Abstract
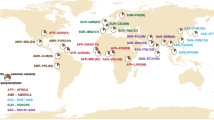
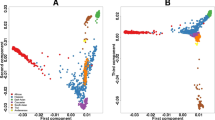
Ethnicity can confound results in pharmacogenomic studies. Allele frequencies of loci that influence drug metabolism can vary substantially between different ethnicities and underlying ancestral genetic differences can lead to spurious findings in pharmacogenomic association studies. We evaluated the application of principal component analysis (PCA) in a pharmacogenomic study in Canada to detect and correct for genetic ancestry differences using genotype data from 2094 loci in 220 key drug biotransformation genes. Using 89 Coriell worldwide reference samples, we observed a strong correlation between principal component values and geographic origin. We further applied PCA to accurately infer the genetic ancestry in our ethnically diverse Canadian cohort of 524 patients from the GATC study of severe adverse drug reactions. We show that PCA can be successfully applied in pharmacogenomic studies using a limited set of markers to detect underlying differences in genetic ancestry thereby maximizing power and minimizing false-positive findings.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 print issues and online access
$259.00 per year
only $43.17 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Abbreviations
- PCA:
-
principal component analysis
- GWAS:
-
genome-wide association study
- ADR:
-
adverse drug reaction
- SNP:
-
single nucleotide polymorphism
- GATC:
-
genotype-specific approaches to therapy in children
References
Gurwitz D, Motulsky AG . Drug reactions, enzymes, and biochemical genetics 50 years later. Pharmacogenomics 2007; 8: 1479–1484.
Ameyaw MM, Regateiro F, Li T, Liu X, Tariq M, Mobarek A et al. MDR1 pharmacogenetics: frequency of the C3435T mutation in exon 26 is significantly influenced by ethnicity. Pharmacogenetics 2001; 11: 217–221.
McLeod HL, Pritchard SC, Githang’a J, Indalo A, Ameyaw MM, Powrie RH et al. Ethnic differences in thiopurine methyltransferase pharmacogenetics: evidence for allele specificity in Caucasian and Kenyan individuals. Pharmacogenetics 1999; 9: 773–776.
Wilson JF, Weale ME, Smith AC, Gratrix F, Fletcher B, Thomas MG et al. Population genetic structure of variable drug response. Nat Genet 2001; 29: 265–269.
Lonjou C, Thomas L, Borot N, Ledger N, de Toma C, LeLouet H et al. A marker for Stevens-Johnson syndrome: ethnicity matters. Pharmacogenomics J 2006; 6: 265–268.
Hindorff L, Junkins H, Mehta J, Manolio TA . Catalog of Published Genome-Wide Association Studies, Available at www.genome.gov/26525384|.
Cooper GM, Johnson JA, Langaee TY, Feng H, Stanaway IB, Schwarz UI et al. A genome-wide scan for common genetic variants with a large influence on warfarin maintenance dose. Blood 2008; 112: 1022–1027.
Link E, Parish S, Armitage J, Bowman L, Heath S, Matsuda F et al. SLCO1B1 variants and statin-induced myopathy—a genomewide study. N Engl J Med 2008; 359: 789–799.
Wellcome Trust Case Control Consortium. Genome-wide association study of 14 000 cases of seven common diseases and 3000 shared controls. Nature 2007; 447: 661–678.
Sladek R, Rocheleau G, Rung J, Dina C, Shen L, Serre D et al. A genome-wide association study identifies novel risk loci for type 2 diabetes. Nature 2007; 445: 881–885.
Pritchard JK, Stephens M, Rosenberg NA, Donnelly P . Association mapping in structured populations. Am J Hum Genet 2000; 67: 170–181.
Balding DJ . A tutorial on statistical methods for population association studies. Nat Rev 2006; 7: 781–791.
Huang RS, Duan S, Shukla SJ, Kistner EO, Clark TA, Chen TX et al. Identification of genetic variants contributing to cisplatin-induced cytotoxicity by use of a genomewide approach. Am J Hum Genet 2007; 81: 427–437.
Ross CJ, Carleton B, Warn DG, Stenton SB, Rassekh SR, Hayden MR . Genotypic approaches to therapy in children: a national active surveillance network (GATC) to study the pharmacogenomics of severe adverse drug reactions in children. Ann N Y Acad Sci 2007; 1110: 177–192.
Statistics Canada. Ethnic Origin and Visible Minorities 2006 Census (2008). Statistics Canada Demography Division: Ottawa.
Suarez-Kurtz G, Vargens DD, Struchiner CJ, Bastos-Rodrigues L, Pena SD . Self-reported skin color, genomic ancestry and the distribution of GST polymorphisms. Pharmacogenet Genomics 2007; 17: 765–771.
Barnholtz-Sloan JS, McEvoy B, Shriver MD, Rebbeck TR . Ancestry estimation and correction for population stratification in molecular epidemiologic association studies. Cancer Epidemiol Biomarkers Prev 2008; 17: 471–477.
Pritchard JK, Stephens M, Donnelly P . Inference of population structure using multilocus genotype data. Genetics 2000; 155: 945–959.
Falush D, Stephens M, Pritchard JK . Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 2003; 164: 1567–1587.
Price AL, Patterson NJ, Plenge RM, Weinblatt ME, Shadick NA, Reich D . Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet 2006; 38: 904–909.
Menozzi P, Piazza A, Cavalli-Sforza L . Synthetic maps of human gene frequencies in Europeans. Science (NY) 1978; 201: 786–792.
Patterson N, Price AL, Reich D . Population structure and eigenanalysis. PLoS Genet 2006; 2: e190.
Novembre J, Johnson T, Bryc K, Kutalik Z, Boyko AR, Auton A et al. Genes mirror geography within Europe. Nature 2008; 456: 98–101, Epub 31 August 2008.
Tian C, Gregersen PK, Seldin MF . Accounting for ancestry: population substructure and genome-wide association studies. Hum Mol Genet 2008; 17: R143–R150.
Nelson MR, Bryc K, King KS, Indap A, Boyko AR, Novembre J et al. The population reference sample, POPRES: a resource for population, disease, and pharmacological genetics research. Am J Hum Genet 2008; 83: 347–358.
Jakobsson M, Scholz SW, Scheet P, Gibbs JR, VanLiere JM, Fung HC et al. Genotype, haplotype and copy-number variation in worldwide human populations. Nature 2008; 451: 998–1003.
Campbell MC, Tishkoff SA . African genetic diversity: implications for human demographic history, modern human origins, and complex disease mapping. Annu Rev Genomics Hum Genet 2008; 9: 403–433.
Singapore Department of Statistics. Population Trends 2008. Department of Statistics: Singapore.
Wang S, Lewis CM, Jakobsson M, Ramachandran S, Ray N, Bedoya G et al. Genetic variation and population structure in native Americans. PLoS Genet 2007; 3: e185.
Li JZ, Absher DM, Tang H, Southwick AM, Casto AM, Ramachandran S et al. Worldwide human relationships inferred from genome-wide patterns of variation. Science (NY) 2008; 319: 1100–1104.
Friedlaender JS, Friedlaender FR, Reed FA, Kidd KK, Kidd JR, Chambers GK et al. The genetic structure of Pacific Islanders. PLoS Genet 2008; 4: e19.
Kimura R, Ohashi J, Matsumura Y, Nakazawa M, Inaoka T, Ohtsuka R et al. Gene flow and natural selection in oceanic human populations inferred from genome-wide SNP typing. Mol Biol Evol 2008; 25: 1750–1761.
Fiji Islands Bureau of Statistics (2008) Census 2007.
Estrela RC, Ribeiro FS, Carvalho RS, Gregorio SP, Dias-Neto E, Struchiner CJ et al. Distribution of ABCB1 polymorphisms among Brazilians: impact of population admixture. Pharmacogenomics 2008; 9: 267–276.
Lao O, Lu TT, Nothnagel M, Junge O, Freitag-Wolf S, Caliebe A et al. Correlation between genetic and geographic structure in Europe. Curr Biol 2008; 18: 1241–1248.
Indian Genome Variation Consortium. Genetic landscape of the people of India: a canvas for disease gene exploration. J Genet 2008; 87: 3–20.
Clayton DG, Walker NM, Smyth DJ, Pask R, Cooper JD, Maier LM et al. Population structure, differential bias and genomic control in a large-scale, case-control association study. Nat Genet 2005; 37: 1243–1246.
Voight BF, Pritchard JK . Confounding from cryptic relatedness in case-control association studies. PLoS Genet 2005; 1: e32.
Carlson CS, Eberle MA, Rieder MJ, Yi Q, Kruglyak L, Nickerson DA . Selecting a maximally informative set of single-nucleotide polymorphisms for association analyses using linkage disequilibrium. Am J Hum Genet 2004; 74: 106–120.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 2007; 81: 559–575.
Acknowledgements
We thank the patients and their families for their participation in the GATC project. We acknowledge the support of the GATC active ADR surveillance network, particularly the site investigators Cheri Nijssen-Jordan, David Johnson, Kevin Hall, Michael Rieder, Shinya Ito, Gideon Koren, Regis Vaillancourt, Pat Elliott-Miller, Jean-Francois Bussières, Denis Lebel, Margaret Murray, Darlene Boliver, Carol Portwine; site surveillance clinicians Linda Verbeek, Rick Kaczowka, Shanna Chan, Becky Malkin, Facundo Garcia, Miho Inoue, Sachi Sakaguchi, Toshihiro Tanaka, Elaine Wong, Brenda Wilson, Pierre Barret, Carol-anne Osborne, Amy Cranston; and research staff at POPI and the CMMT: Anne Smith, Claudette Hildebrand; Lucila Castro, Reza Ghannadan, Catherine Carter, Fudan Miao, Terry Pape and Graeme Honeyman. The study was funded by the Genome Canada, and additional funding was also provided by the Genome British Columbia; the Child & Family Research Institute (Vancouver, BC); the Canadian Institutes of Health Research; Faculties of Pharmaceutical Sciences and Medicine, University of British Columbia; University of Western Ontario; the Canada Gene Cure Foundation; the Canadian Society of Clinical Pharmacology; C17 Research Network: the Childhood Cancer Foundation, Candlelighters Canada; the Canadian Paediatric Society, Merck Frosst; Janssen-Ortho; Illumina; IBM; Eli Lilly; and Pfizer. HV is supported by a postdoctoral research fellowship award from the Michael Smith Foundation for Health Research and the Child and Family Research Institute.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the The Pharmacogenomics Journal website (http://www.nature.com/tpj)
Rights and permissions
About this article
Cite this article
Visscher, H., Ross, C., Dubé, MP. et al. Application of principal component analysis to pharmacogenomic studies in Canada. Pharmacogenomics J 9, 362–372 (2009). https://doi.org/10.1038/tpj.2009.36
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/tpj.2009.36
Keywords
This article is cited by
-
Hybrid autoencoder with orthogonal latent space for robust population structure inference
Scientific Reports (2023)
-
The targetable A1 Huntington disease haplotype has distinct Amerindian and European origins in Latin America
European Journal of Human Genetics (2017)
-
A coding variant in RARG confers susceptibility to anthracycline-induced cardiotoxicity in childhood cancer
Nature Genetics (2015)
-
Applying genome-wide gene-based expression quantitative trait locus mapping to study population ancestry and pharmacogenetics
BMC Genomics (2014)
-
Genetic Markers of Cisplatin-Induced Hearing Loss in Children
Clinical Pharmacology & Therapeutics (2014)