Featured
-
-
Article
| Open Access DNA methylation-based high-resolution mapping of long-distance chromosomal interactions in nucleosome-depleted regions
DNA methylation-based high-resolution mapping of long-distance chromosomal interactions in nucleosome-depleted regions
Here, the authors present MTAC, a method to map chromosomal interactions in budding yeast. By applying MTAC to various viewpoints, they find that most of the long-distance chromosomal interactions detected by MTAC reflect tethering by the nuclear pore complexes.
- Yi Li
- , James Lee
- & Lu Bai
-
Article
| Open Access An improved epigenetic counter to track mitotic age in normal and precancerous tissues
An improved epigenetic counter to track mitotic age in normal and precancerous tissues
DNA methylation (DNAm) clocks can track mitotic age, but their potential use for cancer risk prediction remains less explored. Here, the authors develop a DNAm counter of total mitotic age (stemTOC) that shows an increase of mitotic age in normal tissues and precancerous lesions.
- Tianyu Zhu
- , Huige Tong
- & Andrew E. Teschendorff
-
Article
| Open Access A time-resolved multi-omics atlas of transcriptional regulation in response to high-altitude hypoxia across whole-body tissues
A time-resolved multi-omics atlas of transcriptional regulation in response to high-altitude hypoxia across whole-body tissues
The mechanisms underlying high-altitude acclimatization remain unclear. Here authors use the sheep model to reveal multi-tissue temporal dynamics of gene transcription and regulation during acclimatization, and provide resources for hypoxia-related studies.
- Ze Yan
- , Ji Yang
- & Meng-Hua Li
-
Article
| Open Access Pervasive structural heterogeneity rewires glioblastoma chromosomes to sustain patient-specific transcriptional programs
Pervasive structural heterogeneity rewires glioblastoma chromosomes to sustain patient-specific transcriptional programs
By applying Hi-C to cells derived from the tumors of 24 GBM patients, the authors show pervasive structural variation in GBM chromosomal organization. How such patient-to-patient variation explains the characteristic gene expression patterns in each tumor is investigated.
- Ting Xie
- , Adi Danieli-Mackay
- & Argyris Papantonis
-
Article
| Open Access Disease related changes in ATAC-seq of iPSC-derived motor neuron lines from ALS patients and controls
Disease related changes in ATAC-seq of iPSC-derived motor neuron lines from ALS patients and controls
Amyotrophic Lateral Sclerosis (ALS) is highly heritable but the mechanisms of sporadic ALS are not fully understood. In this study, the authors identify drivers of variation and disease-relevant changes in the epigenomic profile of iPSC-derived motor neuron lines generated from ALS patients and healthy controls as part of the Answer ALS program.
- Stanislav Tsitkov
- , Kelsey Valentine
- & Ernest Fraenkel
-
Article
| Open Access Nonlinear DNA methylation trajectories in aging male mice
Nonlinear DNA methylation trajectories in aging male mice
DNA methylation is an age biomarker, but nonlinear aspects of its age-related dynamics are not well characterized. Here, the authors identify loci that undergo sudden methylation changes at specific life stages in the aging colon of male mice.
- Maja Olecka
- , Alena van Bömmel
- & Steve Hoffmann
-
Article
| Open Access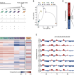 p53 rapidly restructures 3D chromatin organization to trigger a transcriptional response
p53 rapidly restructures 3D chromatin organization to trigger a transcriptional response
Here the authors uncover p53’s role as the master regulator of spatio-temporal genome organization. p53 controls the expression of 340 distal genes through newly formed and pre-existing loops between p53-bound enhancers and promoters.
- François Serra
- , Andrea Nieto-Aliseda
- & Biola M. Javierre
-
Article
| Open Access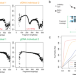 FinaleMe: Predicting DNA methylation by the fragmentation patterns of plasma cell-free DNA
FinaleMe: Predicting DNA methylation by the fragmentation patterns of plasma cell-free DNA
DNA methylation from cell-free DNA (cfDNA) can be profiled using whole genome bisulfite sequencing (WGBS). Here, the authors develop a computational method, FinaleMe, that predicts DNA methylation and tissues of-origin in cfDNA and validate its performance using paired deep and shallow-coverage whole-genome sequencing (WGS) and WGBS data.
- Yaping Liu
- , Sarah C. Reed
- & Manolis Kellis
-
Article
| Open Access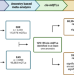 Genetic control of DNA methylation is largely shared across European and East Asian populations
Genetic control of DNA methylation is largely shared across European and East Asian populations
Most functional genomic resources have been based on individuals with European ancestry. Here, the authors perform DNA methylation quantitative trait locus (mQTL) analyses in 3,701 European and 2,099 East Asian individuals to identify thousands of genetic variants, a large degree of which are shared between the two populations.
- Alesha A. Hatton
- , Fei-Fei Cheng
- & Allan F. McRae
-
Article
| Open Access Aberrant non-canonical NF-κB signalling reprograms the epigenome landscape to drive oncogenic transcriptomes in multiple myeloma
Aberrant non-canonical NF-κB signalling reprograms the epigenome landscape to drive oncogenic transcriptomes in multiple myeloma
The downstream molecular mechanisms following the activation of the NF-κB pathway in multiple myeloma (MM) remain to be characterised. Here, it is shown that aberrant non-canonical NF-κB signalling causes epigenomic reprogramming leading to transcriptional changes that favour MM progression.
- Daniel A. Ang
- , Jean-Michel Carter
- & Yinghui Li
-
Article
| Open Access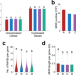 Dynamics of accessible chromatin regions and subgenome dominance in octoploid strawberry
Dynamics of accessible chromatin regions and subgenome dominance in octoploid strawberry
Subgenome dominance is widely observed in allopolyploid species, but the molecular mechanisms remain unclear. Here, the authors generate genome-wide map of accessible chromatin regions (ACRs) in allo-octoploid cultivated strawberry and reveal that dynamics of the ACRs play an important role in its subgenome dominance.
- Chao Fang
- , Ning Jiang
- & Jiming Jiang
-
Article
| Open Access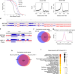 The chromatin factors SET-26 and HCF-1 oppose the histone deacetylase HDA-1 in longevity and gene regulation in C. elegans
The chromatin factors SET-26 and HCF-1 oppose the histone deacetylase HDA-1 in longevity and gene regulation in C. elegans
SET-26, HCF-1, HDA-1 are factors that help package DNA and regulate gene expression. Here, the authors find that SET-26 recruits HCF-1 to specific locations along the genome, where they antagonize the activity of HDA-1. Together, they maintain balanced gene expression and longevity in C. elegans.
- Felicity J. Emerson
- , Caitlin Chiu
- & Siu Sylvia Lee
-
Article
| Open Access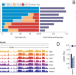 Extensive DNA methylome rearrangement during early lamprey embryogenesis
Extensive DNA methylome rearrangement during early lamprey embryogenesis
DNA methylation plays a major role in establishing cell identity, but the dynamics of DNA methylation patterns are highly variable across species. Here, the authors discover extensive DNA methylation reprogramming during embryonic development of the sea lamprey, a jawless fish with a distinctive, highly disordered methylome.
- Allegra Angeloni
- , Skye Fissette
- & Ozren Bogdanovic
-
Article
| Open Access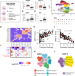 Complex regulatory networks influence pluripotent cell state transitions in human iPSCs
Complex regulatory networks influence pluripotent cell state transitions in human iPSCs
Stem cells exist in vitro in a spectrum of interconvertible pluripotent states. Here, authors show that pluripotency and self-renewal processes have a high level of regulatory complexity and suggest that genetic factors contribute to cell state transitions in human iPSC lines.
- Timothy D. Arthur
- , Jennifer P. Nguyen
- & Kelly A. Frazer
-
Article
| Open Access Multi-omics analysis in human retina uncovers ultraconserved cis-regulatory elements at rare eye disease loci
Multi-omics analysis in human retina uncovers ultraconserved cis-regulatory elements at rare eye disease loci
Ultraconserved non-coding elements (UCNEs) can regulate developmental gene expression. Retinal multi-omics data integration revealed UCNEs to be candidate cis-regulatory elements during retinal development, which may be implicated in rare eye diseases.
- Victor Lopez Soriano
- , Alfredo Dueñas Rey
- & Elfride De Baere
-
Article
| Open Access Chromatin attachment to the nuclear matrix represses hypocotyl elongation in Arabidopsis thaliana
Chromatin attachment to the nuclear matrix represses hypocotyl elongation in Arabidopsis thaliana
The role of the nuclear matrix in plant nuclei is unclear. Here the authors reveal that nuclear matrix-associated proteins act as a regulatory hub, recruiting both DNA and transcriptional repressors to the nuclear matrix
- Linhao Xu
- , Shiwei Zheng
- & Hua Jiang
-
Article
| Open Access Orchestrating chromosome conformation capture analysis with Bioconductor
Orchestrating chromosome conformation capture analysis with Bioconductor
The Bioconductor project aims to develop R packages for analysis of genomic datasets. Here the authors show the HiCExperiment package suite and its companion online book (https://bioconductor.org/books/OHCA/) which present data structures, computational methods and visualization tools available in Bioconductor to investigate chromatin conformation capture (3C) data in R.
- Jacques Serizay
- , Cyril Matthey-Doret
- & Romain Koszul
-
Article
| Open Access The UBP5 histone H2A deubiquitinase counteracts PRCs-mediated repression to regulate Arabidopsis development
The UBP5 histone H2A deubiquitinase counteracts PRCs-mediated repression to regulate Arabidopsis development
The authors demonstrate that UBIQUITIN SPECIFIC PROTEASE 5 (UBP5) is a major H2Aub deubiquitinase that antagonises H3K27me3, leading to transcriptional de-repression and controlling development in Arabidopsis.
- James Godwin
- , Mohan Govindasamy
- & Sara Farrona
-
Article
| Open Access DiffDomain enables identification of structurally reorganized topologically associating domains
DiffDomain enables identification of structurally reorganized topologically associating domains
Topologically associating domains (TADs) are critical structural units in 3D genome organization, and their reorganization between health and disease states is associated with essential genome functions. However, computational methods for identifying reorganized TADs are still in the early stages of development. Here, the authors present an algorithm leveraging random matrix theory to identify reorganized TADs.
- Dunming Hua
- , Ming Gu
- & Dechao Tian
-
Article
| Open Access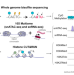 The chromatin landscape of healthy and injured cell types in the human kidney
The chromatin landscape of healthy and injured cell types in the human kidney
Comprehensive integration of gene expression with epigenetic features is needed to understand the transition of kidney cells from health to injury. Here, the authors integrate dual single nucleus RNA expression and chromatin accessibility, DNA methylation, and histone modifications to decipher the chromatin landscape of the kidney in reference and adaptive injury cell states, identifying a transcription factor network of ELF3, KLF6, and KLF10 which regulates adaptive repair and maladaptive failed repair.
- Debora L. Gisch
- , Michelle Brennan
- & Michael T. Eadon
-
Article
| Open Access Mapping nucleosome-resolution chromatin organization and enhancer-promoter loops in plants using Micro-C-XL
Mapping nucleosome-resolution chromatin organization and enhancer-promoter loops in plants using Micro-C-XL
The authors employ Micro-C-XL to investigate chromatin structures in plants, specifically focusing on nucleosome-resolution chromatin organizations and enhancer-promoter chromatin loops in Arabidopsis, rice, and soybean.
- Linhua Sun
- , Jingru Zhou
- & Hang He
-
Article
| Open Access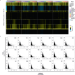 Cell-type differential targeting of SETDB1 prevents aberrant CTCF binding, chromatin looping, and cis-regulatory interactions
Cell-type differential targeting of SETDB1 prevents aberrant CTCF binding, chromatin looping, and cis-regulatory interactions
Here, the authors show how the histone methyltransferase SETDB1 is involved in cell-type specific regulation of chromatin landscape by catalyzing H3K9me3, which antagonizes CTCF binding. They further define the subsequent transcriptomic impact.
- Phoebe Lut Fei Tam
- , Ming Fung Cheung
- & Danny Leung
-
Article
| Open Access Single cell multi-omics reveal intra-cell-line heterogeneity across human cancer cell lines
Single cell multi-omics reveal intra-cell-line heterogeneity across human cancer cell lines
Intra-cell line heterogeneity remains to be characterized. Here, the use of single multi-omics on a large panel of human cell lines identifies copy number variation, epigenetic variation and extrachromosomal DNA distribution as the main contributors to intra-cell line heterogeneity.
- Qionghua Zhu
- , Xin Zhao
- & Liang Wu
-
Article
| Open Access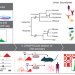 TAD evolutionary and functional characterization reveals diversity in mammalian TAD boundary properties and function
TAD evolutionary and functional characterization reveals diversity in mammalian TAD boundary properties and function
The authors show that the deletion of ultraconserved TAD boundaries affects gene expression and phenotype, highlighting TAD evolution’s function. Human-specific TAD boundaries reveal a role in brain development and disease.
- Mariam Okhovat
- , Jake VanCampen
- & Lucia Carbone
-
Article
| Open Access Outward-oriented sites within clustered CTCF boundaries are key for intra-TAD chromatin interactions and gene regulation
Outward-oriented sites within clustered CTCF boundaries are key for intra-TAD chromatin interactions and gene regulation
The TAD boundaries comprise clustered arrays of CTCF sites with complex orientations. Here the authors show that the outward-oriented CTCF sites within clustered TAD boundaries are central for intra-TAD chromatin spatial contacts and gene regulation.
- Xiao Ge
- , Haiyan Huang
- & Qiang Wu
-
Article
| Open Access SMARCB1 loss activates patient-specific distal oncogenic enhancers in malignant rhabdoid tumors
SMARCB1 loss activates patient-specific distal oncogenic enhancers in malignant rhabdoid tumors
The regulatory landscape of malignant rhabdoid tumor (MRT) due to SMARCB1 loss remains to be explored. Here, the authors perform multi-omics analysis using patient-derived MRT organoids and characterise the epigenetic reprogramming events underlying SMARCB1 loss.
- Ning Qing Liu
- , Irene Paassen
- & Jarno Drost
-
Article
| Open Access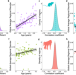 The rate of epigenetic drift scales with maximum lifespan across mammals
The rate of epigenetic drift scales with maximum lifespan across mammals
Epigenetic drift has been hypothesized to contribute to epigenetic clock signals and variation in lifespan across species. Here, the authors show that an empirical measure of epigenetic drift scales with maximum lifespan across four mammal species and accumulates in non-random genomic locations.
- Emily M. Bertucci-Richter
- & Benjamin B. Parrott
-
Article
| Open Access Preneoplastic liver colonization by 11p15.5 altered mosaic cells in young children with hepatoblastoma
Preneoplastic liver colonization by 11p15.5 altered mosaic cells in young children with hepatoblastoma
Paediatric liver cancer is rare, and often associated with a predisposition syndrome. Here, the authors show that 11p15.5 mosaic alteration in the liver is a pre-neoplastic lesion associated with hepatoblastoma, and spatial transcriptomics together with single-nucleus RNAseq identify a an altered zonation in the liver of these patients.
- Jill Pilet
- , Theo Z. Hirsch
- & Jessica Zucman-Rossi
-
Article
| Open Access The admixed brushtail possum genome reveals invasion history in New Zealand and novel imprinted genes
The admixed brushtail possum genome reveals invasion history in New Zealand and novel imprinted genes
The brushtail possum is a treasured Australian marsupial, but also a harmful pest introduced into New Zealand. Here, using functional genomics and a new chromosome-level genome assembly of New Zealand possums, Bond et al. quantify their genome admixture and identify unique parent-specific and weaning associated gene expression.
- Donna M. Bond
- , Oscar Ortega-Recalde
- & Timothy A. Hore
-
Article
| Open Access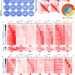 Regulation of CTCF loop formation during pancreatic cell differentiation
Regulation of CTCF loop formation during pancreatic cell differentiation
Here the authors show that differentiation of hESCs into pancreatic cells involves changes in CTCF loops by alteration of pioneer factor recruitment, histone modifications, and DNA methylation, leading to changes in enhancer-promoter interactions and gene expression.
- Xiaowen Lyu
- , M. Jordan Rowley
- & Victor G. Corces
-
Article
| Open Access Epigenomic analysis of formalin-fixed paraffin-embedded samples by CUT&Tag
Epigenomic analysis of formalin-fixed paraffin-embedded samples by CUT&Tag
Conducting epigenomic studies on FFPE samples is traditionally challenging due to chromatin damage caused due to exposure to formaldehyde. Here, the authors show that an optimisation of their previous CUTAC method allows the production of high-resolution maps of regulatory elements from FFPE samples.
- Steven Henikoff
- , Jorja G. Henikoff
- & Eric C. Holland
-
Article
| Open Access Context-dependent perturbations in chromatin folding and the transcriptome by cohesin and related factors
Context-dependent perturbations in chromatin folding and the transcriptome by cohesin and related factors
Enhancer–promoter looping and topologically associating domain are at the base of chromatin structures. Here the authors present a computational workflow in which multi-omics datasets are compared systematically to explore how three-dimensional (3D) structure and gene expression are regulated by cohesin and related factors.
- Ryuichiro Nakato
- , Toyonori Sakata
- & Katsuhiko Shirahige
-
Article
| Open Access Single cell multiomic analysis reveals diabetes-associated β-cell heterogeneity driven by HNF1A
Single cell multiomic analysis reveals diabetes-associated β-cell heterogeneity driven by HNF1A
The mechanism and disease-relevance of pancreatic b-cell heterogeneity remains elusive. Here the authors show that variable HNF1A-FXYD2 activity drives single b-cell heterogeneity at transcriptomic, epigenomic, and electro-physiological levels, which strongly mark the progression of type 2 diabetes.
- Chen Weng
- , Anniya Gu
- & Yan Li
-
Article
| Open Access DNA methylation at the suppressor of cytokine signaling 3 (SOCS3) gene influences height in childhood
DNA methylation at the suppressor of cytokine signaling 3 (SOCS3) gene influences height in childhood
Here the authors identify an association between methylation of the suppressor of cytokine signaling 3 (SOCS3) and height in children from India, Gambia and the UK.
- Prachand Issarapu
- , Manisha Arumalla
- & Stephen Owens
-
Article
| Open Access A deep learning method for replicate-based analysis of chromosome conformation contacts using Siamese neural networks
A deep learning method for replicate-based analysis of chromosome conformation contacts using Siamese neural networks
Siamese neural networks are a powerful deep learning approach for image analysis. Here, the authors adapt this method to the replicate-based analysis of Hi-C data and find that it successfully discriminates technical noise from biological variation.
- Ediem Al-jibury
- , James W. D. King
- & Daniel Rueckert
-
Article
| Open Access Single-cell genomics improves the discovery of risk variants and genes of atrial fibrillation
Single-cell genomics improves the discovery of risk variants and genes of atrial fibrillation
Here the authors combine an experimental and analytical approach that integrates single cell epigenomics with GWAS to prioritize risk variants and genes to provide a comprehensive map of Atrial Fibrillation risk variants and genes.
- Alan Selewa
- , Kaixuan Luo
- & Sebastian Pott
-
Article
| Open Access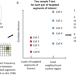 SnapFISH: a computational pipeline to identify chromatin loops from multiplexed DNA FISH data
SnapFISH: a computational pipeline to identify chromatin loops from multiplexed DNA FISH data
Multiplexed DNA FISH technologies are powerful tools to reveal chromatin spatial organisation. Here, the authors developed SnapFISH, a computational pipeline to identify chromatin loops from multiplexed DNA FISH data.
- Lindsay Lee
- , Hongyu Yu
- & Ming Hu
-
Article
| Open Access Droplet-based bisulfite sequencing for high-throughput profiling of single-cell DNA methylomes
Droplet-based bisulfite sequencing for high-throughput profiling of single-cell DNA methylomes
Single-cell DNA methylomic studies offer high resolution to differentiate cell subsets based on their epigenomic features. Here, the authors demonstrate Drop-BS, a droplet-based single-cell bisulfite sequencing library preparation method, for DNA methylome profiling.
- Qiang Zhang
- , Sai Ma
- & Chang Lu
-
Article
| Open Access Increased body mass index is linked to systemic inflammation through altered chromatin co-accessibility in human preadipocytes
Increased body mass index is linked to systemic inflammation through altered chromatin co-accessibility in human preadipocytes
Preadipocytes contribute to the pro-inflammatory environment in obesity, via unknown mechanisms. Here, comparing monozygotic twin pairs, the authors show that co-accessibility of chromatin in preadipocytes is altered in siblings with higher compared to lower BMI, and that variants in these regions contribute to systemic inflammation via interactions with BMI.
- Kristina M. Garske
- , Asha Kar
- & Päivi Pajukanta
-
Article
| Open Access DNA 5-methylcytosine detection and methylation phasing using PacBio circular consensus sequencing
DNA 5-methylcytosine detection and methylation phasing using PacBio circular consensus sequencing
Existing methods for detecting DNA methylation (5mC) are less accurate and robust. Here, the authors develop a deep learning tool ccsmeth and a Nextflow pipeline ccsmethphase for genome-wide 5mCpG detection and phasing with high accuracy from CCS reads in human.
- Peng Ni
- , Fan Nie
- & Jianxin Wang
-
Article
| Open Access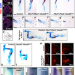 A spatio-temporally constrained gene regulatory network directed by PBX1/2 acquires limb patterning specificity via HAND2
A spatio-temporally constrained gene regulatory network directed by PBX1/2 acquires limb patterning specificity via HAND2
Many key developmental transcriptional regulators are broadly expressed but perform distinct functions in specific tissues. Here they show that ubiquitously expressed PBX factors gain limb bud functionality by interaction with HAND2, uncovering fundamental principles of cooperation between promiscuous and tissue-specific regulators to instruct developmental programs.
- Marta Losa
- , Iros Barozzi
- & Licia Selleri
-
Article
| Open Access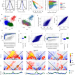 Single cell Hi-C identifies plastic chromosome conformations underlying the gastrulation enhancer landscape
Single cell Hi-C identifies plastic chromosome conformations underlying the gastrulation enhancer landscape
Here the authors use single-cell Hi-C to investigate chromosome conformation in post-gastrulation mouse embryos. They find a distinct genome organization in primitive erythrocytes and conformations matching the mesodermal and ectodermal lineages.
- Nimrod Rappoport
- , Elad Chomsky
- & Amos Tanay
-
Article
| Open Access Histone exchange sensors reveal variant specific dynamics in mouse embryonic stem cells
Histone exchange sensors reveal variant specific dynamics in mouse embryonic stem cells
Eviction of histones from nucleosomes and their exchange with newly synthesized or alternative variants is a central epigenetic determinant. Here the authors implement a molecular sensor that reports on steady-state exchange of histones in mESC and mice revealing dependency between deposition of histone variant H3.3 and exchange of H3.1 and H2B in both open and closed chromatin.
- Marko Dunjić
- , Felix Jonas
- & Yonatan Stelzer
-
Article
| Open Access Impaired histone inheritance promotes tumor progression
Impaired histone inheritance promotes tumor progression
Here the authors show disturbing parental histone inheritance in cancer cells drives tumor progression by reprogramming the epigenetic profile and conferring fitness advantages to some of the newly emerged subclones.
- Congcong Tian
- , Jiaqi Zhou
- & Haiyun Gan
-
Article
| Open Access Chromatin alternates between A and B compartments at kilobase scale for subgenic organization
Chromatin alternates between A and B compartments at kilobase scale for subgenic organization
Ultra-deep mapping of genome organization uncovers precise nuclear compartments and diffuse CTCF loops. This work demonstrates that compartment domains segregate the 5′ and 3′ ends of genes and that CTCF loops create proximal structures.
- Hannah L. Harris
- , Huiya Gu
- & M. Jordan Rowley
-
Article
| Open Access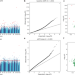 DNA methylation markers for kidney function and progression of diabetic kidney disease
DNA methylation markers for kidney function and progression of diabetic kidney disease
Epigenetic markers are potential biomarkers for diabetes and related complications. Here, the authors identify CpG sites associated with kidney function and its subsequent decline using both single-site and multisite analyses, which are shown to have functional significance in the kidney.
- Kelly Yichen Li
- , Claudia Ha Ting Tam
- & Ronald C. W. Ma
-
Article
| Open Access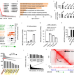 Epigenetic landscape reveals MECOM as an endothelial lineage regulator
Epigenetic landscape reveals MECOM as an endothelial lineage regulator
Errors in vascular development are associated with several congenital defects. Here they systematically investigated the epigenetic landscape of the endothelial lineage and found that MECOM depletion impairs endothelial cell differentiation and angiogenesis.
- Jie Lv
- , Shu Meng
- & Lili Zhang
-
Article
| Open Access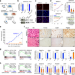 ASPSCR1::TFE3 orchestrates the angiogenic program of alveolar soft part sarcoma
ASPSCR1::TFE3 orchestrates the angiogenic program of alveolar soft part sarcoma
The mechanisms of angiogenesis in alveolar soft part sarcoma (ASPS) remain to be explored. Here, the authors highlight the role of the ASPSCR1::TFE3 fusion in regulating super-enhancer activity during the angiogenic process in ASPS.
- Miwa Tanaka
- , Surachada Chuaychob
- & Takuro Nakamura
-
Article
| Open Access MYC reshapes CTCF-mediated chromatin architecture in prostate cancer
MYC reshapes CTCF-mediated chromatin architecture in prostate cancer
The functional link between MYC and CTCF in prostate cancer remains to be investigated. Here, the authors highlight the role of MYC in rewiring chromatin architecture by interacting with CTCF protein.
- Zhao Wei
- , Song Wang
- & Haiyang Guo

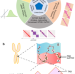 A comprehensive benchmarking with interpretation and operational guidance for the hierarchy of topologically associating domains
A comprehensive benchmarking with interpretation and operational guidance for the hierarchy of topologically associating domains