Featured
-
-
Article
| Open Access Dynamics of extrachromosomal circular DNA in rice
Dynamics of extrachromosomal circular DNA in rice
Comparing to other biological systems, our understanding of plant extrachromosomal circular DNA (eccDNA) is limited. Here, the authors profile eccDNA from six rice tissues and investigate eccDNA characteristics, formation mechanisms, distribution, and functional implications.
- Jundong Zhuang
- , Yaoxin Zhang
- & Tingting Lu
-
Article
| Open Access ProTInSeq: transposon insertion tracking by ultra-deep DNA sequencing to identify translated large and small ORFs
ProTInSeq: transposon insertion tracking by ultra-deep DNA sequencing to identify translated large and small ORFs
Identifying small proteins is challenging. ProTInSeq uses modified transposons to express markers inserted in-frame to protein-coding genes. This method identifies 153 unannotated small proteins in M. pneumoniae and additional proteomic information.
- Samuel Miravet-Verde
- , Rocco Mazzolini
- & Luis Serrano
-
Article
| Open Access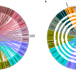 A genomic toolkit for winged bean Psophocarpus tetragonolobus
A genomic toolkit for winged bean Psophocarpus tetragonolobus
Winged bean is a tropical legume that can produce similar level of seed protein to soybean. Here, the authors report the genome assembly, population genetics, QTL mapping of the plant architecture, protein content and phytonutrients for this species.
- Wai Kuan Ho
- , Alberto Stefano Tanzi
- & Sean Mayes
-
Article
| Open Access Evolution and expression patterns of the neo-sex chromosomes of the crested ibis
Evolution and expression patterns of the neo-sex chromosomes of the crested ibis
The evolutionary trajectory of avian sex chromosomes may be more intricate than previously understood. In this study, sequencing and analysis of the neo-sex chromosomes and genome of the Crested Ibis suggests a multidirectional evolution of sex chromosomes in core waterbirds.
- Lulu Xu
- , Yandong Ren
- & Gang Li
-
Article
| Open Access Cepharanthine analogs mining and genomes of Stephania accelerate anti-coronavirus drug discovery
Cepharanthine analogs mining and genomes of Stephania accelerate anti-coronavirus drug discovery
Cepharanthine is a secondary metabolite isolated from Stephania with a variety of medicinal properties. Here, the authors assembled three Stephania genomes, propose cepharanthine biosynthetic pathway, and assess the antiviral potential of cepharanthine-related metabolites.
- Liang Leng
- , Zhichao Xu
- & Shilin Chen
-
Article
| Open Access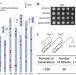 Large-scale genomic rearrangements boost SCRaMbLE in Saccharomyces cerevisiae
Large-scale genomic rearrangements boost SCRaMbLE in Saccharomyces cerevisiae
Synthetic Chromosome Rearrangement and Modification by LoxP-mediated Evolution (SCRaMbLE) is a promising tool to study genomic rearrangements. Here the authors present an engineered yeast strain with 83 sparsely distributed loxPsym sites across the genome can genrerate large-scale genomic rearrangements, which benefits cell fitness under stress and boosts the SCRaMbLE system when combined with synthetic chromosomes.
- Li Cheng
- , Shijun Zhao
- & Junbiao Dai
-
Article
| Open Access Transposable elements mediate genetic effects altering the expression of nearby genes in colorectal cancer
Transposable elements mediate genetic effects altering the expression of nearby genes in colorectal cancer
It has been suggested that transposable elements (TE) play a role in tumourigenesis, but the associated mechanisms remain unclear. Here, the authors show, using colorectal cancer data and Bayesian Networks, that TEs can mediate the effect of expression quantitative trait loci and contribute to the regulation of cancer-related genes.
- Nikolaos M. R. Lykoskoufis
- , Evarist Planet
- & Emmanouil T. Dermitzakis
-
Article
| Open Access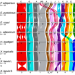 Conserved chromatin and repetitive patterns reveal slow genome evolution in frogs
Conserved chromatin and repetitive patterns reveal slow genome evolution in frogs
Frogs are an ancient and ecologically diverse group of amphibians that include important model systems. This paper reports genome sequences of multiple frog species, revealing remarkable stability of frog chromosomes and centromeres, along with highly recombinogenic extended subtelomeres.
- Jessen V. Bredeson
- , Austin B. Mudd
- & Daniel S. Rokhsar
-
Editorial
| Open AccessEfficient genetic improvement of orphan crops cannot follow the old path
Orphan crops hold the potential to diversify our food systems. Considering their unique characteristics, our deep understanding of major crops, and the availability of modern genomic tools, taking a different research path from what major crops have gone through could accelerate the genetic improvement of orphan crops.
-
Comment
| Open Access Integrative and inclusive genomics to promote the use of underutilised crops
Integrative and inclusive genomics to promote the use of underutilised crops
Underutilised crops or orphan crops are important for diversifying our food systems towards food and nutrition security. Here, the authors discuss how the development of underutilised crop genomic resource should align with their breeding and capacity building strategies, and leverage advances made in major crops.
- Oluwaseyi Shorinola
- , Rose Marks
- & Mark A. Chapman
-
Article
| Open Access Topological structures and syntenic conservation in sea anemone genomes
Topological structures and syntenic conservation in sea anemone genomes
Slowly evolving cnidarians are useful models to study genome architecture. This study shows that sea anemones have a high degree of chromosomal macrosynteny, but poor microsynteny conservation. This is correlated with a small genome size and short distances of cis-regulatory elements to genes.
- Bob Zimmermann
- , Juan D. Montenegro
- & Ulrich Technau
-
Article
| Open Access Mosaic chromosomal alterations in peripheral blood leukocytes of children in sub-Saharan Africa
Mosaic chromosomal alterations in peripheral blood leukocytes of children in sub-Saharan Africa
Mosaic chromosomal alterations (mCAs) in peripheral blood leukocytes are associated with an increased risk of malignancy. Here, the authors use genome-wide genotyping array data to investigate the prevalence of mCAs in sub-Saharan African children with versus those without Burkitt lymphoma.
- Weiyin Zhou
- , Anja Fischer
- & Sam M. Mbulaiteye
-
Article
| Open Access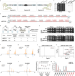 Building a eukaryotic chromosome arm by de novo design and synthesis
Building a eukaryotic chromosome arm by de novo design and synthesis
In Saccharomyces cerevisiae, the left arm of chromosome XII only requires 12 genes to maintain cell viability, whereas 25 genes are needed for robust fitness. Here the authors demonstrate that the entire arm can be replaced by a neochromosome with completely artificial sequences.
- Shuangying Jiang
- , Zhouqing Luo
- & Junbiao Dai
-
Article
| Open Access Pan-genome analysis of 13 Malus accessions reveals structural and sequence variations associated with fruit traits
Pan-genome analysis of 13 Malus accessions reveals structural and sequence variations associated with fruit traits
A pan-genome can reduce bias in genetic diversity analysis inherent in using a single reference genome. Here, the authors assemble genomes of 10 diverse apple accessions, conduct pan-genome analysis together with three existing genomes, and reveal the role of mitogen-activated protein kinase homolog MMK2 in fruit coloration.
- Ting Wang
- , Shiyao Duan
- & Ting Wu
-
Article
| Open Access PRO-IP-seq tracks molecular modifications of engaged Pol II complexes at nucleotide resolution
PRO-IP-seq tracks molecular modifications of engaged Pol II complexes at nucleotide resolution
Tracking molecular modifications of engaged Pol II complexes is important for studying transcriptional regulation. Here, the authors combine run-on sequencing (PRO-seq) and immunoprecipitation, revealing dynamics of Pol II CTD phosphorylation at nucleotide-resolution.
- Anniina Vihervaara
- , Philip Versluis
- & John T. Lis
-
Article
| Open Access Systematic analysis of paralogous regions in 41,755 exomes uncovers clinically relevant variation
Systematic analysis of paralogous regions in 41,755 exomes uncovers clinically relevant variation
Chameleolyser enables the accurate identification of genetic variants hidden within complex regions of the genome. Its application uncovers the disease-explanatory variant in 25 previously undiagnosed patients.
- Wouter Steyaert
- , Lonneke Haer-Wigman
- & Christian Gilissen
-
Article
| Open Access 3D chromatin interactions involving Drosophila insulators are infrequent but preferential and arise before TADs and transcription
3D chromatin interactions involving Drosophila insulators are infrequent but preferential and arise before TADs and transcription
In Drosophila, insulators may be involved in the organization of Topological Associated Domains, but the mechanism of action is a still a matter of investigation. Here the authors investigate the role of insulators in the 3D organization of the Drosophila genome by combining bioinformatics analysis and Hi-M, an imaging-based methods developed to detect the 3D positions of multiple genomic loci in single cells.
- Olivier Messina
- , Flavien Raynal
- & Marcelo Nollmann
-
Article
| Open Access Developing mitochondrial base editors with diverse context compatibility and high fidelity via saturated spacer library
Developing mitochondrial base editors with diverse context compatibility and high fidelity via saturated spacer library
Ddd-Aderived cytosine base editors (DdCBEs) are important for research of mitochondrial DNA mutation diseases. Here the authors report a strategy for screening and characterising dsDNA cytidine deaminases, and identify 7 DddA homologs which they optimise to minimise nuclear and mitochondrial off-target editing.
- Haifeng Sun
- , Zhaojun Wang
- & Bin Shen
-
Article
| Open Access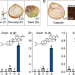 Characterization of the horse chestnut genome reveals the evolution of aescin and aesculin biosynthesis
Characterization of the horse chestnut genome reveals the evolution of aescin and aesculin biosynthesis
Horse chestnut (Aesculus chinensis) is a tree species that can produce medicinal compounds such as aescin and aesculin. Here, the authors assemble its genome, identify key genes involved in the biosynthesis of these two group of compounds, and achieve the de novo synthesis of aesculin in E. coli.
- Wei Sun
- , Qinggang Yin
- & Shilin Chen
-
Article
| Open Access The pan-genome and local adaptation of Arabidopsis thaliana
The pan-genome and local adaptation of Arabidopsis thaliana
Single reference genomes and short-read sequencing data are not enough to harness the full genetic variation of a species. Here, the authors report pan-genome of Arabidopsis thaliana based on chromosomal-level genomes of 32 accessions and identify variations associated with local adaptation.
- Minghui Kang
- , Haolin Wu
- & Jianquan Liu
-
Article
| Open Access Targetable NOTCH1 rearrangements in reninoma
Targetable NOTCH1 rearrangements in reninoma
Reninomas are very rare kidney tumours of juxtaglomerular cells. Here, the authors analyse reninomas using whole-genome and transcriptome sequencing, and reveal the presence and functional effects of NOTCH1 rearrangements.
- Taryn D. Treger
- , John E. G. Lawrence
- & Tanzina Chowdhury
-
Article
| Open Access Evolutionary origin of genomic structural variations in domestic yaks
Evolutionary origin of genomic structural variations in domestic yaks
Yaks have been subject to natural selection, human domestication and interspecific introgression during their evolution. Here, the authors have identified genomic structural variations and the linked genes involved in these processes in domestic yaks, to reveal new insight into genetic basis of phenotypic diversity.
- Xinfeng Liu
- , Wenyu Liu
- & Jianquan Liu
-
Article
| Open Access The mutational landscape of the adult healthy parous and nulliparous human breast
The mutational landscape of the adult healthy parous and nulliparous human breast
While many tissues have been investigated for natural somatic mutations, human breast tissue has not been well studied. Here, the authors characterize somatic mutations in human breast tissue, finding effects of age and parity.
- Biancastella Cereser
- , Angela Yiu
- & Justin Stebbing
-
Article
| Open Access Accurate haplotype construction and detection of selection signatures enabled by high quality pig genome sequences
Accurate haplotype construction and detection of selection signatures enabled by high quality pig genome sequences
Accurate haplotypes with abundant genomic variations benefit genetic research. Here, the authors accurately construct 1,874 pig haplotypes and demonstrate their applications in genome-wide association studies, prediction of breeding values and analyses of evolutionary selection.
- Xinkai Tong
- , Dong Chen
- & Lusheng Huang
-
Article
| Open Access APOGEE 2: multi-layer machine-learning model for the interpretable prediction of mitochondrial missense variants
APOGEE 2: multi-layer machine-learning model for the interpretable prediction of mitochondrial missense variants
APOGEE 2 is a machine-learning tool for assessing the fragility of the mitochondrial genome, evaluating genetic variant pathogenicity and ultimately enhancing our understanding of the clinical heterogeneity of mitochondrial genetic diseases.
- Salvatore Daniele Bianco
- , Luca Parca
- & Tommaso Mazza
-
Article
| Open Access Subtelomeric 5-enolpyruvylshikimate-3-phosphate synthase copy number variation confers glyphosate resistance in Eleusine indica
Subtelomeric 5-enolpyruvylshikimate-3-phosphate synthase copy number variation confers glyphosate resistance in Eleusine indica
Resistance to herbicide glyphosate can be evolved trough copy number variation (CNV) of its target gene EPSPS in goosegrass. Here, the authors assemble the genomes of glyphosate susceptible and resistance lines and provide evidence of sub-telomeric-repeat driven CNV of EPSPS could lead to glyphosate resistance.
- Chun Zhang
- , Nicholas A. Johnson
- & Eric L. Patterson
-
Article
| Open Access Genomic insight into domestication of rubber tree
Genomic insight into domestication of rubber tree
Understanding the genetic basis of rubber tree domestication is critical for improving natural rubber production. Here, the authors assemble the genome of the rubber tree clone CATAS8-79 and conduct population and genetic association analyses to reveal the function of phytosulfokine in regulating number of laticifer rings.
- Jinquan Chao
- , Shaohua Wu
- & Wei-Min Tian
-
Article
| Open Access Rapid gene content turnover on the germline-restricted chromosome in songbirds
Rapid gene content turnover on the germline-restricted chromosome in songbirds
Songbirds have an extra chromosome with unknown function found only in their germline. This study assembles and compares this chromosome in two closely related nightingale species, finding large differences in genetic content and only one conserved gene with probable essential function.
- Stephen A. Schlebusch
- , Jakub Rídl
- & Radka Reifová
-
Article
| Open Access The allotetraploid horseradish genome provides insights into subgenome diversification and formation of critical traits
The allotetraploid horseradish genome provides insights into subgenome diversification and formation of critical traits
Horseradish is a spicy root vegetable and it also produces horseradish peroxidase, an enzyme widely used in biochemistry applications. Here, the authors report its telomere-to-telomere reference genome, reveal subgenome diversification and the effect on the biosynthesis of glucosinolates and horseradish peroxidases.
- Fei Shen
- , Shixiao Xu
- & Martin A. Lysak
-
Article
| Open Access Oncogenic structural aberration landscape in gastric cancer genomes
Oncogenic structural aberration landscape in gastric cancer genomes
Gastric cancers (GC) are driven by genomic alterations, but the underlying molecular mechanisms remain unclear. Here, the authors analyse the structural rearrangement landscape of 170 GCs using whole-genome sequencing, identify recurrent structural variant hotspots and find oncogene amplicons driven by extrachromosomal DNA.
- Mihoko Saito-Adachi
- , Natsuko Hama
- & Tatsuhiro Shibata
-
Article
| Open Access Selective binding of retrotransposons by ZFP352 facilitates the timely dissolution of totipotency network
Selective binding of retrotransposons by ZFP352 facilitates the timely dissolution of totipotency network
During zygotic genome activation the embryo must re-wire the regulatory network that sustains totipotency earlier during development. Here they identify ZFP352 as an essential factor that targets retrotransposon families to facilitate dissolution of the totipotency network and enable ZGA.
- Zhengyi Li
- , Haiyan Xu
- & Hongqing Liang
-
Article
| Open Access Diploid and tetraploid genomes of Acorus and the evolution of monocots
Diploid and tetraploid genomes of Acorus and the evolution of monocots
Acorales is sister to all other monocots and contains only one family with just one genus, Acorus. Here, the authors assemble the genome of the diploid Ac. gramineus and the tetraploid Ac. calamus, reconstruct an ancestral monocot karyotype and gene toolkit, and discuss the origin and evolution of the two species and other monocots.
- Liang Ma
- , Ke-Wei Liu
- & Zhong-Jian Liu
-
Article
| Open Access The genome of Acorus deciphers insights into early monocot evolution
The genome of Acorus deciphers insights into early monocot evolution
Monocots are one of the most diverse and dominant clades of flowering plants. Here, the authors assemble the genome of Acorus gramineus, confirm its phylogenetic position as sister to the rest of monocots and reveal the absence of tau (τ) whole-genome duplication observed in the majority of monocot clades.
- Xing Guo
- , Fang Wang
- & Huan Liu
-
Article
| Open Access Origin of minicircular mitochondrial genomes in red algae
Origin of minicircular mitochondrial genomes in red algae
While the organelle genome is commonly considered to be a single circular DNA molecule, extensive variation exists. Here, the authors report multipartite minicircular genomes in red algae and indicate an origin driven by recombination due to loss of DNA replication, recombination, and repair genes.
- Yongsung Lee
- , Chung Hyun Cho
- & Hwan Su Yoon
-
Article
| Open Access Direct haplotype-resolved 5-base HiFi sequencing for genome-wide profiling of hypermethylation outliers in a rare disease cohort
Direct haplotype-resolved 5-base HiFi sequencing for genome-wide profiling of hypermethylation outliers in a rare disease cohort
HiFi genome sequencing accesses DNA methylation and nucleotide variation in long sequence reads. Here, the authors apply this approach in a rare disease cohort to identify DNA hypermethylation linked to genetic variants including rare disease alleles.
- Warren A. Cheung
- , Adam F. Johnson
- & Tomi Pastinen
-
Article
| Open Access Chromosome-level genome assembly and population genomic resource to accelerate orphan crop lablab breeding
Chromosome-level genome assembly and population genomic resource to accelerate orphan crop lablab breeding
Lablab is a legume native to Africa and cultivated throughout the tropics for food and forage; however, as an orphan crop, limited genomic resources hampers its genetic improvement. Here, an African-led South-North plant genome collaboration produces an improved genome assembly and population genomic resource to accelerate its breeding.
- Isaac Njaci
- , Bernice Waweru
- & Chris S. Jones
-
Article
| Open Access Towards routine chromosome-scale haplotype-resolved reconstruction in cancer genomics
Towards routine chromosome-scale haplotype-resolved reconstruction in cancer genomics
The precise inference of structural variants (SVs) requires suitable sequencing technologies and computational tools. Here, in order to analyse SVs with haplotype resolution, the author applies high-resolution long-read sequencing and long-range Hi-C to a melanoma cell line and develops an efficient graph-based computational framework, pstools.
- Shilpa Garg
-
Article
| Open Access Multitrait genome-wide analyses identify new susceptibility loci and candidate drugs to primary sclerosing cholangitis
Multitrait genome-wide analyses identify new susceptibility loci and candidate drugs to primary sclerosing cholangitis
The genetic basis of primary sclerosing cholangitis has only been partially uncovered. Here, the authors perform a multitrait genome-wide association study to provide insight into the genetic etiology of primary sclerosing cholangitis risk and possible therapeutic drug targets.
- Younghun Han
- , Jinyoung Byun
- & Christopher I. Amos
-
Article
| Open Access Multivariate genomic architecture of cortical thickness and surface area at multiple levels of analysis
Multivariate genomic architecture of cortical thickness and surface area at multiple levels of analysis
The current study identifies five genomic subclusters of brain regions for cortical thickness and surface area characterized by high levels of shared genetic signal. These subclusters map onto biological and functional parcellations of the cortex.
- Andrew D. Grotzinger
- , Travis T. Mallard
- & Jordan W. Smoller
-
Article
| Open Access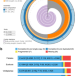 Genomics and biochemical analyses reveal a metabolon key to β-L-ODAP biosynthesis in Lathyrus sativus
Genomics and biochemical analyses reveal a metabolon key to β-L-ODAP biosynthesis in Lathyrus sativus
Grass pea is a multi-stress tolerant orphan crop and developing cultivars with decreased accumulation of the neurotoxin β-L-oxalyl-2,3-diaminopropionic acid (β-L-ODAP) is one of its breeding objectives. Here, the authors assemble its genome and reveal genes involved in the biosynthesis of β-L-ODAP.
- Anne Edwards
- , Isaac Njaci
- & Peter M. F. Emmrich
-
Article
| Open Access Amniotes co-opt intrinsic genetic instability to protect germ-line genome integrity
Amniotes co-opt intrinsic genetic instability to protect germ-line genome integrity
Pachytene Piwi-interacting RNAs (piRNAs) expressed in mammalian germ lines are abundant, but their evolution and function are not fully understood. Here, the authors find that pachytene piRNA loci are hotspots of structural variation, which underlies rapid piRNA birth, divergence, and loss.
- Yu H. Sun
- , Hongxiao Cui
- & Xin Zhiguo Li
-
Article
| Open Access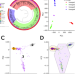 Genome-wide host-pathogen analyses reveal genetic interaction points in tuberculosis disease
Genome-wide host-pathogen analyses reveal genetic interaction points in tuberculosis disease
Few genetic loci have been associated with tuberculosis infection, possibly because of the influence of genetic variation in the pathogen. Here, the authors integrate human and Mycobacterium tuberculosis genetics to find genome-genome interactions associated with infection.
- Jody Phelan
- , Paula Josefina Gomez-Gonzalez
- & Taane G. Clark
-
Article
| Open Access Common evolutionary trajectory of short life-cycle in Brassicaceae ruderal weeds
Common evolutionary trajectory of short life-cycle in Brassicaceae ruderal weeds
Understanding origin and adaptation of weeds is important for their management. Here, via genome assembly, population genomics, and QTL mapping, the authors establish Cardamine occulta as a model to study weed ruderality and show FLC and CRY2 as genetic drivers for the establishment of short life cycle.
- Ling-Zi Li
- , Zhou-Geng Xu
- & Jia-Wei Wang
-
Article
| Open Access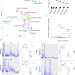 Primate-specific transposable elements shape transcriptional networks during human development
Primate-specific transposable elements shape transcriptional networks during human development
The human genome harbors more than 4.5 million transposable element (TE)-derived insertions, the result of recurrent waves of invasion and internal propagation. Here they show that TEs belonging to evolutionarily recent subfamilies go on to regulate later stages of human embryonic development, notably conditioning the expression of genes involved in gastrulation and early organogenesis.
- Julien Pontis
- , Cyril Pulver
- & Didier Trono
-
Article
| Open Access Transposable elements orchestrate subgenome-convergent and -divergent transcription in common wheat
Transposable elements orchestrate subgenome-convergent and -divergent transcription in common wheat
How subgenome-divergent and -convergent transcription is mediated and harmonized in hexaploid common wheat genome remains unclear. Here, via characterizing the cistrome maps, the authors reveal that transposon elements with transcription factor binding ability have the potential to make the contribution.
- Yuyun Zhang
- , Zijuan Li
- & Yijing Zhang
-
Article
| Open Access Integration of Hi-C with short and long-read genome sequencing reveals the structure of germline rearranged genomes
Integration of Hi-C with short and long-read genome sequencing reveals the structure of germline rearranged genomes
Here the authors characterize structural variations (SVs) in a cohort of individuals with complex genomic rearrangements, identifying breakpoints by employing short- and long-read genome sequencing and investigate their impact on gene expression and the three-dimensional chromatin architecture. They find breakpoints are enriched in inactive regions and can result in chromatin domain fusions.
- Robert Schöpflin
- , Uirá Souto Melo
- & Stefan Mundlos
-
Article
| Open Access Epigenetic control of chromosome-associated lncRNA genes essential for replication and stability
Epigenetic control of chromosome-associated lncRNA genes essential for replication and stability
Heskett et al. describe several members of a class of long non-coding RNAs, known as ASARs, which show distinct epigenetic regulation between subclonal lineages and are essential for normal DNA replication timing and stability of human autosomes.
- Michael B. Heskett
- , Athanasios E. Vouzas
- & Mathew J. Thayer
-
Article
| Open Access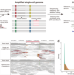 Prevalence and mechanisms of somatic deletions in single human neurons during normal aging and in DNA repair disorders
Prevalence and mechanisms of somatic deletions in single human neurons during normal aging and in DNA repair disorders
DNA damage has been implicated in aging and neurodegeneration. Here, the authors develop a bioinformatic method to detect deletions in single neuron genome sequences and reveal an increased burden of somatic deletions during aging and in DNA repair disorders.
- Junho Kim
- , August Yue Huang
- & Eunjung Alice Lee
-
Article
| Open Access A method for multiplexed full-length single-molecule sequencing of the human mitochondrial genome
A method for multiplexed full-length single-molecule sequencing of the human mitochondrial genome
Accurate analysis of mitochondrial DNA is important for mitochondrial disease clinical research and diagnostics. Here, authors present a method using Cas9 cleavage, nanopore sequencing and a custom pipeline to identify pathogenic variants, deletions and accurately quantify heteroplasmy to below 1%.
- Ieva Keraite
- , Philipp Becker
- & Ivo Glynne Gut

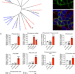 Illuminating the function of the orphan transporter, SLC22A10, in humans and other primates
Illuminating the function of the orphan transporter, SLC22A10, in humans and other primates