Abstract
A timely inflammatory response is crucial for early viral defense, but uncontrolled inflammation harms the host. Retinoic acid-inducible gene I (RIG-I) has a pivotal role in detecting RNA viruses, yet the regulatory mechanisms governing its sensitivity remain elusive. Here we identify PTENα, an N-terminally extended form of PTEN, as an RNA-binding protein with a preference for the CAUC(G/U)UCAU motif. Using both in vivo and in vitro viral infection assays, we demonstrated that PTENα restricted the host innate immune response, relying on its RNA-binding capacity and phosphatase activity. Mechanistically, PTENα directly bound to viral RNA and enzymatically converted its 5′-triphosphate to 5′-monophosphate, thereby reducing RIG-I sensitivity. Physiologically, brain-intrinsic PTENα exerted protective effects against viral inflammation, while peripheral PTENα restricted host antiviral immunity and, to some extent, promoted viral replication. Collectively, our findings underscore the significance of PTENα in modulating viral RNA- and RIG-I-mediated immune recognition, offering potential therapeutic implications for infectious diseases.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Raw RNA-seq data from wild-type and Ptenα–/– MEFs have been submitted to the Gene Expression Omnibus (GEO) database under accession code GSE210684. The RNA-seq data from mouse liver tissues were deposited in the NCBI Sequence Read Archive database under accession code PRJNA792209. UV cross-linking RIP–seq data were uploaded to the GEO dataset under accession number GSE218439. Single-cell RNA-seq data from wild-type and Ptenα–/– brains were submitted to the GEO database under accession code GSE238151. The HyperTRIBE data were also deposited in GEO under accession code GSE237910. The GO database was downloaded from MSigDB (http://www.gsea-msigdb.org/gsea/downloads.jsp). Source data are provided with this paper.
References
Knight, G. M. et al. Antimicrobial resistance and COVID-19: intersections and implications. eLife 10, e64139 (2021).
Gong, T., Liu, L., Jiang, W. & Zhou, R. DAMP-sensing receptors in sterile inflammation and inflammatory diseases. Nat. Rev. Immunol. 20, 95–112 (2020).
Kumar, H., Kawai, T. & Akira, S. Pathogen recognition by the innate immune system. Int. Rev. Immunol. 30, 16–34 (2011).
Loo, Y. M. & Gale, M. Jr. Immune signaling by RIG-I-like receptors. Immunity 34, 680–692 (2011).
Iselin, L. et al. Uncovering viral RNA–host cell interactions on a proteome-wide scale. Trends Biochem. Sci. 47, 23–38 (2022).
Bruderer, T., Tu, L. C. & Lee, M. G. The 5′ end structure of transcripts derived from the rRNA gene and the RNA polymerase I transcribed protein coding genes in Trypanosoma brucei. Mol. Biochem. Parasitol. 129, 69–77 (2003).
Jiao, X., Chang, J. H., Kilic, T., Tong, L. & Kiledjian, M. A mammalian pre-mRNA 5′ end capping quality control mechanism and an unexpected link of capping to pre-mRNA processing. Mol. Cell 50, 104–115 (2013).
Rehwinkel, J. & Gack, M. U. RIG-I-like receptors: their regulation and roles in RNA sensing. Nat. Rev. Immunol. 20, 537–551 (2020).
Wu, B. & Hur, S. How RIG-I like receptors activate MAVS. Curr. Opin. Virol. 12, 91–98 (2015).
Kaida, D. et al. Spliceostatin A targets SF3b and inhibits both splicing and nuclear retention of pre-mRNA. Nat. Chem. Biol. 3, 576–583 (2007).
Takeuchi, O. & Akira, S. Pattern recognition receptors and inflammation. Cell 140, 805–820 (2010).
Chen, N. Y. et al. HIV-1 capsid is involved in post-nuclear entry steps. Retrovirology 13, 28 (2016).
Yehia, L., Keel, E. & Eng, C. The clinical spectrum of PTEN mutations. Annu. Rev. Med. 71, 103–116 (2020).
Worby, C. A. & Dixon, J. E. PTEN. Annu. Rev. Biochem. 83, 641–669 (2014).
Liang, H. et al. PTENα, a PTEN isoform translated through alternative initiation, regulates mitochondrial function and energy metabolism. Cell Metab. 19, 836–848 (2014).
Hopkins, B. D. et al. A secreted PTEN phosphatase that enters cells to alter signaling and survival. Science 341, 399–402 (2013).
Masson, G. R., Perisic, O., Burke, J. E. & Williams, R. L. The intrinsically disordered tails of PTEN and PTEN-L have distinct roles in regulating substrate specificity and membrane activity. Biochem. J 473, 135–144 (2016).
Zhou, R., Liu, L. & Wang, Y. Viral proteins recognized by different TLRs. J. Med. Virol. 93, 6116–6123 (2021).
Luo, M., Terrell, J. R. & McManus, S. A. Nucleocapsid structure of negative strand RNA virus. Viruses 12, 835 (2020).
Korn, S. M., Dhamotharan, K., Jeffries, C. M. & Schlundt, A. The preference signature of the SARS-CoV-2 nucleocapsid NTD for its 5′-genomic RNA elements. Nat. Commun. 14, 3331 (2023).
Wang, S. et al. Targeting liquid–liquid phase separation of SARS-CoV-2 nucleocapsid protein promotes innate antiviral immunity by elevating MAVS activity. Nat. Cell Biol. 23, 718–732 (2021).
Rahman, R., Xu, W., Jin, H. & Rosbash, M. Identification of RNA-binding protein targets with HyperTRIBE. Nat. Protoc. 13, 1829–1849 (2018).
Hornung, V. et al. 5′-Triphosphate RNA is the ligand for RIG-I. Science 314, 994–997 (2006).
Choi, J. H. et al. DUSP11-mediated control of 5′-triphosphate RNA regulates RIG-I sensitivity. Genes Dev. 34, 1697–1712 (2020).
Wilson, M. R. & Peters, C. J. Diseases of the central nervous system caused by lymphocytic choriomeningitis virus and other arenaviruses. Handb. Clin. Neurol. 123, 671–681 (2014).
Jarrous, N. & Rouvinski, A. RNA polymerase III and antiviral innate immune response. Transcription 12, 1–11 (2021).
Sun, Y. et al. PTENα functions as an immune suppressor and promotes immune resistance in PTEN-mutant cancer. Nat. Commun. 12, 5147 (2021).
Chen, E. et al. Poly(I:C) preconditioning protects the heart against myocardial ischemia/reperfusion injury through TLR3/PI3K/AKT-dependent pathway. Signal Transduct. Target. Ther. 5, 216 (2020).
Ehrhardt, C. et al. Bivalent role of the phosphatidylinositol-3-kinase (PI3K) during influenza virus infection and host cell defence. Cell Microbiol. 8, 1336–1348 (2006).
Blanco, J., Cameirao, C., López, M. C. & Muñoz-Barroso, I. Phosphatidylinositol-3-kinase–AKT pathway in negative-stranded RNA virus infection: a minireview. Arch. Virol. 165, 2165–2176 (2020).
Cao, Y. et al. PTEN-L promotes type I interferon responses and antiviral immunity. Cell. Mol. Immunol. 15, 48–57 (2018).
Shen, S. M. et al. PTENα and PTENβ promote carcinogenesis through WDR5 and H3K4 trimethylation. Nat. Cell Biol. 21, 1436–1448 (2019).
Castello, A. et al. Insights into RNA biology from an atlas of mammalian mRNA-binding proteins. Cell 149, 1393–1406 (2012).
Vuzman, D. & Levy, Y. Intrinsically disordered regions as affinity tuners in protein–DNA interactions. Mol. Biosyst. 8, 47–57 (2012).
Liang, H. et al. PTENβ is an alternatively translated isoform of PTEN that regulates rDNA transcription. Nat. Commun. 8, 14771 (2017).
Zhang, Y. et al. β-Arrestin 2 as an activator of cGAS–STING signaling and target of viral immune evasion. Nat. Commun. 11, 6000 (2020).
Longhi, S. Nucleocapsid structure and function. Curr. Top. Microbiol. Immunol. 329, 103–128 (2009).
Malik, Y. A. Properties of coronavirus and SARS-CoV-2. Malays. J. Pathol. 42, 3–11 (2020).
Wang, P. et al. PTENα modulates CaMKII signaling and controls contextual fear memory and spatial learning. Cell Rep. 19, 2627–2641 (2017).
Zhang, Q. et al. PTENε suppresses tumor metastasis through regulation of filopodia formation. EMBO J. 40, e105806 (2021).
Hafner, M. et al. PAR-CliP—a method to identify transcriptome-wide the binding sites of RNA binding proteins. J. Vis. Exp. 2010, 2034 (2010).
Hafner, M. et al. Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell 141, 129–141 (2010).
Kincaid, R. P., Lam, V. L., Chirayil, R. P., Randall, G. & Sullivan, C. S. RNA triphosphatase DUSP11 enables exonuclease XRN-mediated restriction of hepatitis C virus. Proc. Natl Acad. Sci. USA 115, 8197–8202 (2018).
Burke, J. M., Kincaid, R. P., Nottingham, R. M., Lambowitz, A. M. & Sullivan, C. S. DUSP11 activity on triphosphorylated transcripts promotes Argonaute association with noncanonical viral microRNAs and regulates steady-state levels of cellular noncoding RNAs. Genes Dev. 30, 2076–2092 (2016).
Wang, Y. et al. LIMD1 phase separation contributes to cellular mechanics and durotaxis by regulating focal adhesion dynamics in response to force. Dev. Cell 56, 1313–1325 (2021).
Tang, Q. et al. Structure of the receptor-activated human TRPC6 and TRPC3 ion channels. Cell Res. 28, 746–755 (2018).
Xie, M. et al. TREM2 interacts with TDP-43 and mediates microglial neuroprotection against TDP-43-related neurodegeneration. Nat. Neurosci. 25, 26–38 (2022).
Zhao, Y. et al. SARS-CoV-2 spike protein interacts with and activates TLR41. Cell Res. 31, 818–820 (2021).
Laudenbach, B. T. et al. NUDT2 initiates viral RNA degradation by removal of 5′-phosphates. Nat. Commun. 12, 6918 (2021).
Acknowledgements
This work was supported by grants including the National Natural Science Foundation of China (82022032, 81991505, 82221003 and 82171826 to D. L.), Beijing Natural Science Foundation (L222017 to D.L.) and Clinical Medicine Plus X-Young Scholars Project, Peking University (PKU2023LCXQ024 to D. L.).
Author information
Authors and Affiliations
Contributions
Yue Yin, Z.Y. and D.L. conceived and designed the experiments. Yuxin Yin provided the transgenic mice. Yue Yin, Z.Y. and Y.S. performed most of the experiments and analyzed the data. Y. Yang, W.T., Y.Q., Yuxin Yin and F.Y. assisted in some experiments. X. Zhao and X. Zhang performed MS and bioinformatic analyses. Yue Yin, Z.Y., F.Y. and D.L. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Anna Salvetti and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 PTENα selectively interacts with LCMV-NP.
a, Co-immunoprecipitation analysis of HEK293T cells co-transfected with GFP-LCMV-NP as well as FLAG-PTEN, FLAG-PTENα or empty vector, respectively; lysates were immunoprecipitated with anti-FLAG antibody, subsequently analyzed by immunoblot with anti-GFP antibody. GAPDH was used as a loading control. b, Flow diagram of PAR-CliP assay. c, Phosphorimage of an SDS-gel resolving radiolabeled RNA cross-linked to FLAG-tagged Mock, FLAG-PTENα, or FLAG-IGF2BP3 protein in PAR-CliP assay. d–f, Coomassie-stained SDS-PAGE of the recombinant PTENα (d), PTENα-GFP (e), and NP (f). Data were representative of two independent experiments.
Extended Data Fig. 2 PTENα binds to VSV RNA around the N terminus.
a, Predicted PTENα binding motifs at the position 145–1281 nt of VSV-GFP genomes are highlighted with red. b, Enrichment analysis of PTENα-associated mRNAs in HEK293T cells using GO datasets. c, HEK293T cells were transfected with empty vector, PTENα, or PTENα in combination with mutated VSV-NP (ATG to TAA) vector. Cell lysates were immunoprecipitated with anti-FLAG M2 beads and the retrieved HSP90AB1 mRNA was detected by RT-qPCR assay (n = 3 cell cultures). Data were representative of two independent experiments (c). Each dot in bar plots represents one biological sample that is independently treated and assessed at the same time. Statistical significance was assessed by one-sided hypergeometric test (b) or one-way ANOVA followed by Tukey’s multiple comparisons test (c).
Extended Data Fig. 3 Overexpression of PTENα promoted viral infection.
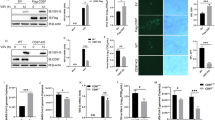
a, Western blot analysis of PTENα expression in PC3 cell line. GAPDH was used as a loading control. b–d, PC3 cells stably expressing PTENα or empty vector were infected with VSV-GFP (MOI = 0.1) for 12 hours. b, c, Percentages of virus-infected cells (GFP+) were measured by flow cytometry (n = 3 cell cultures). d, Transcriptional levels of VSV-GFP normalized by GAPDH mRNA were assessed by RT-qPCR assay (n = 3 cell cultures). Fold change represents the ratio compared to specific value under a single condition. e, Expression levels of the reconstituted proteins in PTEN-deficient Hela cells reconstituted with PTENα and the endogenous proteins were determined by western blot. GAPDH was used as a loading control. Statistical significance was assessed by two-tailed unpaired Student’s t test (c, d). Each dot in bar plots represents one biological sample that is independently treated and assessed at the same time. Data were representative of two independent experiments.
Extended Data Fig. 4 PTENα selectively blocks RIG-I-mediated immune recognition and facilitates viral replication.
a, WT and Ptenα−/− BMDMs were treated with VSV-GFP (MOI = 0.01) for 8 hours or not, then RT-qPCR assay was performed (n = 3 cell cultures). Fold change represents the ratio compared to specific value under a single condition. b, HEK293T cells were transfected with Mock, PTENα, or PTENα-C297S plasmids in combination with vector coding IFNβ (left), MAVS (middle), or RIG-I (right), respectively. Meanwhile, HEK293T cells transfected with an equal amount of Mock plasmid were used as control. After VSV-GFP infection, the percentages of GFP-positive cells were assessed by flow cytometry assay (n = 3 cell cultures). Statistical significance was assessed by two-tailed unpaired Student’s t test (a) or one-way ANOVA followed by Tukey’s multiple comparisons test (b). Each dot in bar plots represents one biological sample that is independently treated and assessed at the same time. Data were representative of two independent experiments.
Extended Data Fig. 5 Loss of PTENα augments peripheral immune response against RNA virus.
a–e, Ptenα−/− and WT mice were infected with VSV. a, Survival rates of Ptenα−/− mice (n = 10) and WT mice (n = 12) were monitored over time. Gross examination (b) and H&E staining (c) of the infected lungs on day 4 post infection. Scale bar, 500 μm. d, RT-qPCR analysis of VSV RNA levels in lungs from the infected mice on day 4 post infection (n = 4 mice). e, Viral titers in lung tissues were determined by TCID50 calculation on day 1 post viral infection (n = 4 mice). f, Ptenα−/− and WT mice were intraperitoneally injected with 5 × 106 PFU VSV. 12 hours later, the peritoneal macrophages from viral infected mice and untreated mice were collected, and the expression levels of ISGs including Ifn-b1, Irf7, Isg56 and Isg15 were measured by RT-qPCR assay (n = 4 mice). g, Ptenα−/− and WT mice were intraperitoneally injected with SeV. 12 hours later, the peritoneal macrophages from viral infected and untreated mice were collected, and the expression levels of ISGs including Ifn-b1, Irf7 and Oas2 were measured by RT-qPCR assay (n = 4 mice). Each dot in bar plots represents one biological sample that is independently treated and assessed at the same time. Fold change in RT-qPCR assay represents the ratio compared to specific value under a single condition. Statistical significance was assessed by log-rank (Mantel-Cox) test (a) or two-tailed unpaired Student’s t test (d–g). UT, untreated control. Data were representative of three independent experiments (b–g) or pooled from two independent experiments (a).
Supplementary information
Supplementary Information
Supplementary Figs. 1–6, Table 1 and caption for Supplementary Video 1 and source data for the Supplementary figures.
Supplementary Video 1a
Phenotypes of WT mice infected with LCMV-ARM were recorded.
Supplementary Video 1b
Remarkable encephalitis-like symptoms, including limbic seizure and paralysis, were observed in Ptenα–/– mice compared to WT mice after exposure to LCMV-ARM.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 3
Unprocessed western blots.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 5
Unprocessed gel.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Unprocessed western blots.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed western blots.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yin, Y., Yang, Z., Sun, Y. et al. RNA-binding protein PTENα blocks RIG-I activation to prevent viral inflammation. Nat Chem Biol (2024). https://doi.org/10.1038/s41589-024-01621-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41589-024-01621-5