Featured
-
-
Article
| Open Access Predicting response to enzalutamide and abiraterone in metastatic prostate cancer using whole-omics machine learning
Predicting response to enzalutamide and abiraterone in metastatic prostate cancer using whole-omics machine learning
Prostate cancer is known to have a variable response to androgen receptor signalling inhibitors. Here, the authors use machine learning to predict response to therapy from genomic, transcriptomic and clinical data.
- Anouk C. de Jong
- , Alexandra Danyi
- & Martijn P. Lolkema
-
Article
| Open Access Systematic comparison of tools used for m6A mapping from nanopore direct RNA sequencing
Systematic comparison of tools used for m6A mapping from nanopore direct RNA sequencing
Direct RNA sequencing using nanopore platform can be used to detect N6-methyladenosine (m6A) modifications on mRNAs. Here the authors systematically compare tools used for m6A detection from nanopore direct sequencing.
- Zhen-Dong Zhong
- , Ying-Yuan Xie
- & Guan-Zheng Luo
-
Article
| Open Access MAIT cell inhibition promotes liver fibrosis regression via macrophage phenotype reprogramming
MAIT cell inhibition promotes liver fibrosis regression via macrophage phenotype reprogramming
Liver cirrhosis is characterised by extensive fibrosis of the liver, and understanding the underpinning immunological processes is important in designing intervention. Here authors show that Mucosal-Associated Invariant T cells are instrumental to controlling the balance between profibrogenic and restorative macrophages and inhibiting their activation might reverse liver fibrosis.
- Morgane Mabire
- , Pushpa Hegde
- & Sophie Lotersztajn
-
Article
| Open Access Muscle cell-type diversification is driven by bHLH transcription factor expansion and extensive effector gene duplications
Muscle cell-type diversification is driven by bHLH transcription factor expansion and extensive effector gene duplications
Different muscle cell types account for specific abilities in animals, yet how their diversification arose remains unclear. Here, the authors show that gene duplications of bHLH transcription factors and effector genes contributed to the diversification of muscle cell types in the sea anemone Nematostella.
- Alison G. Cole
- , Stefan M. Jahnel
- & Ulrich Technau
-
Article
| Open Access IGFBP5 is an ROR1 ligand promoting glioblastoma invasion via ROR1/HER2-CREB signaling axis
IGFBP5 is an ROR1 ligand promoting glioblastoma invasion via ROR1/HER2-CREB signaling axis
Glioblastoma stem-like cells (GSCs) contribute to therapeutic resistance and recurrence of glioblastomas. Here the authors show that Insulin-like Growth Factor-Binding Protein 5 (IGFBP5) is a ligand for Receptor tyrosine kinase-like Orphan Receptor 1 (ROR1) promotes GSCs invasion.
- Weiwei Lin
- , Rui Niu
- & Jinlong Yin
-
Article
| Open Access Spatial transcriptomics using multiplexed deterministic barcoding in tissue
Spatial transcriptomics using multiplexed deterministic barcoding in tissue
Examining the spatially resolved transcriptome of tissue sections promises advances in biomedical research. Here, the authors present xDBiT, a versatile, microfluidics-based approach to cost-effectively measure the spatial transcriptome of multiple tissue sections in parallel.
- Johannes Wirth
- , Nina Huber
- & Matthias Meier
-
Article
| Open Access Immune subset-committed proliferating cells populate the human foetal intestine throughout the second trimester of gestation
Immune subset-committed proliferating cells populate the human foetal intestine throughout the second trimester of gestation
The intestine is an important immunological organ in embryonic life, preparing the infant for the microbial colonization following birth. Here authors show that between gestational weeks 14 and 22, the human foetal intestine is first populated by myeloid and innate lymphoid cells, followed by the development of lymphoid cells and a wider range of proliferation-capable immune cell types.
- Nannan Guo
- , Na Li
- & Frits Koning
-
Article
| Open Access Longitudinal single-cell profiling of chemotherapy response in acute myeloid leukemia
Longitudinal single-cell profiling of chemotherapy response in acute myeloid leukemia
Relapse within acute myeloid leukaemia may be driven by the presence of leukaemia stem cells. Here, the authors use single cell RNA-seq seq to characterise leukemia stem cells, and show miR-126 as a potential marker of resistance.
- Matteo Maria Naldini
- , Gabriele Casirati
- & Bernhard Gentner
-
Article
| Open Access Well-TEMP-seq as a microwell-based strategy for massively parallel profiling of single-cell temporal RNA dynamics
Well-TEMP-seq as a microwell-based strategy for massively parallel profiling of single-cell temporal RNA dynamics
Gene expression of cells is a heterogeneous and dynamic program involved in various biological processes. Here, authors develop Well-TEMPseq, a high-throughput, cost-effective, and accurate method for massively parallel profiling of the temporal dynamics of single-cell gene expression.
- Shichao Lin
- , Kun Yin
- & Chaoyong Yang
-
Article
| Open Access BdLT-Seq as a barcode decay-based method to unravel lineage-linked transcriptome plasticity
BdLT-Seq as a barcode decay-based method to unravel lineage-linked transcriptome plasticity
Cellular plasticity is a core biological process; however, observing diversity in non-genetic inheritance and the resulting phenotypic outputs, is challenging. Here the authors develop a non-genetically based tracing technology which can be used to reveal lineage-linked transcriptome plasticity.
- Yelyzaveta Shlyakhtina
- , Bianca Bloechl
- & Maximiliano M. Portal
-
Article
| Open Access Single-cell transcriptome profiling of the stepwise progression of head and neck cancer
Single-cell transcriptome profiling of the stepwise progression of head and neck cancer
Head and neck squamous cell carcinomas (HNSCCs) undergo a stepwise progression from normal tissues. In order to better understand the molecular mechanisms behind such progression, here the authors profile HNSCC tumors at different stages using single-cell RNA-seq, and observe the role of interactions with the tumor microenvironment.
- Ji-Hye Choi
- , Bok-Soon Lee
- & Chul-Ho Kim
-
Article
| Open Access Application of high-throughput single-nucleus DNA sequencing in pancreatic cancer
Application of high-throughput single-nucleus DNA sequencing in pancreatic cancer
Implementing high-throughput single-cell DNA sequencing for the study of solid tumours has been challenging. Here, the authors present an optimised approach for snap-frozen tissue single nuclei extraction and DNA sequencing, which can be applied to study pancreatic ductal adenocarcinoma evolution and heterogeneity.
- Haochen Zhang
- , Elias-Ramzey Karnoub
- & Christine A. Iacobuzio-Donahue
-
Article
| Open Access Dissecting the immune suppressive human prostate tumor microenvironment via integrated single-cell and spatial transcriptomic analyses
Dissecting the immune suppressive human prostate tumor microenvironment via integrated single-cell and spatial transcriptomic analyses
The immune suppressive tumour microenvironment drives recurrence and metastatic disease in prostate cancer. Here authors provide a detailed analysis of the microenvironment via single cell RNA sequencing and high-resolution spatial transcriptomics to identify tumour-dependent changes compared to healthy tissue.
- Taghreed Hirz
- , Shenglin Mei
- & David B. Sykes
-
Article
| Open Access Single-exonuclease nanocircuits reveal the RNA degradation dynamics of PNPase and demonstrate potential for RNA sequencing
Single-exonuclease nanocircuits reveal the RNA degradation dynamics of PNPase and demonstrate potential for RNA sequencing
Observing the natural process of RNA degradation in real-time is a significant challenge. Here, the authors develop and use single-exonuclease nanocircuits to reveal the single-base degradation behaviour of PNPase, and demonstrate proof-of-principle RNA sequencing using this approach.
- Zhiheng Yang
- , Wenzhe Liu
- & Xuefeng Guo
-
Article
| Open Access Spatially resolved transcriptomic profiling of degraded and challenging fresh frozen samples
Spatially resolved transcriptomic profiling of degraded and challenging fresh frozen samples
Spatial transcriptomics relies on RNA quality, which is variable and dependent on sample handling, storage, and/or intrinsic factors. Here, authors present a genome-wide spatial gene expression profiling method called RNA Rescue Spatial Transcriptomics (RRST), designed for the analysis of moderate to low quality fresh frozen tissue samples and demonstrate its robustness on 7 different tissue types.
- Reza Mirzazadeh
- , Zaneta Andrusivova
- & Joakim Lundeberg
-
Article
| Open Access Epigenetic control of cellular crosstalk defines gastrointestinal organ fate and function
Epigenetic control of cellular crosstalk defines gastrointestinal organ fate and function
Mesenchymal-epithelial crosstalk plays a key role in gut development and stem cell homeostasis, though underlying mechanisms are still unclear. Here, the authors demonstrate that mesenchymal Polycomb Repressive Complex 2 controls niche signals for gut epithelial fate and growth.
- Ryan J. Smith
- , Minggao Liang
- & Tae-Hee Kim
-
Article
| Open Access Decision level integration of unimodal and multimodal single cell data with scTriangulate
Decision level integration of unimodal and multimodal single cell data with scTriangulate
Single-cell genomics has expanded to measure diverse molecular modalities within the same cell. Here the authors provide a computational framework called scTriangulate to integrate cluster annotations from diverse independent sources, algorithms, and modalities to define statistically stable populations.
- Guangyuan Li
- , Baobao Song
- & Nathan Salomonis
-
Article
| Open Access Topological identification and interpretation for single-cell gene regulation elucidation across multiple platforms using scMGCA
Topological identification and interpretation for single-cell gene regulation elucidation across multiple platforms using scMGCA
A major challenge in analyzing scRNA-seq data arises from challenges related to dimensionality and the prevalence of dropout events. Here the authors develop a deep graph learning method called scMGCA based on a graph-embedding autoencoder that simultaneously learns cell-cell topology representation and cluster assignments, outperforming other state-of-the-art models across multiple platforms.
- Zhuohan Yu
- , Yanchi Su
- & Xiangtao Li
-
Article
| Open Access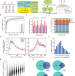 scm6A-seq reveals single-cell landscapes of the dynamic m6A during oocyte maturation and early embryonic development
scm6A-seq reveals single-cell landscapes of the dynamic m6A during oocyte maturation and early embryonic development
Modification of RNA with N6-methyladenosine can regulate RNA metabolism. Here they developed scm6A-seq to profile the methylome and transcriptome in single cells, and reveal the functions of m6A modification during oocyte maturation and early embryo development.
- Huan Yao
- , Chun-Chun Gao
- & Yun-Gui Yang
-
Article
| Open Access Semi-quantitative detection of pseudouridine modifications and type I/II hypermodifications in human mRNAs using direct long-read sequencing
Semi-quantitative detection of pseudouridine modifications and type I/II hypermodifications in human mRNAs using direct long-read sequencing
Pseudouridine (psi) is an RNA modification that can affect its physiology, including increased half-life. Here the authors identify sites of psi modification in the human transcriptome using direct RNA sequencing and provide a “ground truth” list of psi sites, sites of high psi occupancy, and transcripts that may be modified at multiple sites.
- Sepideh Tavakoli
- , Mohammad Nabizadeh
- & Sara H. Rouhanifard
-
Article
| Open Access In vivo induction of activin A-producing alveolar macrophages supports the progression of lung cell carcinoma
In vivo induction of activin A-producing alveolar macrophages supports the progression of lung cell carcinoma
Alveolar macrophages represent a cell type that is physiologic to the lung immune landscape, however, it is not known whether they play an active role to maintain the tumour immune microenvironment. Here authors show by single cell RNA sequencing and functional experiments, that intra-tumour alveolar macrophages are phenotypically and transcriptionally different from the healthy ones, and likely play an aetio-pathologic role in tumorigenesis.
- Seiji Taniguchi
- , Takahiro Matsui
- & Masaru Ishii
-
Article
| Open Access Germline TP53 mutations undergo copy number gain years prior to tumor diagnosis
Germline TP53 mutations undergo copy number gain years prior to tumor diagnosis
Li-Fraumeni syndrome (LFS) is associated with pathogenic germline TP53 variants and predisposes patients to cancer; understanding the evolution and drivers of LFS-related tumours remains crucial. Here, the authors analyse 22 LFS tumours using whole-genome sequencing and reconstruct the evolution and timing of somatic driver alterations.
- Nicholas Light
- , Mehdi Layeghifard
- & Adam Shlien
-
Article
| Open Access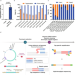 TAPE-seq is a cell-based method for predicting genome-wide off-target effects of prime editor
TAPE-seq is a cell-based method for predicting genome-wide off-target effects of prime editor
Methods to predict genome-wide off-target activities of prime editors (PEs) are currently lacking. Here the authors report a cell-based assay, TAgmentation of Prime Editor sequencing (TAPE-seq), that provides genome-wide off-target candidates for PEs.
- Jeonghun Kwon
- , Minyoung Kim
- & Jungjoon K. Lee
-
Article
| Open Access Simultaneous profiling of histone modifications and DNA methylation via nanopore sequencing
Simultaneous profiling of histone modifications and DNA methylation via nanopore sequencing
The interplay between histone modifications and DNA methylation plays a crucial role in establishing and maintaining the epigenomic landscape. Here, the authors develop a nanopore sequencing based method for mapping histone modifications and DNA methylation from native, long, single DNA molecules.
- Xue Yue
- , Zhiyuan Xie
- & Yimeng Yin
-
Article
| Open Access Dysfunctional Sars-CoV-2-M protein-specific cytotoxic T lymphocytes in patients recovering from severe COVID-19
Dysfunctional Sars-CoV-2-M protein-specific cytotoxic T lymphocytes in patients recovering from severe COVID-19
Cytotoxic T lymphocytes play important roles in the anti-viral immune response in COVID-19, and it is important to know how they contribute to disease outcome. Authors here identify a dominant SARS-CoV-2 M protein epitope, M198–206, and show that M198–206-specific cytotoxic T cells from convalescent patients with severe disease harbour a gene expression pattern indicative of poor functionality.
- Hideki Ogura
- , Jin Gohda
- & Satoshi Ishido
-
Article
| Open Access High-throughput robust single-cell DNA methylation profiling with sciMETv2
High-throughput robust single-cell DNA methylation profiling with sciMETv2
Despite the importance of DNA methylation, accessible and high-throughput methods to profile methylation at the single-cell level are lacking. Here, the authors present sciMETv2, a high-throughput workflow that provides high-quality single-cell methylomes in a robust and simple workflow.
- Ruth V. Nichols
- , Brendan L. O’Connell
- & Andrew C. Adey
-
Article
| Open Access Whole genome sequencing identifies structural variants contributing to hematologic traits in the NHLBI TOPMed program
Whole genome sequencing identifies structural variants contributing to hematologic traits in the NHLBI TOPMed program
Most genetic association studies have been done on single nucleotide polymorphisms and small indels, while other types of variants have been less studied. Here, the authors use whole genome sequencing in a diverse population to identify and provide experimental evidence for associations between structural variants and blood-cell traits.
- Marsha M. Wheeler
- , Adrienne M. Stilp
- & Alex P. Reiner
-
Article
| Open Access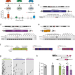 Retrotransposon instability dominates the acquired mutation landscape of mouse induced pluripotent stem cells
Retrotransposon instability dominates the acquired mutation landscape of mouse induced pluripotent stem cells
Retrotransposons are mobile genetic elements normally repressed by DNA methylation in differentiated cells. Here, the authors show that DNA hypomethylation in mouse induced pluripotent stem cells allows retrotransposons to jump, but this can be blocked with a reverse transcriptase inhibitor.
- Patricia Gerdes
- , Sue Mei Lim
- & Geoffrey J. Faulkner
-
Article
| Open Access A unified computational framework for single-cell data integration with optimal transport
A unified computational framework for single-cell data integration with optimal transport
Integrating heterogeneous single-cell multi-omics as well as spatially resolved transcriptomic data remains a major challenge. Here the authors report a unified single-cell data integration framework using an unbalanced optimal transport-based deep network.
- Kai Cao
- , Qiyu Gong
- & Lin Wan
-
Article
| Open Access Low-dose IL-2 reduces IL-21+ T cell frequency and induces anti-inflammatory gene expression in type 1 diabetes
Low-dose IL-2 reduces IL-21+ T cell frequency and induces anti-inflammatory gene expression in type 1 diabetes
Low-dose interleukin-2 is showing promise in the treatment of several autoimmune inflammatory diseases. Here authors map the trajectory of cellular and transcriptional changes in type 1 diabetes patients receiving an interval dosing interleukin-2 regimen, which shows an anti-inflammatory gene expression signature shared by all immune cell types analysed, persisting for at least a month after ending treatment.
- Jia-Yuan Zhang
- , Fiona Hamey
- & Ricardo C. Ferreira
-
Article
| Open Access Leveraging data-driven self-consistency for high-fidelity gene expression recovery
Leveraging data-driven self-consistency for high-fidelity gene expression recovery
Recovering dropout-affected gene expression values is a challenging problem in bioinformatics. Here, the authors propose a data-driven framework, that first learns the underlying data distribution and then recovers the expression values by imposing a self-consistency on the expression matrix.
- Md Tauhidul Islam
- , Jen-Yeu Wang
- & Lei Xing
-
Article
| Open Access Systematic tissue annotations of genomics samples by modeling unstructured metadata
Systematic tissue annotations of genomics samples by modeling unstructured metadata
The 1+ million publicly-available human –omics samples currently remain acutely underused. Here the authors present an approach combining natural language processing and machine learning to infer the source tissue of public genomics samples based on their plain text descriptions, making these samples easy to discover and reuse.
- Nathaniel T. Hawkins
- , Marc Maldaver
- & Arjun Krishnan
-
Article
| Open Access Deep autoencoder for interpretable tissue-adaptive deconvolution and cell-type-specific gene analysis
Deep autoencoder for interpretable tissue-adaptive deconvolution and cell-type-specific gene analysis
Traditional bulk sequencing data lack information about cell-type-specific gene expression. Here, the authors develop a Tissue-AdaPtive autoEncoder (TAPE), a deep learning method connecting bulk RNA-seq and single-cell RNA-seq, and apply it to analyze the cell type fractions and cell-type-specific gene expression in clinical data.
- Yanshuo Chen
- , Yixuan Wang
- & Yu Li
-
Article
| Open Access Analysis of matched primary and recurrent BRCA1/2 mutation-associated tumors identifies recurrence-specific drivers
Analysis of matched primary and recurrent BRCA1/2 mutation-associated tumors identifies recurrence-specific drivers
Carriers of pathogenic BRCA1/2 variants have a higher risk of breast and ovarian cancers, which recur frequently. Here, the authors sequence primary and recurrent tumours of BRCA1/2 mutation carriers, finding PARP1 amplifications, differential BRCA2 isoform usage, and discordant loss of heterozygosity that are associated with recurrence.
- Jennifer B. Shah
- , Dana Pueschl
- & Katherine L. Nathanson
-
Article
| Open Access VeChat: correcting errors in long reads using variation graphs
VeChat: correcting errors in long reads using variation graphs
Consensus sequence-based methods for self-correction of long-read sequencing data are affected by biases that can mask true variants characterizing little-covered or low-frequency haplotypes. Here, to address this issue, the authors develop a variation graph-based method for performing haplotype-aware self-correction of long reads.
- Xiao Luo
- , Xiongbin Kang
- & Alexander Schönhuth
-
Article
| Open Access UniTVelo: temporally unified RNA velocity reinforces single-cell trajectory inference
UniTVelo: temporally unified RNA velocity reinforces single-cell trajectory inference
RNA velocity can detect the differentiation directionality by modelling sparse unspliced RNAs, but suffers from high estimation errors. Here, the authors develop a computational method called UniTVelo to reinforce the velocity estimation by introducing a unified time and a top-down model design.
- Mingze Gao
- , Chen Qiao
- & Yuanhua Huang
-
Article
| Open Access De novo analysis of bulk RNA-seq data at spatially resolved single-cell resolution
De novo analysis of bulk RNA-seq data at spatially resolved single-cell resolution
Current methods to reanalyze bulk RNA-seq at spatially resolved single-cell resolution have limitations. Here, the authors develop Bulk2Space, a spatial deconvolution algorithm using single-cell and spatial transcriptomics as references, providing new insights into spatial heterogeneity within bulk tissue.
- Jie Liao
- , Jingyang Qian
- & Xiaohui Fan
-
Article
| Open Access Defining cellular complexity in human autosomal dominant polycystic kidney disease by multimodal single cell analysis
Defining cellular complexity in human autosomal dominant polycystic kidney disease by multimodal single cell analysis
Autosomal dominant polycystic kidney disease (ADPKD) is a complicated disease that involves numerous cell types. Here the authors used a multiomics approach consisting of single nucleus transcriptomes and epigenomes to redefine cell states in ADPKD and to dissect the cellular interactions and molecular mechanisms of ADPKD.
- Yoshiharu Muto
- , Eryn E. Dixon
- & Benjamin D. Humphreys
-
Article
| Open Access Library adaptors with integrated reference controls improve the accuracy and reliability of nanopore sequencing
Library adaptors with integrated reference controls improve the accuracy and reliability of nanopore sequencing
Adding library adaptors to DNA samples is an essential step in preparing samples for next-generation sequencing. Here, Gunter et al. describe the development of Control Library Adaptors (CAPTORs), that correct sequencing errors and normalise quantitative biases in Nanopore libraries.
- Helen M. Gunter
- , Scott E. Youlten
- & Tim R. Mercer
-
Article
| Open Access HiFi metagenomic sequencing enables assembly of accurate and complete genomes from human gut microbiota
HiFi metagenomic sequencing enables assembly of accurate and complete genomes from human gut microbiota
Here, the authors construct 102 complete metagenome-assembled genomes (cMAGs) from Pacific Biosciences (PacBio) high-accuracy long-read (HiFi) metagenomic sequencing, showing as high nucleotide accuracy as reference genomes and revealing that regions hard to assemble by short-read sequencing comprise mostly of genomic islands and rRNAs.
- Chan Yeong Kim
- , Junyeong Ma
- & Insuk Lee
-
Article
| Open Access Characterization of somatic structural variations in 528 Chinese individuals with Esophageal squamous cell carcinoma
Characterization of somatic structural variations in 528 Chinese individuals with Esophageal squamous cell carcinoma
The biological and clinical implications of high genome instability in esophageal squamous cell carcinoma (ESCC) remain to be explored. Here, the authors analyse 528 whole genomes and investigate the landscape and molecular mechanisms underlying structural variations in ESCC patients.
- Heyang Cui
- , Yong Zhou
- & Qimin Zhan
-
Article
| Open Access Online single-cell data integration through projecting heterogeneous datasets into a common cell-embedding space
Online single-cell data integration through projecting heterogeneous datasets into a common cell-embedding space
Integrative analyses of single-cell datasets are facing new challenges as data size and complexity grow. Here the authors present SCALEX, which projects cells from different datasets into a common latent space, allowing accurate online integration as well as cross-referencing with atlas-scale data.
- Lei Xiong
- , Kang Tian
- & Qiangfeng Cliff Zhang
-
Article
| Open Access Alignment of single-cell trajectory trees with CAPITAL
Alignment of single-cell trajectory trees with CAPITAL
Global alignment of complex cell state trajectories between single-cell datasets remains challenging. Here, the authors present a computational method called CAPITAL to compare branching trajectories, and demonstrate that this method achieves accurate and robust alignments.
- Reiichi Sugihara
- , Yuki Kato
- & Yukio Kawahara
-
Article
| Open Access GWAS, MWAS and mGWAS provide insights into precision agriculture based on genotype-dependent microbial effects in foxtail millet
GWAS, MWAS and mGWAS provide insights into precision agriculture based on genotype-dependent microbial effects in foxtail millet
Plant genotype alone appears to be insufficient to explain trait variations. This study integrates GWAS, MWAS and mGWAS in 827 foxtail millet cultivars, revealing that root-associated microbiota affect plant phenotypes in a host genotype-dependent manner.
- Yayu Wang
- , Xiaolin Wang
- & Huan Liu
-
Article
| Open Access A method for multiplexed full-length single-molecule sequencing of the human mitochondrial genome
A method for multiplexed full-length single-molecule sequencing of the human mitochondrial genome
Accurate analysis of mitochondrial DNA is important for mitochondrial disease clinical research and diagnostics. Here, authors present a method using Cas9 cleavage, nanopore sequencing and a custom pipeline to identify pathogenic variants, deletions and accurately quantify heteroplasmy to below 1%.
- Ieva Keraite
- , Philipp Becker
- & Ivo Glynne Gut
-
Article
| Open Access Single cell and spatial transcriptomic analyses reveal microglia-plasma cell crosstalk in the brain during Trypanosoma brucei infection
Single cell and spatial transcriptomic analyses reveal microglia-plasma cell crosstalk in the brain during Trypanosoma brucei infection
Detailed insight into how the brain responds to Trypanosoma brucei infection is lacking. Here, single cell and spatial transcriptomics are integrated to characterise this response, identifying a unique crosstalk between microglia and plasma cells.
- Juan F. Quintana
- , Praveena Chandrasegaran
- & Annette MacLeod
-
Article
| Open Access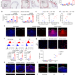 Single-cell transcriptomics reveal cellular diversity of aortic valve and the immunomodulation by PPARγ during hyperlipidemia
Single-cell transcriptomics reveal cellular diversity of aortic valve and the immunomodulation by PPARγ during hyperlipidemia
Identifying the mechanisms underlying the early inflammatory phase of aortic valve disease is crucial for disease prevention. Here the authors perform single-cell RNA sequencing to show the immunomodulatory role of PPARγ in valvular endothelial cells during hyperlipidemia.
- Seung Hyun Lee
- , Nayoung Kim
- & Jae-Hoon Choi
-
Article
| Open Access Multimodal single cell sequencing implicates chromatin accessibility and genetic background in diabetic kidney disease progression
Multimodal single cell sequencing implicates chromatin accessibility and genetic background in diabetic kidney disease progression
Diabetic kidney disease leads to changes in glucose metabolism and inflammation. Here the authors use multimodal single cell sequencing to show that this disease leads to reduced accessibility of glucocorticoid receptor binding sites in the proximal tubule and increased gluconeogenesis.
- Parker C. Wilson
- , Yoshiharu Muto
- & Benjamin D. Humphreys
-
Article
| Open Access PIM1 promotes hepatic conversion by suppressing reprogramming-induced ferroptosis and cell cycle arrest
PIM1 promotes hepatic conversion by suppressing reprogramming-induced ferroptosis and cell cycle arrest
Protein kinase-mediated phosphorylation plays a critical role in many biological processes. Here the authors develop a trans-omics-based algorithm called Central Kinase Inference to integrate quantitative transcriptomic and phosphoproteomic data, finding that PIM1 promotes hepatic conversion by suppressing reprogramming-induced ferroptosis and cell cycle arrest.
- Yangyang Yuan
- , Chenwei Wang
- & Pengyu Huang

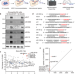 Screening of Drosophila microRNA-degradation sequences reveals Argonaute1 mRNA’s role in regulating miR-999
Screening of Drosophila microRNA-degradation sequences reveals Argonaute1 mRNA’s role in regulating miR-999