Abstract
The C-terminal to LisH (CTLH) complex is a ubiquitin ligase complex that recognizes substrates with Pro/N-degrons via its substrate receptor Glucose-Induced Degradation 4 (GID4), but its function and substrates in humans remain unclear. Here, we report PFI-7, a potent, selective and cell-active chemical probe that antagonizes Pro/N-degron binding to human GID4. Use of PFI-7 in proximity-dependent biotinylation and quantitative proteomics enabled the identification of GID4 interactors and GID4-regulated proteins. GID4 interactors are enriched for nucleolar proteins, including the Pro/N-degron-containing RNA helicases DDX21 and DDX50. We also identified a distinct subset of proteins whose cellular levels are regulated by GID4 including HMGCS1, a Pro/N-degron-containing metabolic enzyme. These data reveal human GID4 Pro/N-degron targets regulated through a combination of degradative and nondegradative functions. Going forward, PFI-7 will be a valuable research tool for investigating CTLH complex biology and facilitating development of targeted protein degradation strategies that highjack CTLH E3 ligase activity.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The mass spectrometry data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD038487. The mass spectrometry data of the chemoproteomics experiment have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD044977. The structure of PFI-7 bound to GID4 was deposited to the PDB with accession number 7SLZ. Source data are provided with this paper.
Code availability
R scripts for analysis of proteomics data are freely available at https://github.com/d0minicO/GID4_analysis.
References
Varshavsky, A. The ubiquitin system, autophagy, and regulated protein degradation. Annu. Rev. Biochem. 86, 123–128 (2017).
Varshavsky, A. N-degron and C-degron pathways of protein degradation. Proc. Natl Acad. Sci. USA 116, 358–366 (2019).
Sherpa, D., Chrustowicz, J. & Schulman, B. A. How the ends signal the end: regulation by E3 ubiquitin ligases recognizing protein termini. Mol. Cell 82, 1424–1438 (2022).
Mészáros, B., Kumar, M., Gibson, T. J., Uyar, B. & Dosztányi, Z. Degrons in cancer. Sci. Signal 10, eaak9982 (2017).
Rechsteiner, M. & Rogers, S. W. PEST sequences and regulation by proteolysis. Trends Biochem. Sci. 21, 267–271 (1996).
Collins, G. A. & Goldberg, A. L. The logic of the 26S proteasome. Cell 169, 792–806 (2017).
Gonda, D. K. et al. Universality and structure of the N-end rule. J. Biol. Chem. 264, 16700–16712 (1989).
Schapira, M., Calabrese, M. F., Bullock, A. N. & Crews, C. M. Targeted protein degradation: expanding the toolbox. Nat. Rev. Drug Discov. 18, 949–963 (2019).
Chen, S.-J., Wu, X., Wadas, B., Oh, J.-H. & Varshavsky, A. An N-end rule pathway that recognizes proline and destroys gluconeogenic enzymes. Science 355, eaal3655 (2017).
Hämmerle, M. et al. Proteins of newly isolated mutants and the amino-terminal proline are essential for ubiquitin-proteasome-catalyzed catabolite degradation of fructose-1,6-bisphosphatase of Saccharomyces cerevisiae. J. Biol. Chem. 273, 25000–25005 (1998).
Dong, C. et al. Molecular basis of GID4-mediated recognition of degrons for the Pro/N-end rule pathway article. Nat. Chem. Biol. 14, 466–473 (2018).
Santt, O. et al. The yeast GID complex, a novel ubiquitin ligase (E3) involved in the regulation of carbohydrate metabolism. Mol. Biol. Cell 19, 3323–3333 (2008).
Francis, O., Han, F. & Adams, J. C. Molecular phylogeny of a RING E3 ubiquitin ligase, conserved in eukaryotic cells and dominated by homologous components, the muskelin/RanBPM/CTLH complex. PLoS ONE 8, e75217 (2013).
Maitland, M. E. R., Lajoie, G. A., Shaw, G. S. & Schild-Poulter, C. Structural and functional insights into GID/CTLH E3 ligase complexes. Int. J. Mol. Sci. 23, 5863 (2022).
Melnykov, A., Chen, S.-J. & Varshavsky, A. Gid10 as an alternative N-recognin of the Pro/N-degron pathway. Proc. Natl Acad. Sci. USA 116, 15914–15923 (2019).
Qiao, S. et al. Interconversion between anticipatory and active GID E3 ubiquitin ligase conformations via metabolically driven substrate receptor assembly. Mol. Cell 77, 150–163.e9 (2020).
Kong, K.-Y. E. et al. Timer-based proteomic profiling of the ubiquitin-proteasome system reveals a substrate receptor of the GID ubiquitin ligase. Mol. Cell 81, 2460–2476.e11 (2021).
Mohamed, W. I. et al. The human GID complex engages two independent modules for substrate recruitment. EMBO Rep. 22, e52981 (2021).
Huffman, N., Palmieri, D. & Coppola, V. The CTLH complex in cancer cell plasticity. J. Oncol. 2019, 4216750 (2019).
Lampert, F. et al. The multi-subunit GID/CTLH E3 ubiquitin ligase promotes cell proliferation and targets the transcription factor Hbp1 for degradation. eLife 7, e35528 (2018).
Leal-Esteban, L. C., Rothé, B., Fortier, S., Isenschmid, M. & Constam, D. B. Role of bicaudal C1 in renal gluconeogenesis and its novel interaction with the CTLH complex. PLoS Genet. 14, e1007487 (2018).
Maitland, M. E. R. et al. The mammalian CTLH complex is an E3 ubiquitin ligase that targets its subunit muskelin for degradation. Sci. Rep. 9, 9864 (2019).
Zhen, R. et al. Wdr26 regulates nuclear condensation in developing erythroblasts. Blood 135, 208–219 (2020).
McTavish, C. et al. Regulation of c-Raf stability through the CTLH Complex. Int. J. Mol. Sci. 20, 934 (2019).
Maitland, M. E. R., Kuljanin, M., Wang, X., Lajoie, G. A. & Schild-Poulter, C. Proteomic analysis of ubiquitination substrates reveals a CTLH E3 ligase complex-dependent regulation of glycolysis. FASEB J. 35, e21825 (2021).
Dong, C. et al. Recognition of nonproline N-terminal residues by the Pro/N-degron pathway. Proc. Natl Acad. Sci. USA 117, 14158–14167 (2020).
Chrustowicz, J. et al. Multifaceted N-degron recognition and ubiquitylation by GID/CTLH E3 ligases. J. Mol. Biol. 434, 167347 (2022).
Oughtred, R. et al. The BioGRID database: a comprehensive biomedical resource of curated protein, genetic, and chemical interactions. Protein Sci. 30, 187–200 (2021).
Chana, C. K. et al. Discovery and structural characterization of small molecule binders of the human CTLH E3 ligase subunit GID4. J. Med. Chem. 65, 12725–12746 (2022).
Yazdi, A. K. et al. Chemical tools for the Gid4 subunit of the human E3 ligase C-terminal to LisH (CTLH) degradation complex. RSC Med. Chem. 15, 1066–1071 (2024).
Sherpa, D. et al. GID E3 ligase supramolecular chelate assembly configures multipronged ubiquitin targeting of an oligomeric metabolic enzyme. Mol. Cell 81, 2445–2459.e13 (2021).
Coyaud, E. et al. BioID-based identification of Skp cullin F-box (SCF)β-TrCP1/2 E3 ligase substrates. Mol. Cell Proteomics 14, 1781–1795 (2015).
Go, C. D. et al. A proximity-dependent biotinylation map of a human cell. Nature 595, 120–124 (2021).
Arrowsmith, C. H. et al. The promise and peril of chemical probes. Nat. Chem. Biol. 11, 536–541 (2015).
Frye, S. V. The art of the chemical probe. Nat. Chem. Biol. 6, 159–161 (2010).
Blagg, J. & Workman, P. Choose and use your chemical probe wisely to explore cancer biology. Cancer Cell 32, 268–270 (2017).
Vu, V., Szewczyk, M. M., Nie, D. Y., Arrowsmith, C. H. & Barsyte-Lovejoy, D. Validating small molecule chemical probes for biological discovery. Annu. Rev. Biochem. 91, 61–87 (2022).
Huttlin, E. L. et al. Dual proteome-scale networks reveal cell-specific remodeling of the human interactome. Cell 184, 3022–3040.e28 (2021).
Cho, N. H. et al. OpenCell: endogenous tagging for the cartography of human cellular organization. Science 375, eabi6983 (2022).
Onea, G., Maitland, M. E. R., Wang, X., Lajoie, G. A. & Schild-Poulter, C. Distinct nuclear and cytoplasmic assemblies and interactomes of the mammalian CTLH E3 ligase complex. J. Cell Sci. 135, jcs259638 (2022).
Kim, D. I. et al. Probing nuclear pore complex architecture with proximity-dependent biotinylation. Proc. Natl Acad. Sci. USA 111, E2453–E2461 (2014).
Gingras, A.-C., Abe, K. T. & Raught, B. Getting to know the neighborhood: using proximity-dependent biotinylation to characterize protein complexes and map organelles. Curr. Opin. Chem. Biol. 48, 44–54 (2019).
Calo, E. et al. RNA helicase DDX21 coordinates transcription and ribosomal RNA processing. Nature 518, 249–253 (2015).
Zhang, Y., Forys, J. T., Miceli, A. P., Gwinn, A. S. & Weber, J. D. Identification of DHX33 as a mediator of rRNA synthesis and cell growth. Mol. Cell. Biol. 31, 4676–4691 (2011).
Gaspar, V. P. et al. Interactome analysis of KIN (Kin17) shows new functions of this protein. Curr. Issues Mol. Biol. 43, 767–781 (2021).
Chen, S.-J., Kim, L., Song, H. K. & Varshavsky, A. Aminopeptidases trim Xaa-Pro proteins, initiating their degradation by the Pro/N-degron pathway. Proc. Natl Acad. Sci. USA 118, e2115430118 (2021).
Mullen, P. J., Yu, R., Longo, J., Archer, M. C. & Penn, L. Z. The interplay between cell signalling and the mevalonate pathway in cancer. Nat. Rev. Cancer 16, 718–731 (2016).
Hondele, M. et al. DEAD-box ATPases are global regulators of phase-separated organelles. Nature 573, 144–148 (2019).
Hansen, S. R., Aderounmu, A. M., Donelick, H. M. & Bass, B. L. Dicer’s helicase domain: a meeting place for regulatory proteins. Cold Spring Harb. Symp. Quant. Biol. 84, 185–193 (2019).
Gregory, R. I. et al. The Microprocessor complex mediates the genesis of microRNAs. Nature 432, 235–240 (2004).
Abdelhaleem, M., Maltais, L. & Wain, H. The human DDX and DHX gene families of putative RNA helicases. Genomics 81, 618–622 (2003).
Georges, A., Marcon, E., Greenblatt, J. & Frappier, L. Identification and characterization of USP7 targets in cancer cells. Sci. Rep. 8, 15833 (2018).
Komander, D. & Rape, M. The ubiquitin code. Annu. Rev. Biochem. 81, 203–229 (2012).
Nguyen, A. T. et al. UBE2O remodels the proteome during terminal erythroid differentiation. Science 357, eaan0218 (2017).
Mashtalir, N. et al. Autodeubiquitination protects the tumor suppressor BAP1 from cytoplasmic sequestration mediated by the atypical ubiquitin ligase UBE2O. Mol. Cell 54, 392–406 (2014).
Havugimana, P. C. et al. A census of human soluble protein complexes. Cell 150, 1068–1081 (2012).
Kabsch, W. XDS. Acta Crystallogr. Sect. D. 66, 125–132 (2010).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. Sec. D. 53, 240–255 (1997).
Bricogne, G. et al. BUSTER v.2.9 (Global Phasing Ltd, 2010).
Smart, O. S. et al. GRADE v.1.102 (Global Phasing Ltd, 2011).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D. Biol. Crystallogr. 60, 2126–2132 (2004).
Williams, C. J. et al. MolProbity: more and better reference data for improved all-atom structure validation. Protein Sci. 27, 293–315 (2018).
Gräslund, S., Savitsky, P. & Müller-Knapp, S. In vivo biotinylation of antigens in E. coli. Methods Mol. Biol. 1586, 337–344 (2017).
Schwalm, M. P. et al. A toolbox for the generation of chemical probes for baculovirus IAP repeat containing proteins. Front Cell Dev. Biol. 10, 886537 (2022).
Rafiee, M.-R. et al. Protease-resistant streptavidin for interaction proteomics. Mol. Syst. Biol. 16, e9370 (2020).
Quast, J. P., Schuster, D. & Picotti, P. protti: an R package for comprehensive data analysis of peptide- and protein-centric bottom-up proteomics data. Bioinformatics Adv. 2, vbab041 (2021).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Durinck, S., Spellman, P. T., Birney, E. & Huber, W. Mapping identifiers for the integration of genomic datasets with the R/Bioconductor package biomaRt. Nat. Protoc. 4, 1184–1191 (2009).
Zhang, X. et al. Proteome-wide identification of ubiquitin interactions using UbIA-MS. Nat. Protoc. 13, 530–550 (2018).
Acknowledgements
This work was supported by Mitacs Elevate Postdoctoral Fellowship to D.D.G.O., funding from the Canadian Institutes for Health Research (grant nos. MOP-142414 and PJT-169101 to C.S.-P.; FDN154328 to C.H.A.), Natural Sciences and Engineering Research Council of Canada grant nos. RGPIN-2021-02728 to J.M. and RGPIN-2021-03435 to D.B.L. and a Cancer Research Society grant no. 25418 to D.B.-L. M.P.S. is funded by the Deutsche Forschungsgemeinschaft CRC1430 (Project ID 424228829). V.R. and M.G. have received support from the EU/EFPIA/OICR/McGill/KTH/Diamond Innovative Medicines Initiative 2 Joint Undertaking (EUbOPEN grant no. 875510). We thank M. Robers and K. Riching from Promega for advising on the NanoBRET and target engagement assays. This research used resources of the Advanced Photon Source, a US Department of Energy (DOE) Office of Science user facility operated for the DOE Office of Science by Argonne National Laboratory under contract no. DE-AC02-06CH11357. Mass spectrometry analyses were performed on equipment funded by a grant from the Canada Foundation for Innovation to G.A.L. The Structural Genomics Consortium is a registered charity (no. 1097737) that receives funds from Bayer AG, Boehringer Ingelheim, Bristol Myers Squibb, Genentech, Genome Canada through the Ontario Genomics Institute (OGI-196), EU/EFPIA/OICR/McGill/KTH/Diamond Innovative Medicines Initiative 2 Joint Undertaking (EUbOPEN grant no. 875510), Janssen, Merck KGaA (also known as EMD in Canada and the United States), Pfizer and Takeda.
Author information
Authors and Affiliations
Contributions
M.E.R.M., D.D.G.O., X.W., E.C.A., M.M.S., R.A.C.M., P.L., D.B.-L. and C.S.-P. designed and performed cellular experiments. M.E.R.M. and G.A.L. designed and performed proteomic experiments. D.D.G.O. conducted proteomic data analysis. V.R. and M.G. designed, performed and analyzed the chemoproteomic experiments. N.B., M.P.S. and S.K. designed and performed GID4-tracer NanoBRET assay and generated NB716. A.K.Y. and M.V. designed and performed biophysical experiments. M.F.C., M.S.D., J.L., J.I.M., T.N.O., D.R.O., C.S. and F.W. designed and synthesized compounds. X.S., C.D., J.M. and A.D. performed crystallography studies and solved structures. C.H.A., M.V., J.M., D.B.-L., M.S., G.A.L. and C.S.-P. supervised research. C.H.A., D.B.-L., C.S.-P. and J.M. provided funding. C.H.A., D.B.-L., M.E.R.M. and D.D.G.O. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
M.F.C., M.S.D., J.L., J.I.M., T.N.O., C.S., F.W. and D.R.O. are or were employees of Pfizer and some of the authors are shareholders in Pfizer Inc. After submission, D.D.G.O. became an employee of Amphista Therapeutics, a company that is developing targeted protein degradation therapeutic platforms. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Nico Dissmeyer, Chao Xu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Enrichment profiling of NB716.
The volcano plot shows the proteins enriched by streptavidin affinity purification in the presence of NB716. GID4 is among the most highly enriched proteins. Many known interaction partners shown in blue are also highly enriched. The differential abundance was calculated and significance was determined using a two-tailed moderated t-test with multiple testing correction using the method of Benjamini-Hochberg. Data are from three independent experiments (n = 3).
Extended Data Fig. 2 Cytotoxic profile of PFI-7.
Cell growth curves for HCT116, HEK293T and U2OS cells treated with increasing concentrations of PFI-7 or PFI-E3H1 are shown over three days. Data are from three independent experiments (n = 3).
Extended Data Fig. 3 Characterization of GID4-BioID2 cell line.
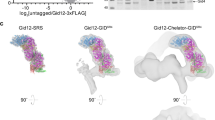
a) Upper, schematic and Alphafold2 predicted structure of GID4-BioID2 fusion protein. Lower, partial CTLH complex structure (PDB 7NSC) showing GID4 C-terminal anchor interacting with ARMC8α (Ref. 31 – Sherpa et al, 2021). b) Immunoprecipitation experiments performed in HeLa cells. Left, HA-pull down of BioID2-GID4 fusion protein with immunoblotting to detect CTLH complex members. Right, immunoprecipitation of RanBP9 with immunoblotting to detect BioID2-GID4 (HA) and other CTLH complex members, *=heavy chain. Data are from one experiment (n = 1). c) Immunoblotting to detect BioID2-GID4 or BioID2 alone after doxycycline induction (top). Bottom, streptavidin-based detection of biotinylated proteins. Representative blot shown from one of five independent experiments (n = 5). d) Immunofluorescence imaging of HA in BioID2 and BioID2-GID4-expressing HeLa cells. HA signal is shown in red, DAPI is shown in blue. Representative images are shown from one of three independent experiments (n = 3). Scale bar: 20 µm. e) Nuclear and cytoplasmic fractionation of HeLa cells expressing BioID2-GID4 or HA-GID4 alone. Immunoblotting of Vinculin, SAP62, RanBP9, and HA is shown. Representative blot shown from one of two independent experiments (n = 23). f) Uniform Manifold Approximation and Projection (UMAP) analysis of proximity-dependent biotinylation samples. BioID2 and BioID2-GID4 cells are shown in gray and blue, respectively. MG132-treated samples are indicated as triangles, and samples treated with vehicle (DMSO) are shown as circles. UMAP was done on 196 high confidence GID4 interacting proteins (MG132 SP > 0.9). Data are from three independent experiments (n = 3, BioID2 + MG132) or four independent experiments (n = 4, BioID2 +DMSO, BioID2-GID4 + DMSO, BioID2-GID4 + MG132). g) Spectral counts of CTLH complex members detected in BioID2-GID4. Bar height represents mean and error bars indicate standard deviation. Data are from three independent experiments (n = 3, BioID2 + MG132) or four independent experiments (n = 4, BioID2 +DMSO, BioID2-GID4 + DMSO, BioID2-GID4 + MG132). h) Overlap between high confidence GID4 interactors and interactors of the CTLH complex present in the BioGRID database.
Extended Data Fig. 4 DDX50 and DDX21 association with GID4 and CTLH complex.
a) Immunoprecipitation of FLAG-GID4 in HEK293T cells expressing FLAG-GID4 treated with 1 µM PFI-7N or PFI-7 for 24 hours. Data are from two experiments (n = 2). b) Immunoprecipitation of RanBP9 in wild type (WT) and ARMC8 knock-out (ARMC8 KO) HeLa cells. Immunoblotting was done for DDX21/50, FLAG, RanBP9, Muskelin and ARMC8α. Data are from one experiment (n = 1). c) Confocal imaging analysis of U2OS cells transduced with a doxycycline inducible lentiviral expression vector coding for N terminally FLAG-tagged GID4. FLAG-GID4 expression was induced for 24 h and is shown in green, DDX21/50 (antibody recognizes both endogenous proteins) is shown in red, and DAPI is shown in blue. Scale bar represents 15 µm. Representative images are shown from four independent experiments. d) Quantification of fluorescence signal intensity over a cross section of cell nuclei. A rolling average with a window size of 8 was used to smooth data and maximum intensities were scaled to 1. One cell from each condition was quantified from one of four independent experiments.
Extended Data Fig. 5 Proteomic samples used and overlap with other datasets.
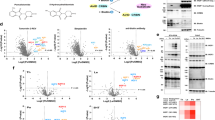
a) Proteomics samples with number of replicates and treatment conditions shown. Data are from four independent experiments (n = 4). These replicate numbers apply to all panels of the figure. b) Number of quantified proteins detected in each sample. Bar height represents mean values and error bars indicate standard deviation. N = 4 independent experiments. c) Protein abundances of CTLH complex members. Median-centered log2 intensities are shown, with blue dots indicating DMSO-treated samples and PFI-7-treated samples shown in red. Boxplot midline indicates median values, bounds of the box indicate 25th and 75th percentiles, and maxima and minima indicate the largest point above or below 1.5 * interquartile range. N = 4 independent experiments. d) Overlap between proteins significantly changed after PFI-7 treatment or GID4 overexpression and medium confidence GID4 interactors (SP > 0.6 in any condition), left. Right, PFI-7-dependent and GID4-dependent proteins overlap with Pro/N-degron-containing proteins. SP scores derived from N = 3 independent experiments in BioID2-GID4 data, protein abundance change significance derived from n = 4 independent expression proteomics experiments.
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2, Tables 1 and 2 and Note.
Supplementary Data 1–6
Proteomics data.
Supplementary Data 7
Unprocessed western blots for Supplementary Fig. 2.
Source data
Source Data Fig. 1
SPR and fluorescence polarization data.
Source Data Fig. 2
NanoBRET quantification of GID4 with Pro/N-degron, GID4-tracer titration and PFI-7 tracer competition.
Source Data Fig. 4
NanoBRET quantification of GID4 interaction with DDX21 and DDX50.
Source Data Fig. 6
Quantification of HMGCS1 levels by western blot after PFI-7 and cycloheximide treatments.
Source Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 3
Unprocessed western blots.
Source Data Extended Data Fig. 4
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Owens, D.D.G., Maitland, M.E.R., Khalili Yazdi, A. et al. A chemical probe to modulate human GID4 Pro/N-degron interactions. Nat Chem Biol (2024). https://doi.org/10.1038/s41589-024-01618-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41589-024-01618-0