FUS unveiled in mitochondrial DNA repair and targeted ligase-1 expression rescues repair-defects in FUS-linked motor neuron disease
FUS unveiled in mitochondrial DNA repair and targeted ligase-1 expression rescues repair-defects in FUS-linked motor neuron disease
Dysfunction of Fused in Sarcoma (FUS) leads to increased mitochondrial DNA (mtDNA) damage, mutations, and mitochondrial dysfunction. This study shows that FUS collaborates with mtDNA Ligase IIIα to maintain mtDNA repair and integrity.
- Manohar Kodavati
- , Haibo Wang
- & Muralidhar L. Hegde
-
Article
| Open Access Topological barrier to Cas12a activation by circular DNA nanostructures facilitates autocatalysis and transforms DNA/RNA sensing
Topological barrier to Cas12a activation by circular DNA nanostructures facilitates autocatalysis and transforms DNA/RNA sensing
The authors find that small circular DNA nanostructures which partially match gRNA sequences only minimally activate Cas12a. They report AutoCAR (Autocatalytic Cas12a Circular DNA Amplification Reaction) which allows a single nucleic acid target to activate multiple ribonucleoproteins, and increases reporter cleavage rates.
- Fei Deng
- , Yi Li
- & Ewa M. Goldys
-
Article
| Open Access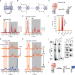 In-cell NMR suggests that DNA i-motif levels are strongly depleted in living human cells
In-cell NMR suggests that DNA i-motif levels are strongly depleted in living human cells
I-Motifs (iM) are non-canonical DNA structures potentially forming in the accessible, single stranded, cytosine-rich genomic region, but the specific contributions of several factors involved in their formation are unknown. Using in-cell NMR, the authors examined DNA i-motif formation in human cells at body temperature, suggesting i-M occur in a small portion (<1%) of genomic sites predisposed to its formation.
- Pavlína Víšková
- , Eva Ištvánková
- & Lukáš Trantírek
-
Article
| Open Access Fluorogenic CRISPR for genomic DNA imaging
Fluorogenic CRISPR for genomic DNA imaging
Conventional CRISPR-based approaches to monitor genomic loci can be hampered by high background and nonspecific nucleolar signal. Here, the authors propose a fluorogenic CRISPR (fCRISPR) tool that allows for high-contrast and sensitive imaging of genomic DNA.
- Zhongxuan Zhang
- , Xiaoxiao Rong
- & Xing Li
-
Article
| Open Access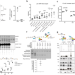 Covalent PARylation of DNA base excision repair proteins regulates DNA demethylation
Covalent PARylation of DNA base excision repair proteins regulates DNA demethylation
The PARylation activity of PARP recruits DNA repair proteins to damaged DNA, most likely via non-covalent protein-PAR interactions. Here, the authors show that PARP1 covalently PARylates base excision repair proteins to modulate their DNA transactions and thus promote active BER DNA demethylation.
- Simon D. Schwarz
- , Jianming Xu
- & Roland Steinacher
-
Article
| Open Access Structural basis for DNA proofreading
Structural basis for DNA proofreading
Here, the authors use cryo-EM to capture nine intermediates along the DNA proofreading pathway using human mitochondrial DNA Polymerase Gamma. The results provide a step-by-step view of the DNA proofreading at single-nucleotide resolution.
- Gina Buchel
- , Ashok R. Nayak
- & Dmitry Temiakov
-
Article
| Open Access The bacterial replication origin BUS promotes nucleobase capture
The bacterial replication origin BUS promotes nucleobase capture
Here, the authors show that a DnaA:origin complex promotes specific nucleobase capture from a single DNA strand. It is proposed that this mechanism may play a key role stimulating opening of bacterial chromosome origins.
- Simone Pelliciari
- , Salomé Bodet-Lefèvre
- & Heath Murray
-
Article
| Open Access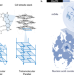 The ALS/FTD-related C9orf72 hexanucleotide repeat expansion forms RNA condensates through multimolecular G-quadruplexes
The ALS/FTD-related C9orf72 hexanucleotide repeat expansion forms RNA condensates through multimolecular G-quadruplexes
A common genetic cause of Amyotrophic lateral sclerosis (ALS) and frontotemporal dementia (FTD) is expansion of the intronic hexanucleotide repeat (GGGGCC)n in C9orf72. Here the authors reveal that the RNA (GGGGCC)n expansion repeat associated with ALS/FTD can generate condensates in the absence of proteins, highlighting the potential relevance of targeting RNA-structures to treat neurodegenerative diseases.
- Federica Raguseo
- , Yiran Wang
- & Marco Di Antonio
-
Article
| Open Access A unified Watson-Crick geometry drives transcription of six-letter expanded DNA alphabets by E. coli RNA polymerase
A unified Watson-Crick geometry drives transcription of six-letter expanded DNA alphabets by E. coli RNA polymerase
Here the authors present the structural mechanism of recognition of unnatural nucleobases in a six-letter expanded genetic system by E. coli RNA polymerase, and provide structural evidence for tautomerization during transcription.
- Juntaek Oh
- , Zelin Shan
- & Dong Wang
-
Article
| Open Access Structures of the DarR transcription regulator reveal unique modes of second messenger and DNA binding
Structures of the DarR transcription regulator reveal unique modes of second messenger and DNA binding
The molecular basis for 2nd messenger binding to the TetR family regulator (TFR), DarR, is unknown. Here the authors obtain DarR structures bound to adenine 2nd messengers and DNA, revealing both a new TFR allosteric binding site and DNA binding mode.
- Maria A. Schumacher
- , Nicholas Lent
- & Raul Salinas
-
Article
| Open Access Mechanism of substrate hydrolysis by the human nucleotide pool sanitiser DNPH1
Mechanism of substrate hydrolysis by the human nucleotide pool sanitiser DNPH1
Inactivation of DNPH1 leads to hmdU incorporation into DNA, sensitising BRCA-deficient cells to PARP inhibitors. Crystal structures of DNPH1 bound to hmdU monophosphate reveal a two-step mechanism for hydrolysis via a glycosyl-enzyme intermediate.
- Neil J. Rzechorzek
- , Simone Kunzelmann
- & Stephen C. West
-
Article
| Open Access Enzymatic synthesis and nanopore sequencing of 12-letter supernumerary DNA
Enzymatic synthesis and nanopore sequencing of 12-letter supernumerary DNA
Unnatural base pairing xenonucleic acids (XNAs) can be used to expand life’s alphabet beyond ATGC. Here, authors show strategies for enzymatic synthesis and next-generation nanopore sequencing of XNA base pairs for reading and writing 12-letter DNA (ATGCBSPZXKJV).
- Hinako Kawabe
- , Christopher A. Thomas
- & Jorge A. Marchand
-
Article
| Open Access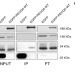 Recognition and coacervation of G-quadruplexes by a multifunctional disordered region in RECQ4 helicase
Recognition and coacervation of G-quadruplexes by a multifunctional disordered region in RECQ4 helicase
In this work, the authors report a three-step charge-driven coacervation model involving dynamic complexes between a positively charged IDR of human RECQ4 and G-quadruplexes. The IDR also interacts with Replication Protein A, implying RECQ4’s regulatory role.
- Anna C. Papageorgiou
- , Michaela Pospisilova
- & Konstantinos Tripsianes
-
Article
| Open Access Structure-guided inhibition of the cancer DNA-mutating enzyme APOBEC3A
Structure-guided inhibition of the cancer DNA-mutating enzyme APOBEC3A
APOBEC3A mutates its host DNA in human cancers to evolve drug resistance. Modified-DNA inhibitors suppress this mutagenic activity in cells, suggesting use as conjuvants in anti-cancer therapies. Here the authors reveal structural insights into how these inhibitors bind APOBEC3A.
- Stefan Harjes
- , Harikrishnan M. Kurup
- & Geoffrey B. Jameson
-
Article
| Open Access Yeast Rad52 is a homodecamer and possesses BRCA2-like bipartite Rad51 binding modes
Yeast Rad52 is a homodecamer and possesses BRCA2-like bipartite Rad51 binding modes
Mediator proteins such as BRCA2 and Rad52 direct formation of Rad51 filaments in Homologous Recombination. Here, the authors present cryoEM structures of Saccharomyces cerevisiae Rad52 revealing a homodecamer and Rad51 binding to two regions in Rad52.
- Jaigeeth Deveryshetty
- , Rahul Chadda
- & Edwin Antony
-
Article
| Open Access A digital twin for DNA data storage based on comprehensive quantification of errors and biases
A digital twin for DNA data storage based on comprehensive quantification of errors and biases
Archiving data in synthetic DNA offers unprecedented storage density and longevity. To understand how experimental choices affect the integrity of digital data stored in DNA, the authors study the evolution of errors and bias and with a digital twin they supply tools for experimental planning and design of error-correcing codes.
- Andreas L. Gimpel
- , Wendelin J. Stark
- & Robert N. Grass
-
Article
| Open Access Synergism between CMG helicase and leading strand DNA polymerase at replication fork
Synergism between CMG helicase and leading strand DNA polymerase at replication fork
Coordinating the activities of replicative helicase and DNA polymerase(s) is crucial for DNA replication. Here, the authors show that DNA translocation by CMG is synchronized with the activities of Polε in the leading-strand replisome.
- Zhichun Xu
- , Jianrong Feng
- & Yuanliang Zhai
-
Article
| Open Access Nongenetic surface engineering of mesenchymal stromal cells with polyvalent antibodies to enhance targeting efficiency
Nongenetic surface engineering of mesenchymal stromal cells with polyvalent antibodies to enhance targeting efficiency
Limited delivery of therapeutic cells to diseased tissue hampers the effective application of mesenchymal stromal cells (MSCs). Here, authors modify the cell surface with polyvalent antibodies using DNA-templated assembly, and show that polyvalent interactions can be used to improve the targeting efficiency of MSCs.
- Tenghui Ye
- , Xi Liu
- & Peng Shi
-
Article
| Open Access Metal-mediated DNA strand displacement and molecular device operations based on base-pair switching of 5-hydroxyuracil nucleobases
Metal-mediated DNA strand displacement and molecular device operations based on base-pair switching of 5-hydroxyuracil nucleobases
The development of dynamic DNA nanodevices, whose configuration and function are regulated by specific chemical inputs, represents a rapidly growing area in molecular science. Herein, the authors report the concept of metal-mediated base-pair switching to induce inter- and intramolecular DNA strand displacement in a metal-responsive manner.
- Yusuke Takezawa
- , Keita Mori
- & Mitsuhiko Shionoya
-
Article
| Open Access DNA framework-engineered chimeras platform enables selectively targeted protein degradation
DNA framework-engineered chimeras platform enables selectively targeted protein degradation
The lack of a universal platform for PROTAC development remains a major bottleneck. Here, the authors report modular DNA framework-based PROTACs (DbTACs) that enable precise control of the linker length and selective degradation of diverse targets in different cellular compartments using various warheads.
- Li Zhou
- , Bin Yu
- & Yi Ma
-
Article
| Open Access Enabling programmable dynamic DNA chemistry using small-molecule DNA binders
Enabling programmable dynamic DNA chemistry using small-molecule DNA binders
The binding of small molecules to the double stranded DNA may significantly alter its stability and functionality, which is the basis for many therapeutic and sensing applications. Here, the authors report that DNA binders can be used to program reaction pathways of a dynamic DNA reaction, where DNA strand displacement can be tuned quantitatively according to the affinity, charge, and concentrations of a given DNA binder.
- Junpeng Xu
- , Guan Alex Wang
- & Feng Li
-
Article
| Open Access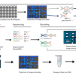 A biological camera that captures and stores images directly into DNA
A biological camera that captures and stores images directly into DNA
DNA data storage has gained recent interest due to the high information density of DNA. Here, the authors have developed a method to directly capture information in the form of light and encode it into DNA via bacteria, analogous to a digital camera.
- Cheng Kai Lim
- , Jing Wui Yeoh
- & Chueh Loo Poh
-
Article
| Open Access Structure and dynamics of an archetypal DNA nanoarchitecture revealed via cryo-EM and molecular dynamics simulations
Structure and dynamics of an archetypal DNA nanoarchitecture revealed via cryo-EM and molecular dynamics simulations
DNA can be folded into rationally designed, unique, and functional materials. Here the authors analyse an archetypal DNA nanoarchitecture with single particle cryo-electron microscopy and molecular dynamics simulations.
- Katya Ahmad
- , Abid Javed
- & Stefan Howorka
-
Article
| Open Access Molecular mechanisms of Holliday junction branch migration catalyzed by an asymmetric RuvB hexamer
Molecular mechanisms of Holliday junction branch migration catalyzed by an asymmetric RuvB hexamer
During homologous recombination, RuvB drives branch migration of the Holliday junction to facilitate DNA repair. With our cryo-EM structure models we have provided a comprehensive mechanism for branch migration that may be universal for all species.
- Anthony D. Rish
- , Zhangfei Shen
- & Tian-Min Fu
-
Article
| Open Access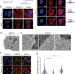 Autophagy receptor NDP52 alters DNA conformation to modulate RNA polymerase II transcription
Autophagy receptor NDP52 alters DNA conformation to modulate RNA polymerase II transcription
An autophagy receptor, NDP52, is recruited to the nucleus where it can bind DNA. The authors show this promotes changes in chromatin accessibility which supports transcription initiation, providing a direct link between autophagy and transcription regulation.
- Ália dos Santos
- , Daniel E. Rollins
- & Christopher P. Toseland
-
Article
| Open Access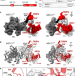 Extended DNA threading through a dual-engine motor module of the activating signal co-integrator 1 complex
Extended DNA threading through a dual-engine motor module of the activating signal co-integrator 1 complex
ASCC3 is a multi-functional helicase that contains two consecutive Ski2-like helicase units. Here, the authors show that ASCC3 can unwind DNA by threading one strand of a substrate duplex through both helicase units, supported by the TRIP4 protein.
- Junqiao Jia
- , Tarek Hilal
- & Markus C. Wahl
-
Article
| Open Access High-throughput single-molecule quantification of individual base stacking energies in nucleic acids
High-throughput single-molecule quantification of individual base stacking energies in nucleic acids
In this work, the authors use a centrifuge force microscope for high-throughput single-molecule experiments to elucidate stacking energies between individual bases of DNA.
- Jibin Abraham Punnoose
- , Kevin J. Thomas
- & Ken Halvorsen
-
Article
| Open Access DNA mechanical flexibility controls DNA potential to activate cGAS-mediated immune surveillance
DNA mechanical flexibility controls DNA potential to activate cGAS-mediated immune surveillance
DNA is well-documented to stimulate immune response. Here the authors show that the activation of cyclic GMP-AMP synthase (cGAS) depends on DNA mechanical flexibility, itself determined by DNA-sequence, damage and length.
- Lina Wang
- , Siru Li
- & Qingkai Yang
-
Article
| Open Access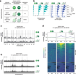 Unscheduled DNA replication in G1 causes genome instability and damage signatures indicative of replication collisions
Unscheduled DNA replication in G1 causes genome instability and damage signatures indicative of replication collisions
Reusswig et al. use engineered systems to force DNA replication in the G1 phase of the cell cycle. This unscheduled G1 replication shows hallmarks of S phase replication, but leads to over-replication and DNA breaks from replication collisions.
- Karl-Uwe Reusswig
- , Julia Bittmann
- & Boris Pfander
-
Article
| Open Access Mechanistic investigation of human maturation of Okazaki fragments reveals slow kinetics
Mechanistic investigation of human maturation of Okazaki fragments reveals slow kinetics
Here, the authors investigate the maturation of Okazaki fragments with human proteins and reveal that initiator RNA removal occurs non-processively and slowly suggesting that additional pathways might exist to accelerate RNA removal in cells.
- Vlad-Stefan Raducanu
- , Muhammad Tehseen
- & Samir M. Hamdan
-
Article
| Open Access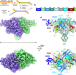 Structural insight into Tn3 family transposition mechanism
Structural insight into Tn3 family transposition mechanism
Here the structure of transposase from Tn3 family is reported in apo form and bound to the transposon ends. The activation of the transposase induces metamorphic refolding of the catalytic domain suggesting the family-specific regulation mechanism.
- Alexander V. Shkumatov
- , Nicolas Aryanpour
- & Rouslan G. Efremov
-
Article
| Open Access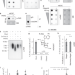 Mammalian N1-adenosine PARylation is a reversible DNA modification
Mammalian N1-adenosine PARylation is a reversible DNA modification
Poly-ADP-ribosylation (PARylation) is a well-known posttranslational modification of proteins. Here the authors show that beyond proteins also mammalian single-stranded DNA is PARylated in vitro and in vivo.
- Michael U. Musheev
- , Lars Schomacher
- & Christof Niehrs
-
Article
| Open Access Structural insight into the bulge-containing KRAS oncogene promoter G-quadruplex bound to berberine and coptisine
Structural insight into the bulge-containing KRAS oncogene promoter G-quadruplex bound to berberine and coptisine
The G-quadruplex formed in KRAS oncogene promoter (KRAS-G4) is a transcriptional modulator and amenable to small molecule targeting. Herein, the authors report the NMR solution structures of a bulge-containing KRAS-G4 that bound to two small molecules. The study provides molecular details of ligand interactions with KRAS-G4 and contributes insight into the design of specific KRAS-G4-interactive drugs.
- Kai-Bo Wang
- , Yushuang Liu
- & Ling-Yi Kong
-
Article
| Open Access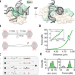 Search and processing of Holliday junctions within long DNA by junction-resolving enzymes
Search and processing of Holliday junctions within long DNA by junction-resolving enzymes
Locating a four-way junction in a high background of genomic DNA is likely to be the rate-limiting step of the resolution process. This study captures the entire reaction trajectory of a nuclease targeting and resolving a DNA junction at single-molecule level.
- Artur P. Kaczmarczyk
- , Anne-Cécile Déclais
- & David S. Rueda
-
Article
| Open Access Structural basis for APE1 processing DNA damage in the nucleosome
Structural basis for APE1 processing DNA damage in the nucleosome
AP endonuclease 1 (APE1) processes genomic AP sites during base excision repair. Here, the authors determine the structural mechanism used by APE1 to process nucleosomal AP sites, providing new insight into DNA repair in chromatin.
- Tyler M. Weaver
- , Nicole M. Hoitsma
- & Bret D. Freudenthal
-
Article
| Open Access Rtt105 regulates RPA function by configurationally stapling the flexible domains
Rtt105 regulates RPA function by configurationally stapling the flexible domains
The single stranded DNA binding protein RPA coordinates DNA metabolism using multiple protein and DNA interaction domains. Here, the authors show that the chaperone-like protein Rtt105 staples RPA domains to prevent untimely protein interactions.
- Sahiti Kuppa
- , Jaigeeth Deveryshetty
- & Edwin Antony
-
Article
| Open Access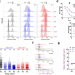 A multi-functional role for the MCM8/9 helicase complex in maintaining fork integrity during replication stress
A multi-functional role for the MCM8/9 helicase complex in maintaining fork integrity during replication stress
The MCM8/9 helicase has been implicated in DNA recombination processes with mutations in these genes causative for infertility and cancer. Here, the authors show that MCM8/9 aids normal fork progression and also stabilizes persistently stalled forks, acting upstream of RAD51 and BRCA1.
- Wezley C. Griffin
- , David R. McKinzey
- & Michael A. Trakselis
-
Article
| Open Access Structure of a nucleosome-bound MuvB transcription factor complex reveals DNA remodelling
Structure of a nucleosome-bound MuvB transcription factor complex reveals DNA remodelling
The MuvB family of protein complexes regulate cell cycle-dependent transcription, and MuvB in complex with the transcription factors B-MYB and FOXM1 activate mitotic genes during G2. Here the authors present cryo-EM data of a MuvB:B-MYB (MMB) complex in the process of remodelling a nucleosome, and define its stoichiometry, assembly, and chromatin binding.
- Marios G. Koliopoulos
- , Reyhan Muhammad
- & Claudio Alfieri
-
Article
| Open Access Efficient DNA fluorescence labeling via base excision trapping
Efficient DNA fluorescence labeling via base excision trapping
Methods for fluorescently labelling DNAs are expensive and labour-intensive. Here the authors report an in situ DNA labelling strategy for oligonucleotides as well as dsDNA that makes use of aldehyde-reactive rotor dyes to trap AP sites resulting from excision of deaminated DNA bases.
- Yong Woong Jun
- , Emily M. Harcourt
- & Eric T. Kool
-
Article
| Open Access Piperazine-derived lipid nanoparticles deliver mRNA to immune cells in vivo
Piperazine-derived lipid nanoparticles deliver mRNA to immune cells in vivo
Next-generation lipid nanoparticles that target non-hepatocytes could be important clinical tools. Using in vivo DNA barcoding, the authors identify piperazine-containing lipids deliver mRNA to immune cells without targeting ligands.
- Huanzhen Ni
- , Marine Z. C. Hatit
- & James E. Dahlman

 Nucleosomal DNA has topological memory
Nucleosomal DNA has topological memory