Abstract
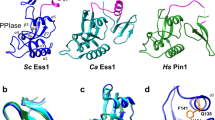
The 5′→3′ exoribonucleases (XRNs) comprise a large family of conserved enzymes in eukaryotes with crucial functions in RNA metabolism and RNA interference1,2,3,4,5. XRN2, or Rat1 in yeast6, functions primarily in the nucleus and also has an important role in transcription termination by RNA polymerase II (refs 7–14). Rat1 exoribonuclease activity is stimulated by the protein Rai1 (refs 15, 16). Here we report the crystal structure at 2.2 Å resolution of Schizosaccharomyces pombe Rat1 in complex with Rai1, as well as the structures of Rai1 and its murine homologue Dom3Z alone at 2.0 Å resolution. The structures reveal the molecular mechanism for the activation of Rat1 by Rai1 and for the exclusive exoribonuclease activity of Rat1. Biochemical studies confirm these observations, and show that Rai1 allows Rat1 to degrade RNAs with stable secondary structure more effectively. There are large differences in the active site landscape of Rat1 compared to related and PIN (PilT N terminus) domain-containing nucleases17,18,19,20. Unexpectedly, we identified a large pocket in Rai1 and Dom3Z that contains highly conserved residues, including three acidic side chains that coordinate a divalent cation. Mutagenesis and biochemical studies demonstrate that Rai1 possesses pyrophosphohydrolase activity towards 5′ triphosphorylated RNA. Such an activity is important for messenger RNA degradation in bacteria21, but this is, to our knowledge, the first demonstration of this activity in eukaryotes and suggests that Rai1/Dom3Z may have additional important functions in RNA metabolism.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Parker, R. & Song, H. The enzymes and control of eukaryotic mRNA turnover. Nature Struct. Mol. Biol. 11, 121–127 (2004)
Newbury, S. F. Control of mRNA stability in eukaryotes. Biochem. Soc. Trans. 34, 30–34 (2006)
Bousquet-Antonelli, C., Presutti, C. & Tollervey, D. Identification of a regulated pathway for nuclear pre-mRNA turnover. Cell 102, 765–775 (2000)
Gatfield, D. & Izaurralde, E. Nonsense-mediated messenger RNA decay is initiated by endonucleolytic cleavage in Drosophila . Nature 429, 575–578 (2004)
Gazzani, S., Lawrenson, T., Woodward, C., Headon, D. & Sablowski, R. A link between mRNA turnover and RNA interference in Arabidopsis . Science 306, 1046–1048 (2004)
Amberg, D. C., Goldstein, A. L. & Cole, C. N. Isolation and characterization of RAT1: an essential gene of Saccharomyces cerevisiae required for the efficient nucleocytoplasmic trafficking of mRNA. Genes Dev. 6, 1173–1189 (1992)
Kim, M. et al. The yeast Rat1 exonuclease promotes transcription termination by RNA polymerase II. Nature 432, 517–522 (2004)
West, S., Gromak, N. & Proudfoot, N. J. Human 5′→3′ exonuclease Xrn2 promotes transcription termination at co-transcriptional cleavage sites. Nature 432, 522–525 (2004)
Connelly, S. & Manley, J. L. A functional mRNA polyadenylation signal is required for transcription termination by RNA polymerase II. Genes Dev. 2, 440–452 (1988)
Luo, W. & Bentley, D. A ribonucleolytic Rat torpedoes RNA polymerase II. Cell 119, 911–914 (2004)
Buratowski, S. Connections between mRNA 3′ end processing and transcription termination. Curr. Opin. Cell Biol. 17, 257–261 (2005)
Luo, W., Johnson, A. W. & Bentley, D. L. The role of Rat1 in coupling mRNA 3′-end processing to transcription termination: implications for a unified allosteric-torpedo model. Genes Dev. 20, 954–965 (2006)
Kaneko, S., Rozenblatt-Rosen, O., Meyerson, M. & Manley, J. L. The multifunctional protein p54nrb/PSF recruits the exonuclease XRN2 to facilitate pre-mRNA 3′ processing and transcription termination. Genes Dev. 21, 1779–1789 (2007)
West, S. & Proudfoot, N. J. Human Pcf11 enhances degradation of RNA polymerase II-associated nascent RNA and transcriptional termination. Nucleic Acids Res. 36, 905–914 (2008)
Stevens, A. & Poole, T. L. 5′-exonuclease-2 of Saccharomyces cerevisiae. Purification and features of ribonuclease activity with comparison to 5′-exonuclease-1. J. Biol. Chem. 270, 16063–16069 (1995)
Xue, Y. et al. Saccharomyces cerevisiae RAI1 (YGL246c) is homologous to human DOM3Z and encodes a protein that binds the nuclear exoribonuclease Rat1p. Mol. Cell. Biol. 20, 4006–4015 (2000)
Mueser, T. C., Nossal, N. G. & Hyde, C. C. Structure of bacteriophage T4 RNase H, a 5′ to 3′ RNA-DNA and DNA-DNA exonuclease with sequence similarity to the RAD2 family of eukaryotic proteins. Cell 85, 1101–1112 (1996)
Devos, J. M., Tomanicek, S. J., Jones, C. E., Nossal, N. G. & Mueser, T. C. Crystal structure of bacteriophage T4 5′ nuclease in complex with a branched DNA reveals how Flap endonuclease-1 family nucleases bind their substrates. J. Biol. Chem. 282, 31713–31724 (2007)
Glavan, F., Behm-Ansmant, I., Izaurralde, E. & Conti, E. Structures of the PIN domains of SMG6 and SMG5 reveal a nuclease within the mRNA surveillance complex. EMBO J. 25, 5117–5125 (2006)
Clissold, P. M. & Ponting, C. P. PIN domains in nonsense-mediated mRNA decay and RNAi. Curr. Biol. 10, R888–R890 (2000)
Deana, A., Celesnik, H. & Belasco, J. G. The bacterial enzyme RppH triggers messenger RNA degradation by 5′ pyrophosphate removal. Nature 451, 355–358 (2008)
Shobuike, T., Tatebayashi, K., Tani, T., Sugano, S. & Ikeda, H. The dhp1+ gene, encoding a putative nuclear 5′→3′ exoribonuclease, is required for proper chromosome segregation in fission yeast. Nucleic Acids Res. 29, 1326–1333 (2001)
Kenna, M., Stevens, A., McCammon, M. & Douglas, M. G. An essential yeast gene with homology to the exonuclease-encoding XRN1/KEM1 gene also encodes a protein with exoribonuclease activity. Mol. Cell. Biol. 13, 341–350 (1993)
Page, A. M., Davis, K., Molineux, C., Kolodner, R. D. & Johnson, A. W. Mutational analysis of exoribonuclease I from Saccharomyces cerevisiae . Nucleic Acids Res. 26, 3707–3716 (1998)
Solinger, J. A., Pascolini, D. & Heyer, W.-D. Active-site mutations in the Xrn1p exoribonuclease of Saccharomyces cerevisiae reveal a specific role in meiosis. Mol. Cell. Biol. 19, 5930–5942 (1999)
Yang, W., Lee, J. Y. & Nowotny, M. Making and breaking nucleic acids: Two-Mg2+-ion catalysis and substrate specificity. Mol. Cell 22, 5–13 (2006)
Lawrence, M. C. & Colman, P. M. Shape complementarity at protein/protein interfaces. J. Mol. Biol. 234, 946–950 (1993)
Poole, T. L. & Stevens, A. Structural modifications of RNA influence the 5′ exoribonucleolytic hydrolysis by XRN1 and HKE1 of Saccharomyces cerevisiae . Biochem. Biophys. Res. Commun. 235, 799–805 (1997)
DeLano, W. L. PyMOL Users Manual (DeLano Scientific, 2002)
Nicholls, A., Sharp, K. A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991)
Doublie, S. et al. Crystallization and preliminary X-ray analysis of the 9 kDa protein of the mouse signal recognition particle and the selenomethionyl-SRP9. FEBS Lett. 384, 219–221 (1996)
Hendrickson, W. A. Determination of macromolecular structures from anomalous diffraction of synchrotron radiation. Science 254, 51–58 (1991)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Weeks, C. M. & Miller, R. The design and implementation of SnB v2.0. J. Appl. Crystallogr. 32, 120–124 (1999)
Terwilliger, T. C. SOLVE and RESOLVE: Automated structure solution and density modification. Methods Enzymol. 374, 22–37 (2003)
Jones, T. A., Zou, J. Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991)
Brunger, A. T. et al. Crystallography & NMR System: A new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
Lamzin, V. S., Perrakis, A. & Wilson, K. S. in Crystallography of Biological Macromolecules. International Tables for Crystallography Vol. F (eds Rossmann, M. G. & Arnold, E.) 720–721 (Kluwer, 2001)
Vagin, A. A. & Teplyakov, A. An approach to multi-copy search in molecular replacement. Acta Crystallogr. D 56, 1622–1624 (2000)
Jiao, X., Wang, Z. & Kiledjian, M. Identification of an mRNA-decapping regulator implicated in X-linked mental retardation. Mol. Cell 24, 713–722 (2006)
Piccirillo, C., Khanna, R. & Kiledjian, M. Functional characterization of the mammalian mRNA decapping enzyme hDcp2. RNA 9, 1138–1147 (2003)
Acknowledgements
We thank R. Abramowitz and J. Schwanof for setting up the X4A and X4C beamlines and H. Robinson for setting up the X29A beamline at the NSLS, and S. Jia for providing the S. pombe genome. This research is supported in part by grants from the NIH to L.T. (GM077175), J.L.M. (GM28983) and M.K. (GM67005).
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Results and a Supplementary Discussion, Supplementary Table 1, Supplementary References and Supplementary Figures 1-12 with Legends. (PDF 5105 kb)
Rights and permissions
About this article
Cite this article
Xiang, S., Cooper-Morgan, A., Jiao, X. et al. Structure and function of the 5′→3′ exoribonuclease Rat1 and its activating partner Rai1. Nature 458, 784–788 (2009). https://doi.org/10.1038/nature07731
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature07731
This article is cited by
-
Ceg1 depletion reveals mechanisms governing degradation of non-capped RNAs in Saccharomyces cerevisiae
Communications Biology (2023)
-
Xrn1 is a deNADding enzyme modulating mitochondrial NAD-capped RNA
Nature Communications (2022)
-
Observation of conformational changes that underlie the catalytic cycle of Xrn2
Nature Chemical Biology (2022)
-
Extensive 5′-surveillance guards against non-canonical NAD-caps of nuclear mRNAs in yeast
Nature Communications (2020)
-
XRN2 promotes EMT and metastasis through regulating maturation of miR-10a
Oncogene (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.