Featured
-
-
Article
| Open Access A molecular staging model for accurately dating the endometrial biopsy
A molecular staging model for accurately dating the endometrial biopsy
Natural variability in menstrual cycle length with rapid changes in gene expression makes it difficult to accurately compare different stages of the endometrial cycle. Here, the authors show a method for precisely determining endometrial cycle stage based on global gene expression that reveals remarkably synchronised daily changes for over 3,400 endometrial genes.
- W. T. Teh
- , J. Chung
- & P. A. W. Rogers
-
Article
| Open Access Targetable NOTCH1 rearrangements in reninoma
Targetable NOTCH1 rearrangements in reninoma
Reninomas are very rare kidney tumours of juxtaglomerular cells. Here, the authors analyse reninomas using whole-genome and transcriptome sequencing, and reveal the presence and functional effects of NOTCH1 rearrangements.
- Taryn D. Treger
- , John E. G. Lawrence
- & Tanzina Chowdhury
-
Article
| Open Access A CCL2+DPP4+ subset of mesenchymal stem cells expedites aberrant formation of creeping fat in humans
A CCL2+DPP4+ subset of mesenchymal stem cells expedites aberrant formation of creeping fat in humans
Extra-intestinal “creeping fat” is a hallmark of Crohn’s disease. Here, using single-cell transcriptomics and lipid metabolomics, the authors identify a subset of mesenchymal stem cells that promote adipogenesis in creeping fat formation.
- Fengfei Wu
- , Fangting Wu
- & Lan Bai
-
Article
| Open Access Circular RNAs in the human brain are tailored to neuron identity and neuropsychiatric disease
Circular RNAs in the human brain are tailored to neuron identity and neuropsychiatric disease
Dopamine neurons control movements while pyramidal neurons regulate memory and language. Here the authors show that circular RNAs production in these neurons appears tailored to neuron identity and genetically linked to neuropsychiatric disease such as Parkinson’s and Alzheimer’s disease.
- Xianjun Dong
- , Yunfei Bai
- & Clemens R. Scherzer
-
Article
| Open Access Divergent single cell transcriptome and epigenome alterations in ALS and FTD patients with C9orf72 mutation
Divergent single cell transcriptome and epigenome alterations in ALS and FTD patients with C9orf72 mutation
Non-coding repeat expansion in the C9ORF72 gene is the most frequent cause of ALS and frontotemporal dementia. Here, the authors performed single cell analyses of gene expression and epigenetic regulation in these patients’ brains and emphasized the role of astrocytes and neurons in neurodegeneration.
- Junhao Li
- , Manoj K. Jaiswal
- & Stella Dracheva
-
Article
| Open Access Single nucleus transcriptomics of ventral midbrain identifies glial activation associated with chronic opioid use disorder
Single nucleus transcriptomics of ventral midbrain identifies glial activation associated with chronic opioid use disorder
The cellular signatures of the opioid exposed human midbrain remain unexplored. Here, authors show by single nuclei transcriptomics activation of the glial immune response and dysregulation of synaptic signaling in opioid exposed individuals
- Julong Wei
- , Tova Y. Lambert
- & Schahram Akbarian
-
Article
| Open Access Cell diversity and plasticity during atrioventricular heart valve EMTs
Cell diversity and plasticity during atrioventricular heart valve EMTs
Lotto et al. delineate cell diversity and mechanisms during heart valve development using scRNA-seq. They identify distinct cell types and states, the emergence of epithelial-mesenchymal plasticity, and cell interactions that may govern this process.
- Jeremy Lotto
- , Rebecca Cullum
- & Pamela A. Hoodless
-
Article
| Open Access Coordination of alternative splicing and alternative polyadenylation revealed by targeted long read sequencing
Coordination of alternative splicing and alternative polyadenylation revealed by targeted long read sequencing
In this study using a targeted long read RNA sequencing approach called PL-Seq, the authors uncover coordination of alternative splicing and alternative polyadenylation within individual genes in Drosophila melanogaster.
- Zhiping Zhang
- , Bongmin Bae
- & Pedro Miura
-
Article
| Open Access Single cell multiomic analysis reveals diabetes-associated β-cell heterogeneity driven by HNF1A
Single cell multiomic analysis reveals diabetes-associated β-cell heterogeneity driven by HNF1A
The mechanism and disease-relevance of pancreatic b-cell heterogeneity remains elusive. Here the authors show that variable HNF1A-FXYD2 activity drives single b-cell heterogeneity at transcriptomic, epigenomic, and electro-physiological levels, which strongly mark the progression of type 2 diabetes.
- Chen Weng
- , Anniya Gu
- & Yan Li
-
Article
| Open Access Functionally distinct cancer-associated fibroblast subpopulations establish a tumor promoting environment in squamous cell carcinoma
Functionally distinct cancer-associated fibroblast subpopulations establish a tumor promoting environment in squamous cell carcinoma
During the progression of cutaneous squamous cell carcinoma (cSCC), dermal fibroblasts become activated into cancer-associated fibroblasts (CAFs) which are pro-tumorigenic. Here, using single-cell RNA sequencing of patients’ samples at different stages of cSCC progression, the authors identify two main CAF subsets and deduce their potential functions using bioinformatics.
- Sabrina Schütz
- , Llorenç Solé-Boldo
- & Frank Lyko
-
Article
| Open Access Notch and retinoic acid signals regulate macrophage formation from endocardium downstream of Nkx2-5
Notch and retinoic acid signals regulate macrophage formation from endocardium downstream of Nkx2-5
A subset of endocardial cells gives rise to hematopoietic cells which are important for the formation of cardiac valves. In this follow up study, authors report the molecular regulatory mechanisms behind the formation of macrophages in the heart.
- Norika Liu
- , Naofumi Kawahira
- & Atsushi Nakano
-
Article
| Open Access A nuclear receptor HR96-related gene underlies large trans-driven differences in detoxification gene expression in a generalist herbivore
A nuclear receptor HR96-related gene underlies large trans-driven differences in detoxification gene expression in a generalist herbivore
Adaptation to toxins in agricultural pests is often caused by increased expression of detoxification genes. Here, the authors reveal that variation in a family of transcriptional regulators facilitates rapid evolution to diverse pesticides and host plants.
- Meiyuan Ji
- , Marilou Vandenhole
- & Thomas Van Leeuwen
-
Article
| Open Access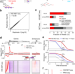 High-sensitive nascent transcript sequencing reveals BRD4-specific control of widespread enhancer and target gene transcription
High-sensitive nascent transcript sequencing reveals BRD4-specific control of widespread enhancer and target gene transcription
Here the authors reveal that high-sensitive nascent transcript sequencing provides an extended high-resolution view on transcription, including lowly transcribed enhancers. Widespread transcription at enhancers and their target genes depends on the BET family protein BRD4.
- Annkatrin Bressin
- , Olga Jasnovidova
- & Andreas Mayer
-
Article
| Open Access Ontogenetically distinct neutrophils differ in function and transcriptional profile in zebrafish
Ontogenetically distinct neutrophils differ in function and transcriptional profile in zebrafish
Neutrophil ontogeny in zebrafish may be a continuum or consist of distinct lineages. Here the authors characterise neutrophils derived from rostral blood island and caudal haematopoietic tissue lineages and show differential gene expression and function in steady state and during wound healing.
- Juan P. García-López
- , Alexandre Grimaldi
- & Carmen G. Feijoo
-
Article
| Open Access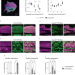 REPTOR and CREBRF encode key regulators of muscle energy metabolism
REPTOR and CREBRF encode key regulators of muscle energy metabolism
Obesity and cancer-induced cachexia are linked to an impairment in the ability of muscle to use glucose or lipids interchangeably as energy substrates. Here, the authors propose that Drosophila REPTOR and its mammalian ortholog CREBRF act as key transcriptional regulators of fuel choice in muscle.
- Pedro Saavedra
- , Phillip A. Dumesic
- & Norbert Perrimon
-
Article
| Open Access Spatial transcriptomics reveal markers of histopathological changes in Duchenne muscular dystrophy mouse models
Spatial transcriptomics reveal markers of histopathological changes in Duchenne muscular dystrophy mouse models
Duchenne muscular dystrophy is a fatal neuromuscular disorder affecting one in 5000 male births. To enrich our understanding of the underlying pathology, the authors apply spatial transcriptomics on dystrophic skeletal muscle to unravel markers related to histopathological changes in Duchenne mouse models.
- L.G.M. Heezen
- , T. Abdelaal
- & P. Spitali
-
Article
| Open Access TEQUILA-seq: a versatile and low-cost method for targeted long-read RNA sequencing
TEQUILA-seq: a versatile and low-cost method for targeted long-read RNA sequencing
The authors report TEQUILA-seq, a versatile, easy-to-implement, and low-cost method for targeted long-read RNA sequencing. TEQUILA-seq uncovers transcript isoforms and RNA mechanisms associated with human health and disease.
- Feng Wang
- , Yang Xu
- & Lan Lin
-
Article
| Open Access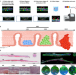 Spatial transcriptomics analysis of esophageal squamous precancerous lesions and their progression to esophageal cancer
Spatial transcriptomics analysis of esophageal squamous precancerous lesions and their progression to esophageal cancer
Understanding the molecular changes in the transition from esophageal squamous precancerous lesions (ESPL) to esophageal squamous cell carcinoma (ESCC) remains essential. Here, the authors analyze ESPL samples using spatial transcriptomics and reveal expression changes in TAGLN2 and CRNN during progression to ESCC.
- Xuejiao Liu
- , Simin Zhao
- & Zigang Dong
-
Article
| Open Access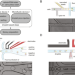 spinDrop: a droplet microfluidic platform to maximise single-cell sequencing information content
spinDrop: a droplet microfluidic platform to maximise single-cell sequencing information content
Droplet microfluidics enables high-throughput single-cell sequencing, but often with increased noise. Here the authors report spinDrop (sorting picoinjection inDrop) to increase gene detection and reduce noise; they use this to generate a high-quality molecular atlas of mouse brain development.
- Joachim De Jonghe
- , Tomasz S. Kaminski
- & Florian Hollfelder
-
Article
| Open Access Single-cell transcriptomics of human cholesteatoma identifies an activin A-producing osteoclastogenic fibroblast subset inducing bone destruction
Single-cell transcriptomics of human cholesteatoma identifies an activin A-producing osteoclastogenic fibroblast subset inducing bone destruction
This study identified a subset of osteoclastogenic fibroblasts expressing INHBA/activin A in human cholesteatoma. It further elucidated the mechanism behind the induction of inflammatory bone destruction, suggesting a potential therapeutic target.
- Kotaro Shimizu
- , Junichi Kikuta
- & Masaru Ishii
-
Article
| Open Access Integrative analysis of transcriptome dynamics during human craniofacial development identifies candidate disease genes
Integrative analysis of transcriptome dynamics during human craniofacial development identifies candidate disease genes
Craniofacial disorders are among the most common congenital defects. Here, the authors examined the genetic causes of non-syndromic craniofacial disorders during human development through analysis of gene expression and epigenomics.
- Tara N. Yankee
- , Sungryong Oh
- & Justin Cotney
-
Article
| Open Access Three-dimensional molecular architecture of mouse organogenesis
Three-dimensional molecular architecture of mouse organogenesis
Qu et al. present a detailed three-dimensional spatial transcriptome atlas of all major organs in the mouse embryo at E13.5, providing a better understanding of organ development and cellular interactions during mammalian development.
- Fangfang Qu
- , Wenjia Li
- & Guangdun Peng
-
Article
| Open Access Identification of transcriptional programs using dense vector representations defined by mutual information with GeneVector
Identification of transcriptional programs using dense vector representations defined by mutual information with GeneVector
In single-cell RNA-seq analyses, it would be critical to measure the relationships between genes. Here, the authors develop a framework for single-cell dimensionality reduction that incorporates gene-specific relationships - GeneVector -, and use it for tasks such as annotating cell types and analysing pathway variation after treatment.
- Nicholas Ceglia
- , Zachary Sethna
- & Andrew McPherson
-
Article
| Open Access High throughput single cell long-read sequencing analyses of same-cell genotypes and phenotypes in human tumors
High throughput single cell long-read sequencing analyses of same-cell genotypes and phenotypes in human tumors
There is a need for methods that allow the analysis of single-cell long-read sequencing data without depending on known barcode lists or short-read sequencing. Here, the authors develop scNanoGPS, a tool that can independently deconvolute long reads into single cells and single molecules, and apply it on tumour and cell line data.
- Cheng-Kai Shiau
- , Lina Lu
- & Ruli Gao
-
Article
| Open Access nnSVG for the scalable identification of spatially variable genes using nearest-neighbor Gaussian processes
nnSVG for the scalable identification of spatially variable genes using nearest-neighbor Gaussian processes
The identification of top spatially variable genes is a key step in the analysis of spatially-resolved transcriptomics data. Here, the authors develop a scalable method based on nearest-neighbor Gaussian processes and evaluate performance compared to existing and baseline methods.
- Lukas M. Weber
- , Arkajyoti Saha
- & Stephanie C. Hicks
-
Article
| Open Access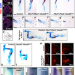 A spatio-temporally constrained gene regulatory network directed by PBX1/2 acquires limb patterning specificity via HAND2
A spatio-temporally constrained gene regulatory network directed by PBX1/2 acquires limb patterning specificity via HAND2
Many key developmental transcriptional regulators are broadly expressed but perform distinct functions in specific tissues. Here they show that ubiquitously expressed PBX factors gain limb bud functionality by interaction with HAND2, uncovering fundamental principles of cooperation between promiscuous and tissue-specific regulators to instruct developmental programs.
- Marta Losa
- , Iros Barozzi
- & Licia Selleri
-
Article
| Open Access Extensive diversity in RNA termination and regulation revealed by transcriptome mapping for the Lyme pathogen Borrelia burgdorferi
Extensive diversity in RNA termination and regulation revealed by transcriptome mapping for the Lyme pathogen Borrelia burgdorferi
Transcription termination can tune bacterial gene expression in response to diverse signals. Here, the authors use several RNA-seq approaches to map RNA ends for the transcriptome of the spirochete Borrelia burgdorferi, providing insights into various modes of transcription termination and identifying potential RNA regulators in this pathogen.
- Emily Petroni
- , Caroline Esnault
- & Philip P. Adams
-
Article
| Open Access Single-cell profiling of lncRNA expression during Ebola virus infection in rhesus macaques
Single-cell profiling of lncRNA expression during Ebola virus infection in rhesus macaques
Long non-coding RNAs (lncRNAs) play key roles in the immune response but their properties at the single-cell level are less well understood. Here, the authors characterize differential features of lncRNAs and protein-coding genes upon Ebola infection in macaques at single-cell resolution.
- Luisa Santus
- , Maria Sopena-Rios
- & Marta Melé
-
Article
| Open Access Single-cell transcriptomics and epigenomics unravel the role of monocytes in neuroblastoma bone marrow metastasis
Single-cell transcriptomics and epigenomics unravel the role of monocytes in neuroblastoma bone marrow metastasis
The bone marrow is a common site of metastasis for neuroblastoma patients. Here, the authors perform single cell RNA-seq and ATAC-seq of bone marrow aspirates from 16 subjects and show conservation of tumor cell plasticity in metastases and identify tumor-to-bone marrow cell signals that trigger tumor promoting monocytes.
- Irfete S. Fetahu
- , Wolfgang Esser-Skala
- & Sabine Taschner-Mandl
-
Article
| Open Access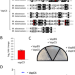 Mycobacterium abscessus VapC5 toxin potentiates evasion of antibiotic killing by ribosome overproduction and activation of multiple resistance pathways
Mycobacterium abscessus VapC5 toxin potentiates evasion of antibiotic killing by ribosome overproduction and activation of multiple resistance pathways
Mycobacterium abscessus (Mab) infections are difficult to clear with antibiotics. Here the authors show that clinical Mab strains can acquire a toxin-antitoxin system that enhances survival upon treatment with current first-line antibiotics through depletion of tRNASerCGA and subsequent ribosome overproduction.
- Eduardo A. Troian
- , Heather M. Maldonado
- & Nancy A. Woychik
-
Article
| Open Access Signaling mechanisms in renal compensatory hypertrophy revealed by multi-omics
Signaling mechanisms in renal compensatory hypertrophy revealed by multi-omics
The authors used a multi-omic approach in a mouse unilateral nephrectomy model to identify signaling processes associated with compensatory hypertrophy of the renal proximal tubule. The results indicate that PPARα is an important determinant of proximal tubule cell size and is a likely mediator of compensatory proximal tubule hypertrophy.
- Hiroaki Kikuchi
- , Chung-Lin Chou
- & Mark A. Knepper
-
Article
| Open Access grandR: a comprehensive package for nucleotide conversion RNA-seq data analysis
grandR: a comprehensive package for nucleotide conversion RNA-seq data analysis
Nucleotide conversion approaches facilitate metabolic RNA labelling experiments but complicate computational analysis. Here, the authors develop a methodology and software package to enable specific analysis methods for nucleotide conversion RNA-seq data.
- Teresa Rummel
- , Lygeri Sakellaridi
- & Florian Erhard
-
Article
| Open Access Immune resilience despite inflammatory stress promotes longevity and favorable health outcomes including resistance to infection
Immune resilience despite inflammatory stress promotes longevity and favorable health outcomes including resistance to infection
The response to infectious and inflammatory challenges differs among people but the reasons for this are poorly understood. Here the authors explore the impact of variables such as age, sex, and the capacity for controlling inflammation and maintaining immunocompetence, linking this capacity to favourable health outcomes and lifespan.
- Sunil K. Ahuja
- , Muthu Saravanan Manoharan
- & Weijing He
-
Article
| Open Access Dynamic chromatin architecture of the porcine adipose tissues with weight gain and loss
Dynamic chromatin architecture of the porcine adipose tissues with weight gain and loss
Here the authors study diet-induced weight gain/loss to identify chromatin architectures in adipose tissue (AT) associated obesity in a pig model. They found parallels and species-specific regulatory elements in humans and pigs that underpin AT specialization.
- Long Jin
- , Danyang Wang
- & Mingzhou Li
-
Article
| Open Access Cell-attribute aware community detection improves differential abundance testing from single-cell RNA-Seq data
Cell-attribute aware community detection improves differential abundance testing from single-cell RNA-Seq data
Single-cell technologies allow quantification of cell-types in human tissues, yet detecting if their abundance changes in aging or disease is challenging. By using cell-attribute aware clustering, this work presents a differential abundance testing algorithm with increased power.
- Alok K. Maity
- & Andrew E. Teschendorff
-
Article
| Open Access M1BP is an essential transcriptional activator of oxidative metabolism during Drosophila development
M1BP is an essential transcriptional activator of oxidative metabolism during Drosophila development
The transcriptional regulation of mitochondrial oxidative phosphorylation gene expression is poorly understood. Using the developing Drosophila flight muscle, the authors identify the transcription factor M1BP as a new major regulator of this process.
- Gabriela Poliacikova
- , Marine Barthez
- & Andrew J. Saurin
-
Article
| Open Access Cell type specific transcriptomic differences in depression show similar patterns between males and females but implicate distinct cell types and genes
Cell type specific transcriptomic differences in depression show similar patterns between males and females but implicate distinct cell types and genes
Sex differences in brain transcriptomics have unknown cell type specificity. Here, authors show concordant cortical transcriptomic patterns in depression within individual cell types between sexes, but distinctly affected top cell types and genes.
- Malosree Maitra
- , Haruka Mitsuhashi
- & Corina Nagy
-
Article
| Open Access High-throughput and high-accuracy single-cell RNA isoform analysis using PacBio circular consensus sequencing
High-throughput and high-accuracy single-cell RNA isoform analysis using PacBio circular consensus sequencing
Long-read single-cell RNA isoform sequencing can elucidate the intricate landscape of alternative RNA splicing in individual cells, but it suffers from a low read throughput. Here, the authors develop circular consensus sequencing methods to allow high-throughput and high-accuracy single-cell RNA isoform sequencing.
- Zhuo-Xing Shi
- , Zhi-Chao Chen
- & Yi-Zhi Liu
-
Article
| Open Access Single-cell RNA-seq uncovers dynamic processes orchestrated by RNA-binding protein DDX43 in chromatin remodeling during spermiogenesis
Single-cell RNA-seq uncovers dynamic processes orchestrated by RNA-binding protein DDX43 in chromatin remodeling during spermiogenesis
Germ cells undergo dramatic chromatin and transcriptomic changes during spermatogenesis, though how this is controlled is not well established. Here they show that RNA-binding protein DDX43 directs the spermatid differentiation trajectory and regulates chromatin remodeling in spermiogenesis, which is partially mediated by its target gene Elfn2.
- Huanhuan Tan
- , Weixu Wang
- & Ke Zheng
-
Article
| Open Access Single-nucleus RNA-sequencing of autosomal dominant Alzheimer disease and risk variant carriers
Single-nucleus RNA-sequencing of autosomal dominant Alzheimer disease and risk variant carriers
Mutations in amyloid precursor protein (APP) and presenilin 1 (PSEN1) cause autosomal dominant AD (ADAD). Here, the authors perform single-nucleus RNA-sequencing of ADAD and other disease risk modifying variant carriers and report altered expression states of specific brain cell types.
- Logan Brase
- , Shih-Feng You
- & Oscar Harari
-
Article
| Open Access Multi-omic underpinnings of epigenetic aging and human longevity
Multi-omic underpinnings of epigenetic aging and human longevity
Here, the authors integrate genomic, bulk and single-cell transcriptomic, and metabolomic data sets to compare the biological underpinning of four epigenetic clocks and human longevity, offering novel insights into aging biology.
- Lucas A. Mavromatis
- , Daniel B. Rosoff
- & Falk W. Lohoff
-
Article
| Open Access Time-of-day defines NAD+ efficacy to treat diet-induced metabolic disease by synchronizing the hepatic clock in mice
Time-of-day defines NAD+ efficacy to treat diet-induced metabolic disease by synchronizing the hepatic clock in mice
The timing of NAD + supply determines its efficacy to treat metabolic disease. Here, the authors show that increasing NAD + at the early active phase maximizes weight loss and glucose regulation in mice. NAD + can displace the phase of the liver clock which can cause circadian misalignment.
- Quetzalcoatl Escalante-Covarrubias
- , Lucía Mendoza-Viveros
- & Lorena Aguilar-Arnal
-
Article
| Open Access Pan-cancer classification of single cells in the tumour microenvironment
Pan-cancer classification of single cells in the tumour microenvironment
The accuracy and granularity of classifying cell types in the tumour microenvironment (TME) from single-cell RNA-seq data is impacted by heterogeneity among cancer cells and similarities among functionally related immune cells. Here, the authors develop scATOMIC, a tumour and TME cell type classifier based on a hierarchical approach that can be applied to pan-cancer datasets.
- Ido Nofech-Mozes
- , David Soave
- & Sagi Abelson
-
Article
| Open Access Spatial transcriptomics using multiplexed deterministic barcoding in tissue
Spatial transcriptomics using multiplexed deterministic barcoding in tissue
Examining the spatially resolved transcriptome of tissue sections promises advances in biomedical research. Here, the authors present xDBiT, a versatile, microfluidics-based approach to cost-effectively measure the spatial transcriptome of multiple tissue sections in parallel.
- Johannes Wirth
- , Nina Huber
- & Matthias Meier
-
Article
| Open Access Well-TEMP-seq as a microwell-based strategy for massively parallel profiling of single-cell temporal RNA dynamics
Well-TEMP-seq as a microwell-based strategy for massively parallel profiling of single-cell temporal RNA dynamics
Gene expression of cells is a heterogeneous and dynamic program involved in various biological processes. Here, authors develop Well-TEMPseq, a high-throughput, cost-effective, and accurate method for massively parallel profiling of the temporal dynamics of single-cell gene expression.
- Shichao Lin
- , Kun Yin
- & Chaoyong Yang
-
Article
| Open Access Quantitative analysis of C. elegans transcripts by Nanopore direct-cDNA sequencing reveals terminal hairpins in non trans-spliced mRNAs
Quantitative analysis of C. elegans transcripts by Nanopore direct-cDNA sequencing reveals terminal hairpins in non trans-spliced mRNAs
C. elegans long read transcriptomic analysis provides evidence that non-trans-spliced mRNAs display a terminal a hairpin structure mimicking the Spliced Leader. This provides an explanation how the main maturation system might be bypassed.
- Florian Bernard
- , Delphine Dargère
- & Denis Dupuy
-
Article
| Open Access RNA splicing analysis using heterogeneous and large RNA-seq datasets
RNA splicing analysis using heterogeneous and large RNA-seq datasets
Here the authors develop MAJIQ v2 to address challenges in detection, quantification, and visualization of RNA splicing variations from large heterogeneous RNA-Seq datasets. They then apply it to analyze 2,335 samples from 13 brain subregions.
- Jorge Vaquero-Garcia
- , Joseph K. Aicher
- & Yoseph Barash
-
Article
| Open Access BdLT-Seq as a barcode decay-based method to unravel lineage-linked transcriptome plasticity
BdLT-Seq as a barcode decay-based method to unravel lineage-linked transcriptome plasticity
Cellular plasticity is a core biological process; however, observing diversity in non-genetic inheritance and the resulting phenotypic outputs, is challenging. Here the authors develop a non-genetically based tracing technology which can be used to reveal lineage-linked transcriptome plasticity.
- Yelyzaveta Shlyakhtina
- , Bianca Bloechl
- & Maximiliano M. Portal
-
Article
| Open Access Reconstruction of the tumor spatial microenvironment along the malignant-boundary-nonmalignant axis
Reconstruction of the tumor spatial microenvironment along the malignant-boundary-nonmalignant axis
Delineating the cellular composition of tumour boundaries in spatial transcriptomics (ST) data is challenging. Here, the authors develop Cottrazm to integrate ST with histological imaging and single-cell data, identify the malignant and non-malignant tissue boundaries, deconvolute cell-type composition, and reconstruct cell type-specific gene expression profiles.
- Zhenzhen Xun
- , Xinyu Ding
- & Youqiong Ye

 Positive regulation of oxidative phosphorylation by nuclear myosin 1 protects cells from metabolic reprogramming and tumorigenesis in mice
Positive regulation of oxidative phosphorylation by nuclear myosin 1 protects cells from metabolic reprogramming and tumorigenesis in mice