Featured
-
-
Article
| Open Access Invariant γδTCR natural killer-like effector T cells in the naked mole-rat
Invariant γδTCR natural killer-like effector T cells in the naked mole-rat
Naked mole-rats are long-lived rodents known to be resistant to the development of cancer, yet their immune system remains poorly explored. Here, the authors identify natural killer-like effector γδ T cells that express a dominant γδ T cell receptor and may serve a role in tumour immunosurveillance.
- Guillem Sanchez Sanchez
- , Stephan Emmrich
- & David Vermijlen
-
Article
| Open Access Bioinformatics leading to conveniently accessible, helix enforcing, bicyclic ASX motif mimics (BAMMs)
Bioinformatics leading to conveniently accessible, helix enforcing, bicyclic ASX motif mimics (BAMMs)
Researchers mimic protein interface helices by stapling peptide side chains, or replacing hydrogen bonds with covalent ones, and synthetic helical mimics are heavily biased towards stapling. Here the authors describe bioinformatic discovery of hydrophobic triangles at helix N-termini, and rigid, bicyclic synthetic mimics of them.
- Tianxiong Mi
- , Duyen Nguyen
- & Kevin Burgess
-
Article
| Open Access Enzyme-assisted high throughput sequencing of an expanded genetic alphabet at single base resolution
Enzyme-assisted high throughput sequencing of an expanded genetic alphabet at single base resolution
The expansion of the genetic code with synthetic nucleotides has broadened our ability to evolve DNA as a functional material, but we lack analytical tools for the expanded alphabet. Here the authors demonstrate an enzyme-assisted method for the sequencing of six-letter DNA.
- Bang Wang
- , Kevin M. Bradley
- & Steven A. Benner
-
Article
| Open Access Metabolic phenotyping reveals an emerging role of ammonia abnormality in Alzheimer’s disease
Metabolic phenotyping reveals an emerging role of ammonia abnormality in Alzheimer’s disease
Metabolic implications in AD are unclear. Here, authors found significant correlations between cognitive impairment and metabolic features in a Chinese aging cohort (n = 1397). The study highlights ammonia disturbance as a potential therapeutic target for AD.
- Tianlu Chen
- , Fengfeng Pan
- & Wei Jia
-
Article
| Open Access Teacher-student collaborated multiple instance learning for pan-cancer PDL1 expression prediction from histopathology slides
Teacher-student collaborated multiple instance learning for pan-cancer PDL1 expression prediction from histopathology slides
PDL1 expression is a common biomarker for immunotherapy response in cancer, and it is usually quantified using immunohistochemistry. Here, the authors develop a weakly supervised learning approach combining multiple instance learning and a teacher-student framework to predict PDL1 expression from histopathological imaging.
- Darui Jin
- , Shangying Liang
- & Xiangzhi Bai
-
Article
| Open Access PLMSearch: Protein language model powers accurate and fast sequence search for remote homology
PLMSearch: Protein language model powers accurate and fast sequence search for remote homology
Homologous protein search is one of the most commonly used methods for protein analysis. Here, authors propose PLMSearch, a search method that takes only sequences as input and can search millions of protein pairs in seconds while maintaining sensitivity comparable to SOTA structure search methods.
- Wei Liu
- , Ziye Wang
- & Shanfeng Zhu
-
Article
| Open Access DELVE: feature selection for preserving biological trajectories in single-cell data
DELVE: feature selection for preserving biological trajectories in single-cell data
Characteristic genes or proteins driving continuous biological processes are difficult to uncover from noisy single-cell data. Here, authors present DELVE, an unsupervised feature selection method to identify core molecular features driving cell fate decisions.
- Jolene S. Ranek
- , Wayne Stallaert
- & Jeremy E. Purvis
-
Article
| Open Access InPACT: a computational method for accurate characterization of intronic polyadenylation from RNA sequencing data
InPACT: a computational method for accurate characterization of intronic polyadenylation from RNA sequencing data
Intronic polyadenylation (IPA) can produce transcripts with truncated coding regions and has been implicated in diverse biological processes and diseases. Here, the authors present a computational method for the accurate delineation of IPA events using RNA-sequencing data.
- Xiaochuan Liu
- , Hao Chen
- & Yang Yang
-
Article
| Open Access A universal molecular control for DNA, mRNA and protein expression
A universal molecular control for DNA, mRNA and protein expression
Multi-omics analyses powerfully combine gene expression and translation, however no available controls can be used across these techniques. Here the authors develop pREF, a universal control construct designed for use in DNA, RNA and protein analyses.
- Helen M. Gunter
- , Scott E. Youlten
- & Tim R. Mercer
-
Article
| Open Access Biomimetic nanocluster photoreceptors for adaptative circular polarization vision
Biomimetic nanocluster photoreceptors for adaptative circular polarization vision
All-in-one multi-task photoperception is desirable for artificial vision systems. Wen et al. present wafer-scale high density integration of artificial photoreceptors that combine photoadaptation and circular polarized light vision, enabled by chiral-nanocluster-conjugated molecule heterostructures.
- Wei Wen
- , Guocai Liu
- & Yunqi Liu
-
Article
| Open Access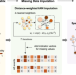 Interrogations of single-cell RNA splicing landscapes with SCASL define new cell identities with physiological relevance
Interrogations of single-cell RNA splicing landscapes with SCASL define new cell identities with physiological relevance
RNA splicing serves as a critical layer of gene expression regulation. Here, authors introduce SCASL for investigating the heterogeneity of RNA splicing landscapes at single-cell resolution, offering a novel scheme for classifying cell identities with physiological relevance.
- Xianke Xiang
- , Yao He
- & Xuerui Yang
-
Article
| Open Access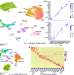 Predicting proximal tubule failed repair drivers through regularized regression analysis of single cell multiomic sequencing
Predicting proximal tubule failed repair drivers through regularized regression analysis of single cell multiomic sequencing
A profibrotic, proinflammatory kidney cell population has been identified as a driver of chronic kidney disease. Here, authors generate a human kidney single cell multiomic dataset and apply a regularised regression approach to identify transcription factors underpinning this cell population.
- Nicolas Ledru
- , Parker C. Wilson
- & Benjamin D. Humphreys
-
Article
| Open Access SiFT: uncovering hidden biological processes by probabilistic filtering of single-cell data
SiFT: uncovering hidden biological processes by probabilistic filtering of single-cell data
Cells simultaneously encode multiple signals, some harder to recover. Here, authors introduce SiFT (Signal FilTering), a kernel-based projection method, revealing underlying biological processes in single-cell data.
- Zoe Piran
- & Mor Nitzan
-
Article
| Open Access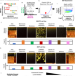 Anti-correlated feature selection prevents false discovery of subpopulations in scRNAseq
Anti-correlated feature selection prevents false discovery of subpopulations in scRNAseq
Typical single-cell RNAseq pipelines will subcluster homogeneous cells. Here, authors present a computational algorithm for accurately identifying cell-type marker genes in single-cell data analysis with a low false discovery rate.
- Scott R. Tyler
- , Daniel Lozano-Ojalvo
- & Eric E. Schadt
-
Article
| Open Access DiffDomain enables identification of structurally reorganized topologically associating domains
DiffDomain enables identification of structurally reorganized topologically associating domains
Topologically associating domains (TADs) are critical structural units in 3D genome organization, and their reorganization between health and disease states is associated with essential genome functions. However, computational methods for identifying reorganized TADs are still in the early stages of development. Here, the authors present an algorithm leveraging random matrix theory to identify reorganized TADs.
- Dunming Hua
- , Ming Gu
- & Dechao Tian
-
Article
| Open Access Direct RNA sequencing coupled with adaptive sampling enriches RNAs of interest in the transcriptome
Direct RNA sequencing coupled with adaptive sampling enriches RNAs of interest in the transcriptome
It can be difficult to find rare transcripts when sequencing a transcriptome. Here the authors show adaptive sampling on direct RNA runs to increase the likelihood of finding less frequent ones while selectively ejecting the higher-abundance transcripts.
- Jiaxu Wang
- , Lin Yang
- & Yue Wan
-
Article
| Open Access Open access repository-scale propagated nearest neighbor suspect spectral library for untargeted metabolomics
Open access repository-scale propagated nearest neighbor suspect spectral library for untargeted metabolomics
Interpreting untargeted mass spectrometry (MS) data is challenging due to incomplete reference libraries. Here, the authors created the nearest neighbor suspect spectral library from largescale public MS data, significantly enhancing the ability to hypothesize structures for unknown mass spectra.
- Wout Bittremieux
- , Nicole E. Avalon
- & Pieter C. Dorrestein
-
Article
| Open Access Single-cell spatial metabolomics with cell-type specific protein profiling for tissue systems biology
Single-cell spatial metabolomics with cell-type specific protein profiling for tissue systems biology
The authors developed a framework for joint protein-metabolite profiling at the single-cell level in human tissue combining targeted multiplexed protein imaging and untargeted spatial metabolomics in a single pipeline.
- Thomas Hu
- , Mayar Allam
- & Ahmet F. Coskun
-
Article
| Open Access GNTD: reconstructing spatial transcriptomes with graph-guided neural tensor decomposition informed by spatial and functional relations
GNTD: reconstructing spatial transcriptomes with graph-guided neural tensor decomposition informed by spatial and functional relations
Reconstructing transcriptome-wide spatially-resolved gene expressions requires modelling nonlinear patterns and spatial structures in RNA profiling data. Here, authors introduce a graph-guided neural hierarchical tensor decomposition model that incorporates spatial and functional relations for the task.
- Tianci Song
- , Charles Broadbent
- & Rui Kuang
-
Comment
| Open Access Is Protein BLAST a thing of the past?
Is Protein BLAST a thing of the past?
Will protein structure search tools like AlphaFold replace protein sequence search with BLAST? We discuss the promises, using structure search for remote homology detection, and why protein BLAST, as the leading sequence search tool, should strive to incorporate structural information
- Ali Al-Fatlawi
- , Martin Menzel
- & Michael Schroeder
-
Article
| Open Access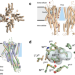 Elucidation of the structural basis for ligand binding and translocation in conserved insect odorant receptor co-receptors
Elucidation of the structural basis for ligand binding and translocation in conserved insect odorant receptor co-receptors
Insects rely on olfaction for behavior control. Recent structural studies of receptors provide insight into ligand binding. Here, the authors identify dynamic binding mechanism to Orco, explaining its high selectivity with insights in compound screening.
- Jody Pacalon
- , Guillaume Audic
- & Jérémie Topin
-
Article
| Open Access Spatial transcriptomics deconvolution at single-cell resolution using Redeconve
Spatial transcriptomics deconvolution at single-cell resolution using Redeconve
Computational deconvolution with single-cell RNA sequencing data as a reference is pivotal for interpreting spatial transcriptomics data. Here, authors present Redeconve, which improves the resolution by more than 100-fold with higher accuracy and speed.
- Zixiang Zhou
- , Yunshan Zhong
- & Xianwen Ren
-
Article
| Open Access Integrating spatial and single-cell transcriptomics data using deep generative models with SpatialScope
Integrating spatial and single-cell transcriptomics data using deep generative models with SpatialScope
Spatial transcriptomics (ST) is transforming tissue analysis but has limitations. Here, authors introduce SpatialScope, an integrated approach combining scRNA-seq and ST data using deep generative models, enabling comprehensive spatial characterisation at transcriptome-wide single-cell resolution.
- Xiaomeng Wan
- , Jiashun Xiao
- & Can Yang
-
Article
| Open Access PhenoSV: interpretable phenotype-aware model for the prioritization of genes affected by structural variants
PhenoSV: interpretable phenotype-aware model for the prioritization of genes affected by structural variants
Here, authors present PhenoSV, a phenotype-aware machine-learning model for the functional interpretation of various types of structural variants (SVs) and genes within or outside SVs, facilitating the extraction of biological insights from coding and noncoding SVs.
- Zhuoran Xu
- , Quan Li
- & Kai Wang
-
Article
| Open Access Improved in situ characterization of protein complex dynamics at scale with thermal proximity co-aggregation
Improved in situ characterization of protein complex dynamics at scale with thermal proximity co-aggregation
Vast majority of cellular activities are carried out by protein complexes that assembled dynamically in response to cellular needs and environmental cues. Here, the authors present Slim-TPCA, an effective and readily deployable strategy to unravel the functional roles of protein complexes en masse across various cellular processes.
- Siyuan Sun
- , Zhenxiang Zheng
- & Chris Soon Heng Tan
-
Article
| Open Access SPACEL: deep learning-based characterization of spatial transcriptome architectures
SPACEL: deep learning-based characterization of spatial transcriptome architectures
Spatial transcriptomics (ST) technologies detect transcript distribution in space. Here, authors present a deep learning based method SPACEL for cell type deconvolution, spatial domain identification and 3D alignment, showcasing it as a valuable toolkit for ST data analysis
- Hao Xu
- , Shuyan Wang
- & Kun Qu
-
Article
| Open Access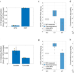 Inferring mitochondrial and cytosolic metabolism by coupling isotope tracing and deconvolution
Inferring mitochondrial and cytosolic metabolism by coupling isotope tracing and deconvolution
Studying metabolism in distinct subcellular compartments typically involves isolating organelles. Here, the authors demonstrate a quantitative approach to infer cytosolic and mitochondrial metabolic activities based on experiments with intact cells, maintaining physiological conditions.
- Alon Stern
- , Mariam Fokra
- & Tomer Shlomi
-
Article
| Open Access Simultaneous selection of nanobodies for accessible epitopes on immune cells in the tumor microenvironment
Simultaneous selection of nanobodies for accessible epitopes on immune cells in the tumor microenvironment
INSPIREseq discovers accessible epitopes in complex environments and leads to nanobodies targeting thousands of epitopes. This in vivo diversity selection identifies selective nanobodies, potentially expediting target discovery and drug development.
- Thillai V. Sekar
- , Eslam A. Elghonaimy
- & Todd A. Aguilera
-
Article
| Open Access Dimension-agnostic and granularity-based spatially variable gene identification using BSP
Dimension-agnostic and granularity-based spatially variable gene identification using BSP
Identifying spatially variable genes (SVGs) is essential for linking molecular cell functions with tissue phenotypes. Here, authors introduce a non-parametric model that detects SVGs from two or three-dimensional spatial transcriptomics data by comparing gene expression patterns at granularities.
- Juexin Wang
- , Jinpu Li
- & Dong Xu
-
Article
| Open Access Spatial-linked alignment tool (SLAT) for aligning heterogenous slices
Spatial-linked alignment tool (SLAT) for aligning heterogenous slices
Spatial omics technologies reveal the organisation of cells in various biological systems. Here, authors propose SLAT, a graph-based algorithm for aligning heterogenous data across technologies, modalities and timepoints, enabling spatiotemporal reconstruction of complex developmental processes.
- Chen-Rui Xia
- , Zhi-Jie Cao
- & Ge Gao
-
Article
| Open Access CamoTSS: analysis of alternative transcription start sites for cellular phenotypes and regulatory patterns from 5' scRNA-seq data
CamoTSS: analysis of alternative transcription start sites for cellular phenotypes and regulatory patterns from 5' scRNA-seq data
Five-prime single-cell RNA-seq, especially the read 1, has precise capture of transcription start sites (TSS), but such information is often overlooked. Here, authors present a computational method suite, CamoTSS, to precisely identify TSS and quantify its expression, enabling effective detection of alternative TSS usage in different biological processes.
- Ruiyan Hou
- , Chung-Chau Hon
- & Yuanhua Huang
-
Article
| Open Access Identification of errors in draft genome assemblies at single-nucleotide resolution for quality assessment and improvement
Identification of errors in draft genome assemblies at single-nucleotide resolution for quality assessment and improvement
A high-quality genome assembly is essential for various genomic studies in life sciences. Here the authors develop CRAQ, a reference-free method that facilitates the evaluation and improvement of any de novo genome assembly with single nucleotide resolution.
- Kunpeng Li
- , Peng Xu
- & Yuannian Jiao
-
Article
| Open Access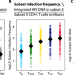 Estimating the contribution of CD4 T cell subset proliferation and differentiation to HIV persistence
Estimating the contribution of CD4 T cell subset proliferation and differentiation to HIV persistence
The authors used mathematical modeling of human data to study how HIV persists despite suppressive antiretroviral therapy. They found that when latently infected CD4+ T cells proliferate or differentiate, they can create HIV DNA and passage it into other subsets. More mature CD4 cell subsets then clear HIV DNA faster.
- Daniel B. Reeves
- , Charline Bacchus-Souffan
- & Peter W. Hunt
-
Article
| Open Access CRUSTY: a versatile web platform for the rapid analysis and visualization of high-dimensional flow cytometry data
CRUSTY: a versatile web platform for the rapid analysis and visualization of high-dimensional flow cytometry data
CRUSTY is an interactive webtool for flow cytometry data analysis, offering popular algorithms and visualizations, and generating publication-quality figures in minutes. It enables users without bioinformatics expertize to mine complex datasets, supports real-time exploration, and is freely available online.
- Simone Puccio
- , Giorgio Grillo
- & Enrico Lugli
-
Comment
| Open Access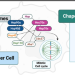 Epichaperomics reveals dysfunctional chaperone protein networks
Epichaperomics reveals dysfunctional chaperone protein networks
Molecular chaperones establish essential protein-protein interaction networks. Modified versions of these assemblies are generally enriched in certain maladies. A study published in Nature Communications used epichaperomics to identify unique changes occurring in chaperone-formed protein networks during mitosis in cancer cells.
- Mark R. Woodford
- , Dimitra Bourboulia
- & Mehdi Mollapour
-
Article
| Open Access A deep learning method for replicate-based analysis of chromosome conformation contacts using Siamese neural networks
A deep learning method for replicate-based analysis of chromosome conformation contacts using Siamese neural networks
Siamese neural networks are a powerful deep learning approach for image analysis. Here, the authors adapt this method to the replicate-based analysis of Hi-C data and find that it successfully discriminates technical noise from biological variation.
- Ediem Al-jibury
- , James W. D. King
- & Daniel Rueckert
-
Article
| Open Access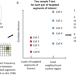 SnapFISH: a computational pipeline to identify chromatin loops from multiplexed DNA FISH data
SnapFISH: a computational pipeline to identify chromatin loops from multiplexed DNA FISH data
Multiplexed DNA FISH technologies are powerful tools to reveal chromatin spatial organisation. Here, the authors developed SnapFISH, a computational pipeline to identify chromatin loops from multiplexed DNA FISH data.
- Lindsay Lee
- , Hongyu Yu
- & Ming Hu
-
Article
| Open Access Cell-type-specific co-expression inference from single cell RNA-sequencing data
Cell-type-specific co-expression inference from single cell RNA-sequencing data
Inferring co-expressions with scRNA-seq data is challenging, and existing methods suffer from inflated false positives and biases. Here, the authors proposed CS-CORE, which yields unbiased estimates and identifies co-expressions that are more reproducible and biologically relevant for scRNA-seq data.
- Chang Su
- , Zichun Xu
- & Jingfei Zhang
-
Article
| Open Access Robust phenotyping of highly multiplexed tissue imaging data using pixel-level clustering
Robust phenotyping of highly multiplexed tissue imaging data using pixel-level clustering
Multiplexed imaging studies are typically focused on cell-level phenotypes. Here, the authors propose Pixie, a cross-platform and open-source pipeline that achieves robust and quantitative annotation of both pixel-level and cell-level features in multiplexed imaging data.
- Candace C. Liu
- , Noah F. Greenwald
- & Michael Angelo
-
Article
| Open Access Atlas-scale single-cell multi-sample multi-condition data integration using scMerge2
Atlas-scale single-cell multi-sample multi-condition data integration using scMerge2
Recent advances in multi-condition single-cell multi-cohort studies enable exploration of diverse cell states. Here, authors present scMerge2, an algorithm that allows integration of a large COVID-19 data collection with over five million cells to uncover distinct signatures of disease progression.
- Yingxin Lin
- , Yue Cao
- & Jean Y. H. Yang
-
Article
| Open Access Multi-batch single-cell comparative atlas construction by deep learning disentanglement
Multi-batch single-cell comparative atlas construction by deep learning disentanglement
Comparing single-cell RNA-seq and ATAC-seq data from multiple batches is challenging due to technical artifacts. Here, the authors propose a method that disentangles technical and biological effects, facilitating batch-confounded chromatin and gene expression state discovery and enhancing the analysis of perturbation effects on cell populations.
- Allen W. Lynch
- , Myles Brown
- & Clifford A. Meyer
-
Article
| Open Access SpatialDM for rapid identification of spatially co-expressed ligand–receptor and revealing cell–cell communication patterns
SpatialDM for rapid identification of spatially co-expressed ligand–receptor and revealing cell–cell communication patterns
Spatial omics are increasingly being recognised to study cell-cell communications. Here, the authors present a bioinformatics toolbox for rapid identification of spatially co-expressed ligand-receptor and revealing cell-cell communication patterns.
- Zhuoxuan Li
- , Tianjie Wang
- & Yuanhua Huang
-
Article
| Open Access Systems-level analyses of protein-protein interaction network dysfunctions via epichaperomics identify cancer-specific mechanisms of stress adaptation
Systems-level analyses of protein-protein interaction network dysfunctions via epichaperomics identify cancer-specific mechanisms of stress adaptation
Epichaperomics allow the study of protein-protein interactions and their alterations, but probes have been limited to capturing HSP90 epichaperomes. Here, the authors introduce and validate a toolset of HSP70 epichaperome ligands, and use them in epichaperomics to identify a mechanism with which cancer cells can enhance the fitness of mitotic protein networks.
- Anna Rodina
- , Chao Xu
- & Gabriela Chiosis
-
Article
| Open Access Multi-omic features of oesophageal adenocarcinoma in patients treated with preoperative neoadjuvant therapy
Multi-omic features of oesophageal adenocarcinoma in patients treated with preoperative neoadjuvant therapy
It remains critical to understand the genomic events in response to treatment of oesophageal adenocarcinoma (OAC). Here, the authors perform a multi-omics analysis of OAC patients from the DOCTOR phase II clinical trial, finding genomic features and immune clusters associated with survival.
- Marjan M. Naeini
- , Felicity Newell
- & Nicola Waddell
-
Article
| Open Access Inference of cell type-specific gene regulatory networks on cell lineages from single cell omic datasets
Inference of cell type-specific gene regulatory networks on cell lineages from single cell omic datasets
Cell type-specific gene expression patterns are outputs of transcriptional gene regulatory networks (GRNs) that connect transcription factors and signaling proteins to target genes. Here, the authors present single-cell Multi-Task Network Inference (scMTNI), a multi-task learning framework to infer cell type-specific GRN dynamics from scRNA-seq and scATAC-seq datasets collected for diverse cell fate specification trajectories.
- Shilu Zhang
- , Saptarshi Pyne
- & Sushmita Roy
-
Article
| Open Access Using mass spectrometry imaging to map fluxes quantitatively in the tumor ecosystem
Using mass spectrometry imaging to map fluxes quantitatively in the tumor ecosystem
Isotopologue spectral analysis was originally designed to assess metabolic fluxes from bulk samples. Here, the authors adapted this approach to infer fluxes from discrete regions in tissue by using mass spectrometry imaging, showing increased fatty acid synthesis flux in brain tumors of mice.
- Michaela Schwaiger-Haber
- , Ethan Stancliffe
- & Gary J. Patti
-
Article
| Open Access NetBID2 provides comprehensive hidden driver analysis
NetBID2 provides comprehensive hidden driver analysis
It’s challenging to capture “hidden” drivers that may not be genetically-altered or differentially-expressed from omics data. Here the authors developed NetBID2, a comprehensive network-based toolbox with versatile features, enabling the integration of multi-omics data to expose such hidden drivers.
- Xinran Dong
- , Liang Ding
- & Jiyang Yu
-
Article
| Open Access Explainable multi-task learning for multi-modality biological data analysis
Explainable multi-task learning for multi-modality biological data analysis
Multimodal biological data is challenging to analyze. Here, the authors develop UnitedNet, an explainable deep neural network for analyzing single-cell multimodal biological data and estimating relationships between gene expression and other modalities with cell-type specificity.
- Xin Tang
- , Jiawei Zhang
- & Jia Liu
-
Article
| Open Access Reconstruction of the cell pseudo-space from single-cell RNA sequencing data with scSpace
Reconstruction of the cell pseudo-space from single-cell RNA sequencing data with scSpace
Methods to reanalyze scRNA-seq data in a spatial perspective are vital but lacking. Here, the authors develop scSpace, an integrative method that uses ST data as spatial reference to reconstruct the pseudo-space of scRNA-seq data and identify spatially variable cell subpopulations, providing insights into spatial heterogeneity from scRNA-seq data.
- Jingyang Qian
- , Jie Liao
- & Xiaohui Fan

 Representation of genomic intratumor heterogeneity in multi-region non-small cell lung cancer patient-derived xenograft models
Representation of genomic intratumor heterogeneity in multi-region non-small cell lung cancer patient-derived xenograft models