Featured
-
-
Article
| Open Access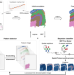 Pianno: a probabilistic framework automating semantic annotation for spatial transcriptomics
Pianno: a probabilistic framework automating semantic annotation for spatial transcriptomics
Recognising spatial spots’ biological identity in spatial transcriptomics remains a challenge. Here, authors introduce Pianno, a tool that helps annotate the biological structures or cell-type constructions across diverse tissues, offering new perspectives on understanding spatial transcriptomics.
- Yuqiu Zhou
- , Wei He
- & Ying Zhu
-
Article
| Open Access DELVE: feature selection for preserving biological trajectories in single-cell data
DELVE: feature selection for preserving biological trajectories in single-cell data
Characteristic genes or proteins driving continuous biological processes are difficult to uncover from noisy single-cell data. Here, authors present DELVE, an unsupervised feature selection method to identify core molecular features driving cell fate decisions.
- Jolene S. Ranek
- , Wayne Stallaert
- & Jeremy E. Purvis
-
Article
| Open Access NAP-seq reveals multiple classes of structured noncoding RNAs with regulatory functions
NAP-seq reveals multiple classes of structured noncoding RNAs with regulatory functions
The genome-wide prevalence, mechanism and function of noncapped RNAs (napRNAs) are currently poorly understood. Here, the authors develop a method called NAP-seq, to globally profile the full-length sequences of napRNAs, revealing several classes of structured noncoding RNAs.
- Shurong Liu
- , Junhong Huang
- & Jianhua Yang
-
Article
| Open Access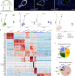 Single-cell division tracing and transcriptomics reveal cell types and differentiation paths in the regenerating lung
Single-cell division tracing and transcriptomics reveal cell types and differentiation paths in the regenerating lung
This study uses single-cell transcriptomics to examine how lung cells respond to targeted damage. The authors employ genetically modified mouse models and cell sorting to enrich for rare, actively dividing cells, revealing cell types/states and alternative differentiation paths.
- Leila R. Martins
- , Lina Sieverling
- & Claudia Scholl
-
Article
| Open Access An organism-wide atlas of hormonal signaling based on the mouse lemur single-cell transcriptome
An organism-wide atlas of hormonal signaling based on the mouse lemur single-cell transcriptome
Endocrinologists have traditionally focused on studying one hormone or organ system at a time. Here the authors use transcriptomic data from the mouse lemur to globally characterize primate hormonal signaling, describing hormone sources and targets, identifying conserved and primate specific regulation, and elucidating principles of the network.
- Shixuan Liu
- , Camille Ezran
- & James E. Ferrell Jr.
-
Article
| Open Access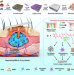 Endogenous stimuli-responsive separating microneedles to inhibit hypertrophic scar through remodeling the pathological microenvironment
Endogenous stimuli-responsive separating microneedles to inhibit hypertrophic scar through remodeling the pathological microenvironment
The treatment of hypertrophic scar (HS) is hindered by the low bioavailability of drugs and the pathological microenvironment. Here the authors report a separating microneedle drug delivery system responsive to high reactive oxygen species levels and overexpression of matrix metalloproteinases to remodel the pathological microenvironment for HS treatment.
- Zhuo-Ran Yang
- , Huinan Suo
- & Jintao Zhu
-
Article
| Open Access A single-cell atlas of Drosophila trachea reveals glycosylation-mediated Notch signaling in cell fate specification
A single-cell atlas of Drosophila trachea reveals glycosylation-mediated Notch signaling in cell fate specification
Studying Drosophila trachea development can inform the mechanisms of growth of all tubular structures. Here, the authors generate a transcriptomic cell atlas of the developing fly trachea and establish roles for Notch signaling, which may be disrupted by diet-induced glycosylation.
- Yue Li
- , Tianfeng Lu
- & Hai Huang
-
Article
| Open Access TFvelo: gene regulation inspired RNA velocity estimation
TFvelo: gene regulation inspired RNA velocity estimation
Most RNA velocity models extract dynamics from the phase delay between unspliced and spliced mRNA for each gene. Here, authors propose TFvelo, broadening RNA velocity beyond splicing information to include gene regulation. TFvelo accurately models genes dynamics and infers cell pseudo-time from RNA abundance data.
- Jiachen Li
- , Xiaoyong Pan
- & Hong-Bin Shen
-
Article
| Open Access The dynamic genetic determinants of increased transcriptional divergence in spermatids
The dynamic genetic determinants of increased transcriptional divergence in spermatids
Here the authors show that genetic changes between species often alter gene expression in a cell type-specific manner. Most of this variability is driven by locally functioning cis-acting variation, and this contributes to the speed at which cell types accumulate expression changes.
- Jasper Panten
- , Tobias Heinen
- & Duncan T. Odom
-
Article
| Open Access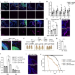 Mechanistic characterization of a Drosophila model of paraneoplastic nephrotic syndrome
Mechanistic characterization of a Drosophila model of paraneoplastic nephrotic syndrome
The fruit fly Drosophila melanogaster has emerged as a model to characterize the mechanisms of tumor-induced host organ dysfunction. Here, Xu, Liu et al. describe a mechanism of tumor-induced kidney dysfunction through hyper-activation of the PvR/JNK/Jra pathway in the Principal cells of the fly kidney/Malpighian tubules.
- Jun Xu
- , Ying Liu
- & Norbert Perrimon
-
Article
| Open Access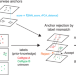 Semi-supervised integration of single-cell transcriptomics data
Semi-supervised integration of single-cell transcriptomics data
Batch effects hinder multi-sample single-cell data analyses. Here, authors present STACAS, a scalable single-cell RNA-seq data integration tool that uses prior cell type knowledge to preserve biological variability, demonstrating robustness to noisy input cell type labels.
- Massimo Andreatta
- , Léonard Hérault
- & Santiago J. Carmona
-
Article
| Open Access PROST: quantitative identification of spatially variable genes and domain detection in spatial transcriptomics
PROST: quantitative identification of spatially variable genes and domain detection in spatial transcriptomics
Understanding biological mechanisms requires a thorough exploration of spatiotemporal transcriptional patterns in complex tissues. Here, authors present PROST to quantify spatial gene expression patterns and detect spatial domains using spatial transcriptomics data of varying resolutions.
- Yuchen Liang
- , Guowei Shi
- & Zhonghui Tang
-
Article
| Open Access BIDCell: Biologically-informed self-supervised learning for segmentation of subcellular spatial transcriptomics data
BIDCell: Biologically-informed self-supervised learning for segmentation of subcellular spatial transcriptomics data
Subcellular in situ spatial transcriptomics offers the promise to address biological problems that were previously inaccessible but requires accurate cell segmentation to uncover insights. Here, authors present BIDCell, a biologically informed, deep learning-based cell segmentation framework.
- Xiaohang Fu
- , Yingxin Lin
- & Jean Y. H. Yang
-
Article
| Open Access Direct RNA sequencing coupled with adaptive sampling enriches RNAs of interest in the transcriptome
Direct RNA sequencing coupled with adaptive sampling enriches RNAs of interest in the transcriptome
It can be difficult to find rare transcripts when sequencing a transcriptome. Here the authors show adaptive sampling on direct RNA runs to increase the likelihood of finding less frequent ones while selectively ejecting the higher-abundance transcripts.
- Jiaxu Wang
- , Lin Yang
- & Yue Wan
-
Article
| Open Access MarsGT: Multi-omics analysis for rare population inference using single-cell graph transformer
MarsGT: Multi-omics analysis for rare population inference using single-cell graph transformer
Identifying rare cell populations is key to understanding cancer progression and response to therapy. Here, authors introduce MarsGT, an end-to-end deep learning model for rare cell population identification from scMulti-omics data.
- Xiaoying Wang
- , Maoteng Duan
- & Qin Ma
-
Article
| Open Access MENDER: fast and scalable tissue structure identification in spatial omics data
MENDER: fast and scalable tissue structure identification in spatial omics data
Identifying tissue structure in large-scale spatial omics datasets from multiple slices is challenging. Here, authors present MENDER, an optimisation-free spatial clustering method that can scale to million-level spatial data, enabling efficient analysis of spatial cell atlases.
- Zhiyuan Yuan
-
Article
| Open Access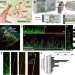 Angiogenesis-on-a-chip coupled with single-cell RNA sequencing reveals spatially differential activations of autophagy along angiogenic sprouts
Angiogenesis-on-a-chip coupled with single-cell RNA sequencing reveals spatially differential activations of autophagy along angiogenic sprouts
The functional heterogeneity of autophagy in endothelial cells during angiogenesis remains incompletely understood. Here, the authors apply a 3D angiogenesis-on-a-chip coupled with single-cell RNA sequencing to find distinct autophagy functions in two different endothelial cell populations during angiogenic sprouting.
- Somin Lee
- , Hyunkyung Kim
- & Noo Li Jeon
-
Article
| Open Access Terminal modifications independent cell-free RNA sequencing enables sensitive early cancer detection and classification
Terminal modifications independent cell-free RNA sequencing enables sensitive early cancer detection and classification
Cell free RNA is a potentially valuable resource to detect cancer, however, its low concentration in plasma can limit usefulness. Here, the authors devise a library preparation method from 100ul of plasma, and apply to multiple cancer types to detect and classify cancer patients
- Jun Wang
- , Jinyong Huang
- & Deming Gou
-
Article
| Open Access Pathway centric analysis for single-cell RNA-seq and spatial transcriptomics data with GSDensity
Pathway centric analysis for single-cell RNA-seq and spatial transcriptomics data with GSDensity
Clustering-based analysis has limited power in highly dynamic single-cell data, which is a common situation in tumour samples. Here, authors introduce GSDensity, enabling pathway-centric analysis for the direct integration of data with their domain knowledge.
- Qingnan Liang
- , Yuefan Huang
- & Ken Chen
-
Article
| Open Access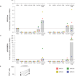 Potent latency reversal by Tat RNA-containing nanoparticle enables multi-omic analysis of the HIV-1 reservoir
Potent latency reversal by Tat RNA-containing nanoparticle enables multi-omic analysis of the HIV-1 reservoir
Reactivating latent HIV reservoirs could be beneficial towards a functional cure. Here, the authors show that Tat-LNP effectively reactivates HIV while preserving the cell transcriptome. Upon reactivation, p24+ cells exhibit distinct genes and pathways potentially contributing to their persistence.
- Marion Pardons
- , Basiel Cole
- & Linos Vandekerckhove
-
Article
| Open Access A patterned human primitive heart organoid model generated by pluripotent stem cell self-organization
A patterned human primitive heart organoid model generated by pluripotent stem cell self-organization
Pluripotent stem cell-derived organoids can recapitulate significant hallmarks of human organ development and are becoming critical tools for human research. Here, the authors report significant technical steps for generating sophisticated synthetic human primitive heart organoids.
- Brett Volmert
- , Artem Kiselev
- & Aitor Aguirre
-
Article
| Open Access STalign: Alignment of spatial transcriptomics data using diffeomorphic metric mapping
STalign: Alignment of spatial transcriptomics data using diffeomorphic metric mapping
Spatial transcriptomics (ST) enables gene expression characterisation within tissue sections, but comparing across sections and technologies remains challenging. Here, authors develop STalign to spatially align ST data and demonstrate applications including aligning to common coordinate frameworks.
- Kalen Clifton
- , Manjari Anant
- & Jean Fan
-
Article
| Open Access Secretory GFP reconstitution labeling of neighboring cells interrogates cell–cell interactions in metastatic niches
Secretory GFP reconstitution labeling of neighboring cells interrogates cell–cell interactions in metastatic niches
Methodologies to study the mechanisms of cell–cell interactions in metastatic niches remain scarce. Here, the authors develop a secretory GFP reconstitution-based system to tag tissue-resident cells neighboring on cancer cells within the metastatic niche.
- Misa Minegishi
- , Takahiro Kuchimaru
- & Satoshi Nishimura
-
Article
| Open Access Spatial transcriptomics deconvolution at single-cell resolution using Redeconve
Spatial transcriptomics deconvolution at single-cell resolution using Redeconve
Computational deconvolution with single-cell RNA sequencing data as a reference is pivotal for interpreting spatial transcriptomics data. Here, authors present Redeconve, which improves the resolution by more than 100-fold with higher accuracy and speed.
- Zixiang Zhou
- , Yunshan Zhong
- & Xianwen Ren
-
Article
| Open Access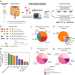 Detection of isoforms and genomic alterations by high-throughput full-length single-cell RNA sequencing in ovarian cancer
Detection of isoforms and genomic alterations by high-throughput full-length single-cell RNA sequencing in ovarian cancer
Long-read single-cell RNA sequencing is capable of detecting isoform-level gene expression and genomic alterations such as mutations and gene fusions, thereby providing cell-specific genotype-phenotype information. Here, the authors use long-read scRNA-seq on metastatic ovarian cancer samples and detect cell-type specific isoforms and gene fusions that may otherwise be misclassified in short-read data.
- Arthur Dondi
- , Ulrike Lischetti
- & Niko Beerenwinkel
-
Article
| Open Access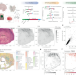 High-sensitive spatially resolved T cell receptor sequencing with SPTCR-seq
High-sensitive spatially resolved T cell receptor sequencing with SPTCR-seq
Understanding T cell behaviour in cancers is vital for improving immunotherapies. Here, the authors present spatially resolved T cell receptor sequencing (SPTCR-seq), a technology that annotates T cell receptors within the tumour ecosystem.
- Jasim Kada Benotmane
- , Jan Kueckelhaus
- & Dieter Henrik Heiland
-
Article
| Open Access Mucosal-associated invariant T cells contribute to suppression of inflammatory myeloid cells in immune-mediated kidney disease
Mucosal-associated invariant T cells contribute to suppression of inflammatory myeloid cells in immune-mediated kidney disease
Mucosal associated invariant T (MAIT) cells reside in barrier organs, but their contribution to inflammatory processes in the kidneys is not fully known. Here authors find by single cell RNA sequencing that among the different MAIT cell subtypes found at steady state, a population with MAIT17 signature is expanded in both human crescentic glomerulonephritis and its mouse model, and these cells may play protective role in the disease.
- Ann-Christin Gnirck
- , Marie-Sophie Philipp
- & Jan-Eric Turner
-
Article
| Open Access Dimension-agnostic and granularity-based spatially variable gene identification using BSP
Dimension-agnostic and granularity-based spatially variable gene identification using BSP
Identifying spatially variable genes (SVGs) is essential for linking molecular cell functions with tissue phenotypes. Here, authors introduce a non-parametric model that detects SVGs from two or three-dimensional spatial transcriptomics data by comparing gene expression patterns at granularities.
- Juexin Wang
- , Jinpu Li
- & Dong Xu
-
Article
| Open Access EASTR: Identifying and eliminating systematic alignment errors in multi-exon genes
EASTR: Identifying and eliminating systematic alignment errors in multi-exon genes
The study reveals limitations in widely used RNA-seq aligners, which create 'phantom' introns in reference databases. The authors introduce EASTR, a computational tool that not only enhances alignment accuracy but also uncovers existing annotation errors. This improvement bolsters the dependability of subsequent RNA-seq analyses.
- Ida Shinder
- , Richard Hu
- & Mihaela Pertea
-
Article
| Open Access MePMe-seq: antibody-free simultaneous m6A and m5C mapping in mRNA by metabolic propargyl labeling and sequencing
MePMe-seq: antibody-free simultaneous m6A and m5C mapping in mRNA by metabolic propargyl labeling and sequencing
Methylation is the dominant modification in mRNA and occurs at a variety of sites. Here, Hartstock et al. show that a clickable analogue of the key cosubstrate S-adenosyl-L-methionine (SAM) can be produced in cells, allowing for identification and mapping of different methylated nucleosides in mRNA.
- Katja Hartstock
- , Nadine A. Kueck
- & Andrea Rentmeister
-
Article
| Open Access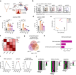 IL-21R-STAT3 signalling initiates a differentiation program in uterine tissue-resident NK cells to support pregnancy
IL-21R-STAT3 signalling initiates a differentiation program in uterine tissue-resident NK cells to support pregnancy
Uterine natural killer (NK) cells support tissue homeostasis in the uterus during pregnancy, but it is not fully known how they differentiate into potentially cytotoxic effector cells while avoiding tissue damage. Here authors show that Il21 receptor signalling via STAT3 activation governs their differentiation, while an apoptotic cell death program ensures that harm is limited.
- Mengwei Han
- , Luni Hu
- & Chao Zhong
-
Article
| Open Access Mass production of lumenogenic human embryoid bodies and functional cardiospheres using in-air-generated microcapsules
Mass production of lumenogenic human embryoid bodies and functional cardiospheres using in-air-generated microcapsules
Current methods to generate spheroids are associated with low production throughputs, limiting clinical and industrial translation. Here the authors present a clean ultra-high-throughput in-air microfluidic platform for mass production of lumenogenic embryoid bodies and functional cardiospheres.
- Bas van Loo
- , Simone A. ten Den
- & Jeroen Leijten
-
Article
| Open Access A spatial sequencing atlas of age-induced changes in the lung during influenza infection
A spatial sequencing atlas of age-induced changes in the lung during influenza infection
Ageing is known to impair the immune response against infectious pathogens. Here, Kasmani et al. present a spatial and transcriptomic atlas of immune changes in the lungs of young and aged mice in response to influenza virus infection.
- Moujtaba Y. Kasmani
- , Paytsar Topchyan
- & Weiguo Cui
-
Article
| Open Access mRNA vaccine quality analysis using RNA sequencing
mRNA vaccine quality analysis using RNA sequencing
mRNA vaccines must be rigorously analysed to measure their integrity and detect contaminants, which can be time-consuming and costly. Here, authors describe a method to analyse mRNA vaccine quality using long-read sequencing and a custom bioinformatic pipeline.
- Helen M. Gunter
- , Senel Idrisoglu
- & Tim R. Mercer
-
Article
| Open Access Systematic transcriptional analysis of human cell lines for gene expression landscape and tumor representation
Systematic transcriptional analysis of human cell lines for gene expression landscape and tumor representation
During preclinical drug development, the ability of cancer cell lines to faithfully model human disease is important for identifying potential therapeutic strategies. Here, using transcriptomic datasets of over 1000 cell lines, the authors evaluate how representative each line is of its cancer type and present their cell line selection tool.
- Han Jin
- , Cheng Zhang
- & Adil Mardinoglu
-
Article
| Open Access Projecting RNA measurements onto single cell atlases to extract cell type-specific expression profiles using scProjection
Projecting RNA measurements onto single cell atlases to extract cell type-specific expression profiles using scProjection
Many expression deconvolution approaches have been developed to estimate % RNA contributions of diverse cell types to mixed RNA measurements. Here, the authors have developed a complementary approach called scProjection to recover cell type-specific expression profiles from mixed RNA measurements.
- Nelson Johansen
- , Hongru Hu
- & Gerald Quon
-
Article
| Open Access Droplet-based high-throughput single microbe RNA sequencing by smRandom-seq
Droplet-based high-throughput single microbe RNA sequencing by smRandom-seq
Population level transcriptomics measurements miss bacterial heterogeneity. Here the authors report smRandom-seq, a droplet-based high-throughput single-microbe RNA-seq assay, using random primers for in situ cDNA generation, droplets for single-microbe barcoding, and CRISPR-based rRNA depletion.
- Ziye Xu
- , Yuting Wang
- & Yongcheng Wang
-
Article
| Open Access TEQUILA-seq: a versatile and low-cost method for targeted long-read RNA sequencing
TEQUILA-seq: a versatile and low-cost method for targeted long-read RNA sequencing
The authors report TEQUILA-seq, a versatile, easy-to-implement, and low-cost method for targeted long-read RNA sequencing. TEQUILA-seq uncovers transcript isoforms and RNA mechanisms associated with human health and disease.
- Feng Wang
- , Yang Xu
- & Lan Lin
-
Article
| Open Access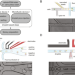 spinDrop: a droplet microfluidic platform to maximise single-cell sequencing information content
spinDrop: a droplet microfluidic platform to maximise single-cell sequencing information content
Droplet microfluidics enables high-throughput single-cell sequencing, but often with increased noise. Here the authors report spinDrop (sorting picoinjection inDrop) to increase gene detection and reduce noise; they use this to generate a high-quality molecular atlas of mouse brain development.
- Joachim De Jonghe
- , Tomasz S. Kaminski
- & Florian Hollfelder
-
Article
| Open Access SONAR enables cell type deconvolution with spatially weighted Poisson-Gamma model for spatial transcriptomics
SONAR enables cell type deconvolution with spatially weighted Poisson-Gamma model for spatial transcriptomics
Spatial transcriptomics reveal cellular profiles with spatial context. Here the authors present SONAR, a computational model that utilizes spatial information to decipher cell types in tissues and validate on various spatial patterns and fine-mapped cell types in complex tissues.
- Zhiyuan Liu
- , Dafei Wu
- & Liang Ma
-
Article
| Open Access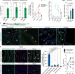 MAVS signaling is required for preventing persistent chikungunya heart infection and chronic vascular tissue inflammation
MAVS signaling is required for preventing persistent chikungunya heart infection and chronic vascular tissue inflammation
Mosquito-borne viruses are serious global public health threats associated with severe atypical cardiovascular manifestations. Here, the authors dissect how chikungunya virus directly infects cardiac tissue leading to heart disease and define key host pathways involved in viral cardiac persistence and tissue damage.
- Maria G. Noval
- , Sophie N. Spector
- & Kenneth A. Stapleford
-
Article
| Open Access Guided construction of single cell reference for human and mouse lung
Guided construction of single cell reference for human and mouse lung
Accurate cell-type identification is vital for single-cell analysis. Here, the authors develop a computational pipeline called “LungMAP CellRef” for efficient, automated cell-type annotation of normal and disease human and mouse lung single-cell datasets.
- Minzhe Guo
- , Michael P. Morley
- & Yan Xu
-
Article
| Open Access High throughput single cell long-read sequencing analyses of same-cell genotypes and phenotypes in human tumors
High throughput single cell long-read sequencing analyses of same-cell genotypes and phenotypes in human tumors
There is a need for methods that allow the analysis of single-cell long-read sequencing data without depending on known barcode lists or short-read sequencing. Here, the authors develop scNanoGPS, a tool that can independently deconvolute long reads into single cells and single molecules, and apply it on tumour and cell line data.
- Cheng-Kai Shiau
- , Lina Lu
- & Ruli Gao
-
Article
| Open Access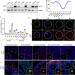 Maternal NAT10 orchestrates oocyte meiotic cell-cycle progression and maturation in mice
Maternal NAT10 orchestrates oocyte meiotic cell-cycle progression and maturation in mice
Generation of mature oocytes requires tight regulation of a discontinuous meiotic cell cycle. Here they show that the acetyltransferase Nat10 mediates modification of RNAs targeted for degradation and find that this process is essential for female oocyte meiosis and maturation.
- Xue Jiang
- , Yu Cheng
- & Jianqiang Bao
-
Article
| Open Access Co-translational binding of importins to nascent proteins
Co-translational binding of importins to nascent proteins
Importins are known to facilitate nucleocytoplasmic transport and cytoplasmic chaperoning of some proteins. Here, the authors uncover that these proteins also act as co-translational chaperones for specific sets of proteins, for example ribonucleic acid binding factors.
- Maximilian Seidel
- , Natalie Romanov
- & Martin Beck
-
Article
| Open Access Systematic detection of tertiary structural modules in large RNAs and RNP interfaces by Tb-seq
Systematic detection of tertiary structural modules in large RNAs and RNP interfaces by Tb-seq
Compact RNA structural motifs control many aspects of gene expression, but methods for their identification are lacking. Here the authors present a sequencing-based terbium probing approach to detect complex 3D structural elements, which can be used to pinpoint potential riboregulatory elements.
- Shivali Patel
- , Alec N. Sexton
- & Anna Marie Pyle
-
Article
| Open Access A village in a dish model system for population-scale hiPSC studies
A village in a dish model system for population-scale hiPSC studies
Village cultures, where multiple stem cell lines are cultured in a single dish, provide an elegant solution for population-scale studies. Here, authors show the utility of village models – showing that expression heterogeneity is largely a result of line-specific effects and not village cultures.
- Drew R. Neavin
- , Angela M. Steinmann
- & Joseph E. Powell
-
Article
| Open Access PIE-seq: identifying RNA-binding protein targets by dual RNA-deaminase editing and sequencing
PIE-seq: identifying RNA-binding protein targets by dual RNA-deaminase editing and sequencing
Tracking protein-RNA interaction across cell types is challenging. Here, Ruan et al develop a dual-deaminase method called PIE-Seq, where protein targets are marked by both C-to-U and A-to-I RNA base editors, and apply it to 25 human RNA-binding proteins.
- Xiangbin Ruan
- , Kaining Hu
- & Xiaochang Zhang
-
Article
| Open Access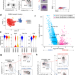 Single cell transcriptomics clarifies the basophil differentiation trajectory and identifies pre-basophils upstream of mature basophils
Single cell transcriptomics clarifies the basophil differentiation trajectory and identifies pre-basophils upstream of mature basophils
Single cell sequencing can be used to better characterize immune cell progenitors. Here the authors characterize CLEC12Ahi pre-basophils downstream of pre-basophil and mast cell progenitors (pre-BMPs) but upstream of mature basophils and this population includes basophil progenitors (BaPs).
- Kensuke Miyake
- , Junya Ito
- & Hajime Karasuyama

 scButterfly: a versatile single-cell cross-modality translation method via dual-aligned variational autoencoders
scButterfly: a versatile single-cell cross-modality translation method via dual-aligned variational autoencoders