Abstract
Adaptations to new pollinators involve multiple floral traits, each requiring coordinated changes in multiple genes. Despite this genetic complexity, shifts in pollination syndromes have happened frequently during angiosperm evolution. Here we study the genetic basis of floral UV absorbance, a key trait for attracting nocturnal pollinators. In Petunia, mutations in a single gene, MYB-FL, explain two transitions in UV absorbance. A gain of UV absorbance in the transition from bee to moth pollination was determined by a cis-regulatory mutation, whereas a frameshift mutation caused subsequent loss of UV absorbance during the transition from moth to hummingbird pollination. The functional differences in MYB-FL provide insight into the process of speciation and clarify phylogenetic relationships between nascent species.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Grant, V. Pollination systems as isolating mechanisms in angiosperms. Evolution 3, 82–97 (1949).
Faegri, K. & van der Pijl, L. The Principles of Pollination Ecology (Pergamon Press, 1979).
Fenster, C.B., Armbruster, W.S., Wilson, P., Dudash, M.R. & Thomson, J.D. Pollination syndromes and floral specialization. Annu. Rev. Ecol. Evol. Syst. 35, 375–403 (2004).
Kay, K. & Schemske, D. Pollinator assemblages and visitation rates for 11 species of neotropical Costus (Costaceae). Biotropica 35, 198–207 (2003).
Gegear, R.J. & Burns, J.G. The birds, the bees, and the virtual flowers: can pollinator behavior drive ecological speciation in flowering plants? Am. Nat. 170, 551–566 (2007).
Whittall, J.B. & Hodges, S.A. Pollinator shifts drive increasingly long nectar spurs in columbine flowers. Nature 447, 706–709 (2007).
Hopkins, R. & Rausher, M.D. Pollinator-mediated selection on flower color allele drives reinforcement. Science 335, 1090–1092 (2012).
Knapp, S. On 'various contrivances': pollination, phylogeny and flower form in the Solanaceae. Phil. Trans. R. Soc. Lond. B 365, 449–460 (2010).
Orr, H.A. The genetic theory of adaptation: a brief history. Nat. Rev. Genet. 6, 119–127 (2005).
Bradshaw, H.D. & Schemske, D.W. Allele substitution at a flower colour locus produces a pollinator shift in monkeyflowers. Nature 426, 176–178 (2003).
Hoballah, M.E. et al. Single gene–mediated shift in pollinator attraction in Petunia. Plant Cell 19, 779–790 (2007).
Hopkins, R. & Rausher, M.D. Identification of two genes causing reinforcement in the Texas wildflower Phlox drummondii. Nature 469, 411–414 (2011).
Klahre, U. et al. Pollinator choice in Petunia depends on two major genetic loci for floral scent production. Curr. Biol. 21, 730–739 (2011).
Yuan, Y.W., Sagawa, J.M., Young, R.C., Christensen, B.J. & Bradshaw, H.D. Jr. Genetic dissection of a major anthocyanin QTL contributing to pollinator-mediated reproductive isolation between sister species of Mimulus. Genetics 194, 255–263 (2013).
Shang, Y. et al. The molecular basis for venation patterning of pigmentation and its effect on pollinator attraction in flowers of Antirrhinum. New Phytol. 189, 602–615 (2011).
Lorenz-Lemke, A.P. et al. Diversification of plant species in a subtropical region of eastern South American highlands: a phylogeographic perspective on native Petunia (Solanaceae). Mol. Ecol. 19, 5240–5251 (2010).
Stehmann, J.R., Lorenz-Lemke, A.P., Freitas, L.B. & Semir, J. in Petunia: Evolutionary, Developmental and Physiological Genetics (eds. Gerats, T. & Strommer, J.) 1–28 (Springer, 2009).
Reck-Kortmann, M. et al. Multilocus phylogeny reconstruction: new insights into the evolutionary history of the genus Petunia. Mol. Phylogenet. Evol. 81, 19–28 (2014).
Gübitz, T., Hoballah, M.E., Dell'Olivo, A. & Kuhlemeier, C. in Petunia: Evolutionary, Developmental and Physiological Genetics (eds. Gerats, T. & Strommer, J.) 29–50 (Springer, 2009).
Wijsman, H.J.W. On the interrelationships of certain species of Petunia. II. Experimental data: crosses between different taxa. Acta Bot. Neerl. 32, 97–107 (1983).
Ando, T. et al. Reproductive isolation in a native population of Petunia sensu Jussieu (Solanaceae). Ann. Bot. (Lond.) 88, 403–413 (2001).
Watanabe, H., Ando, T., Tsukamoto, T., Hashimoto, G. & Marchesi, E. Cross-compatibility of Petunia exserta with other Petunia taxa. J. Jpn. Soc. Hortic. Sci. 70, 33–40 (2001).
Dell'Olivo, A., Hoballah, M.E., Gübitz, T. & Kuhlemeier, C. Isolation barriers between Petunia axillaris and Petunia integrifolia (Solanaceae). Evolution 65, 1979–1991 (2011).
Goldsmith, T.H. Hummingbirds see near ultraviolet light. Science 207, 786–788 (1980).
White, R.H., Brown, P.K., Hurley, A.K. & Bennett, R.R. Rhodopsins, retinula cell ultrastructure, and receptor potentials in the developing pupal eye of the moth Manduca sexta. J. Comp. Physiol. A Neuroethol. Sens. Neural Behav. Physiol. 150, 153–163 (1983).
Peitsch, D. et al. The spectral input systems of hymenopteran insects and their receptor-based colour vision. J. Comp. Physiol. A 170, 23–40 (1992).
Winter, Y., López, J. & Von Helversen, O. Ultraviolet vision in a bat. Nature 425, 612–614 (2003).
Herrera, G. et al. Spectral sensitivities of photoreceptors and their role in colour discrimination in the green-backed firecrown hummingbird (Sephanoides sephaniodes). J. Comp. Physiol. A Neuroethol. Sens. Neural Behav. Physiol. 194, 785–794 (2008).
Kevan, P., Giurfa, M. & Chittka, L. Why are there so many and so few white flowers? Trends Plant Sci. 1, 280–284 (1996).
Guldberg, L.D. & Atsatt, P.R. Frequency of reflection and absorption of ultraviolet light in flowering plants. Am. Midl. Nat. 93, 35–43 (1975).
Chittka, L., Shmida, A., Troje, N. & Menzel, R. Ultraviolet as a component of flower reflections, and the colour perception of hymenoptera. Vision Res. 34, 1489–1508 (1994).
White, R.H., Stevenson, R.D., Bennett, R.R., Cutler, D.E. & Haber, W.A. Wavelength discrimination and the role of ultraviolet vision in the feeding behavior of hawkmoths. Biotropica 26, 427–435 (1994).
Raguso, R.A. & Willis, M.A. Synergy between visual and olfactory cues in nectar feeding by wild hawkmoths, Manduca sexta. Anim. Behav. 69, 407–418 (2005).
Kelber, A. & Hénique, U. Trichromatic colour vision in the hummingbird hawkmoth, Macroglossum stellatarum L. J. Comp. Physiol. A Neuroethol. Sens. Neural Behav. Physiol. 184, 535–541 (1999).
Gould, K.S. & Lister, C. in Flavonoids: Chemistry, Biochemistry and Applications (eds. Andersen, Ø.M. & Markham, K.R.) 397–441 (CRC Press, 2006).
Winkel-Shirley, B. Flavonoid biosynthesis. A colorful model for genetics, biochemistry, cell biology, and biotechnology. Plant Physiol. 126, 485–493 (2001).
Hermann, K. et al. Tight genetic linkage of prezygotic barrier loci creates a multifunctional speciation island in Petunia. Curr. Biol. 23, 873–877 (2013).
Rausher, M.D. Evolutionary transitions in floral color. Int. J. Plant Sci. 169, 7–21 (2008).
Thomson, J.D. & Wilson, P. Explaining evolutionary shifts between bee and hummingbird pollination: convergence, divergence and directionality. Int. J. Plant Sci. 169, 23–38 (2008).
Quattrocchio, F. et al. Molecular analysis of the anthocyanin2 gene of petunia and its role in the evolution of flower color. Plant Cell 11, 1433–1444 (1999).
Dell'Olivo, A. & Kuhlemeier, C. Asymmetric effects of loss and gain of a floral trait on pollinator preference. Evolution 67, 3023–3031 (2013).
Venail, J., Dell'Olivo, A. & Kuhlemeier, C. Speciation genes in the genus Petunia. Phil. Trans. R. Soc. Lond. B 365, 461–468 (2010).
Holton, T.A., Brugliera, F. & Tanaka, Y. Cloning and expression of flavonol synthase from Petunia hybrida. Plant J. 4, 1003–1010 (1993).
Mehrtens, F., Kranz, H., Bednarek, P. & Weisshaar, B. The Arabidopsis transcription factor MYB12 is a flavonol-specific regulator of phenylpropanoid biosynthesis. Plant Physiol. 138, 1083–1096 (2005).
Stracke, R. et al. Differential regulation of closely related R2R3-MYB transcription factors controls flavonol accumulation in different parts of the Arabidopsis thaliana seedling. Plant J. 50, 660–677 (2007).
Quattrocchio, F., Wing, J.F., Leppen, H., Mol, J. & Koes, R.E. Regulatory genes controlling anthocyanin pigmentation are functionally conserved among plant species and have distinct sets of target genes. Plant Cell 5, 1497–1512 (1993).
Davies, K.M. et al. Enhancing anthocyanin production by altering competition for substrate between flavonol synthase and dihydroflavonol 4-reductase. Euphytica 131, 259–268 (2003).
Lorenz-Lemke, A.P. et al. Diversity and natural hybridization in a highly endemic species of Petunia (Solanaceae): a molecular and ecological analysis. Mol. Ecol. 15, 4487–4497 (2006).
Segatto, A.L.A. et al. Nuclear and plastid markers reveal the persistence of genetic identity: a new perspective on the evolutionary history of Petunia exserta. Mol. Phylogenet. Evol. 70, 504–512 (2014).
Pollastri, S. & Tattini, M. Flavonols: old compounds for old roles. Ann. Bot. 108, 1225–1233 (2011).
Rausher, M.D. in The Science of Flavonoids (ed. Grotewold, E.) 175–211 (Springer, 2006).
Cooley, A.M., Modliszewski, J.L., Rommel, M.L. & Willis, J.H. Gene duplication in Mimulus underlies parallel floral evolution via independent trans-regulatory changes. Curr. Biol. 21, 700–704 (2011).
Schwinn, K. et al. A small family of MYB-regulatory genes controls floral pigmentation intensity and patterning in the genus Antirrhinum. Plant Cell 18, 831–851 (2006).
Albert, N.W. et al. A conserved network of transcriptional activators and repressors regulates anthocyanin pigmentation in eudicots. Plant Cell 26, 962–980 (2014).
Streisfeld, M.A., Young, W.N. & Sobel, J.M. Divergent selection drives genetic differentiation in an R2R3-MYB transcription factor that contributes to incipient speciation in Mimulus aurantiacus. PLoS Genet. 9, e1003385 (2013).
Yuan, Y.W., Byers, K.J.R.P. & Bradshaw, H.D. Jr. The genetic control of flower-pollinator specificity. Curr. Opin. Plant Biol. 16, 422–428 (2013).
Edwards, S.V. Is a new and general theory of molecular systematics emerging? Evolution 63, 1–19 (2009).
Yang, Z. & Rannala, B. Molecular phylogenetics: principles and practice. Nat. Rev. Genet. 13, 303–314 (2012).
Dziedzioch, C., Stevens, A.D. & Gottsberger, G. The hummingbird plant community of a tropical montane rain forest in southern Ecuador. Plant Biol. 5, 331–337 (2003).
Cronk, Q. & Ojeda, I. Bird-pollinated flowers in an evolutionary and molecular context. J. Exp. Bot. 59, 715–727 (2008).
Colosimo, P.F. et al. Widespread parallel evolution in sticklebacks by repeated fixation of Ectodysplasin alleles. Science 307, 1928–1933 (2005).
Kronforst, M.R. et al. Linkage of butterfly mate preference and wing color preference cue at the genomic location of wingless. Proc. Natl. Acad. Sci. USA 103, 6575–6580 (2006).
Seehausen, O. et al. Speciation through sensory drive in cichlid fish. Nature 455, 620–626 (2008).
Marchinko, K.B. Predation's role in repeated phenotypic and genetic divergence of armor in threespine stickleback. Evolution 63, 127–138 (2009).
Stuurman, J. et al. Dissection of floral pollination syndromes in Petunia. Genetics 168, 1585–1599 (2004).
Galliot, C., Hoballah, M.E., Kuhlemeier, C. & Stuurman, J. Genetics of flower size and nectar volume in Petunia pollination syndromes. Planta 225, 203–212 (2006).
Hermann, K., Klahre, U., Venail, J., Brandenburg, A. & Kuhlemeier, C. The genetics of reproductive organ morphology in two Petunia species with contrasting pollination syndromes. Planta 241, 1241–1254 (2015).
Bouck, A., Wessler, S.R. & Arnold, M.L. QTL analysis of floral traits in Louisiana iris hybrids. Evolution 61, 2308–2319 (2007).
Brothers, A.N. et al. Genetic architecture of floral traits in Iris hexagona and Iris fulva. J. Hered. 104, 853–861 (2013).
Wessinger, C.A., Hileman, L.C. & Rausher, M.D. Identification of major quantitative trait loci underlying floral pollination syndrome divergence in Penstemon. Phil. Trans. R. Soc. Lond. B 369, 20130349 (2014).
Bradshaw, H.D. Jr., Otto, K.G., Frewen, B.E., McKay, J.K. & Schemske, D.W. Quantitative trait loci affecting differences in floral morphology between two species of monkeyflower (Mimulus). Genetics 149, 367–382 (1998).
Orr, H.A. The population genetics of adaptation: the distribution of factors fixed during adaptive evolution. Evolution 52, 935–949 (1998).
Charlesworth, D. & Charlesworth, B. Selection on recombination in clines. Genetics 91, 581–589 (1979).
Murray, M.G. & Thompson, W.F. Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res. 8, 4321–4325 (1980).
Manly, K.F., Cudmore, R.H. Jr. & Meer, J.M. Map Manager QTX, cross-platform software for genetic mapping. Mamm. Genome 12, 930–932 (2001).
Vandenbussche, M., Zethof, J. & Gerats, T. in Plant Transposable Elements. Methods and Protocols (ed. Peterson, T.) 239–250 (Humana Press, 2013).
Mallona, I., Lischewski, S., Weiss, J., Hause, B. & Egea-Cortines, M. Validation of reference genes for quantitative real-time PCR during leaf and flower development in Petunia hybrida. BMC Plant Biol. 10, 4 (2010).
Andersen, C.L., Jensen, J.L. & Ørntoft, T.F. Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res. 64, 5245–5250 (2004).
Langmead, B. & Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Wu, T.D. & Nacu, S. Fast and SNP-tolerant detection of complex variants and splicing in short reads. Bioinformatics 26, 873–881 (2010).
DePristo, M.A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Grabherr, M.G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Anders, S., Pyl, P.T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Love, M.I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Castel, S.E., Levy-Moonshine, A., Mohammadi, P., Banks, E. & Lappalainen, T. Tools and best practices for data processing in allelic expression analysis. Genome Biol. 16, 195 (2015).
Mayba, O. et al. MBASED: allele-specific expression detection in cancer tissues and cell lines. Genome Biol. 15, 405 (2014).
Acknowledgements
We thank C. Ball, J. Sekulovski, N. Signer and R. Zimmermann for expert care of our plants; M. Saxenhofer and A. Feller for help with genotyping and screening transposon populations; P. Morel and J. Zethof for help with the amplification and sequencing of dTph1 transposon-flanking sequences; A.L. Cazé for help with phenotyping hybrid progeny plants; and A.L. Segatto and C. Turchetto for field collections and DNA extraction. K. Snowden, A. Amrad, K. Hermann, H. Summers and S. Robinson provided constructive comments on manuscript drafts; A. Amrad, R. Koes, J. Stuurman and T. Gerats provided advice and suggestions throughout the project; and R. Köpfli assisted with figures. M.V. was financially supported by an ATIP-AVENIR (CNRS) and ANR-BLANC grant. L.F. was financed by Science without Borders, CNPq. The remaining authors were financed by the National Centre of Competence in Research “Plant Survival” and grants from the Swiss National Science Foundation and the University of Bern.
Author information
Authors and Affiliations
Contributions
Conceptualization: H.S., U.K., L.F. and C.K. Methodology: H.S., M.M., U.K., A.D., M.V. and C.K. Software: M.M. Formal analysis: H.S., M.M. and U.K. Performed all experiments: H.S., M.M., U.K., A.D., K.E., T.M., S.M. and M.V. Resources: M.V., L.F. and C.K. Writing: H.S., M.M., S.M., M.V. and C.K. Visualization: H.S. Funding acquisition: C.K.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
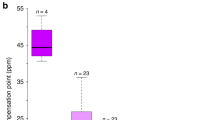
Supplementary Figure 1 UV-absorbance differs significantly between P. axillaris Rio Arapey (Arapey) and P. axillaris S7 (S7) but floral traits important for pollinator attraction do not differ substantially.
(a) UV-absorbance of the corolla limb. (b) Visible absorbance of the corolla limb. (c) Nectar volume. Note that the difference between Arapey and S7 does not affect the time spent feeding Fig. 1e). (d) Quantity of scent (methylbenzoate) emission. Scent had high variability within S7 but this did not affect the overall preference of M. sexta for Arapey flowers (Fig. 1c,d). (e) Surface area of the corolla limb. Bars show mean ± s.d.; for Rio Arapey samples, (a,b,e) n = 5, (c) n = 4, (d) n = 6; for S7 samples, (a-d) n = 4, (e) n = 5. Statistical tests were carried out using a student’s t test and bars with different letters are significantly different (p < 0.05).
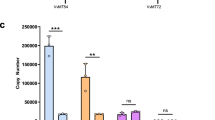
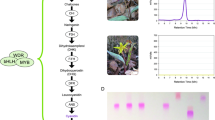
Supplementary Figure 2 Protein sequence alignment of flavonol synthase in P. inflata, P. exserta and P. axillaris, and developmental series of floral buds used for gene expression analyses.
(a) Flavonol synthase (FLS) is a member of the 2-oxoglutarate iron-dependent oxygenase (2-ODD) proteins43. The predicted protein sequence of FLS shows no differences between P. axillaris and P. exserta, and only two residue changes between P. inflata and P. axillaris or P. exserta. Neither residue corresponds to conserved and/or functionally important residues, making differences in enzymatic activity unlikely. Dark grey boxes indicate residues that are conserved between 2-ODD proteins from different plant species87. His34, Asp36 and His290 are required for iron binding, and Arg300 and Ser302 bind the 2-oxoglutarate substrate88. (b,c) Developmental stages of floral buds of P. axillaris (b, top), P. exserta (b, bottom) and P. inflata (c). Numbers define the stages of development, as used for quantitative RT-PCR and RNA-seq experiments. For P. axillaris and P. exserta, stages are separated by approximately 1 d each. For P. inflata, stages 2-4 are separated by 1 to 2 d each, and stages 4-8 are separated by approximately 1 d each. Scale bars, 2 cm.
Supplementary Figure 3 Appearance of various somatic and germline mutants in UV and visible light, including a revertant sector, and details of associated MYB-FL alleles.
(a-d) Four examples of sectors obtained from P. axillaris x W138 F1 (a,c) and IL2-1Pax x W138 F1 (b,d) individuals. Each example is shown in UV light (left) and visible light (right), with UV and visible light absorbance measurements below. (e) A flower from the germline mutant, A1-95 (genotype: MYB-FLPax-dTph4/MYB-FLW138) shown in UV (left) and visible (right) light. (f) The germline mutant, A1-95, was crossed with P. exserta to test for genetic complementation (Fig. 4e) and one of the progeny from this cross that is heterozygous for MYB-FLPax-dTph4 and MYB-FLPex is shown in UV (left) and visible (right) light. This individual has a primarily UV-reflective corolla limb but a UV-absorbent sector is present caused by excision of the dTph4 transposon from the MYB-FLPax-dTph4 allele. (g) DNA was extracted from the sector and the surrounding corolla tissue of the flower shown in f, and the alleles of MYB-FL that were derived from P. axillaris were sequenced from each of these different tissues and are represented in the schematic. The UV-reflective surrounding tissue showed two alleles: allele 1 contained the dTph4 transposon and allele 2 contained a 5 bp footprint left by excision of this transposon (which ultimately causes a frameshift resulting in 21 non-homologous amino acid residues before a stop codon). In the UV-absorbent sector tissue, an allele with a 6 bp footprint due to dTph4 excision was found. This results in the addition of two amino acid residues (phenylalanine and lysine), but the protein sequence remains otherwise intact, allowing the MYB-FLPax-dTph4 allele to return to a functional state after excision of the dTph4 transposon.
Supplementary Figure 4 Protein sequence alignment of MYB111 from Arabidopsis and MYB-FL from P. inflata, P. exserta and P. axillaris.
The N terminus of R2R3-MYBs is highly conserved, with two repeats (R2 and R3; green) which each contain three α helices (yellow) and are involved in DNA-binding, whereas the C terminus is typically less conserved89. The motif that defines subgroup 7 of the R2R3-MYBs in Arabidopsis (AtMYB11, AtMYB12 and AtMYB111; [K/R][R/x][R/K]xGRT[S/x][R/G]xx[M/x]K) is present in Petunia sequences (light blue) with two mismatches (dark blue)90,91. Residues that were conserved in the N-terminus in over 90% of the R2R3-MYBs from Arabidopsis that were analysed are shown in purple90. Only one was not conserved in the Petunia sequences (grey). The positioning of the first two introns (black triangles) is the most common intron pattern in R2R3-MYBs92 and is conserved in the Arabidopsis MYB111 and Petunia MYB-FL sequences. The presence of a third intron (red triangle) is present only in MYB-FL but conserved in all three alleles. The numbers inside the triangles indicate the phase of the introns. Alignment was made using Geneious (Biomatters) with manual modifications.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4, Supplementary Tables 1–6 and Supplementary Note. (PDF 1841 kb)
Rights and permissions
About this article
Cite this article
Sheehan, H., Moser, M., Klahre, U. et al. MYB-FL controls gain and loss of floral UV absorbance, a key trait affecting pollinator preference and reproductive isolation. Nat Genet 48, 159–166 (2016). https://doi.org/10.1038/ng.3462
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3462
This article is cited by
-
Natural variation in Beauty Mark is associated with UV-based geographical adaptation in Gossypium species
BMC Biology (2023)
-
Flower color polymorphism of a wild Iris on the Qinghai-Tibet plateau
BMC Plant Biology (2023)
-
Genetic architecture of a pollinator shift and its fate in secondary hybrid zones of two Petunia species
BMC Biology (2023)
-
R2R3-MYB transcription factors, StmiR858 and sucrose mediate potato flavonol biosynthesis
Horticulture Research (2021)
-
Competition between anthocyanin and kaempferol glycosides biosynthesis affects pollen tube growth and seed set of Malus
Horticulture Research (2021)