Article
|
Open Access
Featured
-
-
Article
| Open Access Insights into the modulation of bacterial NADase activity by phage proteins
Insights into the modulation of bacterial NADase activity by phage proteins
The defense-associated sirtuin 2 (DSR2) effector protects bacteria from phage infection by depleting NAD+. Here, the authors employ biochemical and structural approaches to reveal the inhibition and activation mechanisms of DSR2 by the phage anti-DSR2 protein (DSAD1) and tail tube protein (TTP).
- Hang Yin
- , Xuzichao Li
- & Heng Zhang
-
Article
| Open Access Bacteriophage DNA induces an interrupted immune response during phage therapy in a chicken model
Bacteriophage DNA induces an interrupted immune response during phage therapy in a chicken model
Bacteriophage are potential therapeutics to target bacterial infections, but recent studies suggest that bacteriophage may induce immune responses in eukaryotic cells. Here the authors show that bacteriophage DNA induces interrupted host immunity in a chicken infection model.
- Magdalena Podlacha
- , Lidia Gaffke
- & Alicja Węgrzyn
-
Article
| Open Access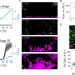 Cell-lysis sensing drives biofilm formation in Vibrio cholerae
Cell-lysis sensing drives biofilm formation in Vibrio cholerae
Bacteria form matrix-encapsulated communities, called biofilms, which protect resident cells from environmental challenges. Here, the authors show that Vibrio cholerae cells detect environmental threats by sensing a cellular component released through kin cell lysis, which induces formation of biofilms by surviving cells.
- Jojo A. Prentice
- , Robert van de Weerd
- & Andrew A. Bridges
-
Article
| Open Access Anti-phage defence through inhibition of virion assembly
Anti-phage defence through inhibition of virion assembly
Bacteria have evolved various defence mechanisms to protect themselves against viral infection. Here, Patel et al. identify a type of antiviral defence that blocks virion assembly, thus preventing formation of infectious virions and allowing destruction of the infected cell.
- Pramalkumar H. Patel
- , Véronique L. Taylor
- & Karen L. Maxwell
-
Article
| Open Access Phage-plasmids promote recombination and emergence of phages and plasmids
Phage-plasmids promote recombination and emergence of phages and plasmids
Phage-plasmids are mobile genetic elements that transfer horizontally between bacterial cells as viruses, and vertically within bacterial lineages as plasmids. Here, Pfeifer & Rocha show that phage-plasmids can mediate gene transfer across mobile elements within their hosts, and can act as intermediates in the conversion of one type of element into another.
- Eugen Pfeifer
- & Eduardo P. C. Rocha
-
Article
| Open Access Structural and biochemical insights into the mechanism of the Gabija bacterial immunity system
Structural and biochemical insights into the mechanism of the Gabija bacterial immunity system
The Gabija system is a newly discovered bacterial immune system. Here, the authors report the EM structure of the Gabija complex at 3.6 Å. Here, the authors show that when invading phages depletes cellular NTP and dNTP, the nuclease activity of Gabija complex is activated and cleaves the circular DNA to prevent phage DNA replication.
- Yanwu Huo
- , Lingfei Kong
- & Taotao Wei
-
Article
| Open Access Phage-inducible chromosomal minimalist islands (PICMIs), a novel family of small marine satellites of virulent phages
Phage-inducible chromosomal minimalist islands (PICMIs), a novel family of small marine satellites of virulent phages
Phage satellites are bacterial genetic elements that co-opt phage machinery for their own dissemination. Here, Barcia-Cruz et al. identify a family of satellites, named PICMIs, that are characterized by reduced gene content and are broadly distributed in marine bacteria of the family Vibrionaceae.
- Rubén Barcia-Cruz
- , David Goudenège
- & Frédérique Le Roux
-
Article
| Open Access Structure-guided discovery of anti-CRISPR and anti-phage defense proteins
Structure-guided discovery of anti-CRISPR and anti-phage defense proteins
Bacteria use various defense systems to protect themselves from phage infection, and phages have evolved diverse counter-defense measures to overcome host defenses. Here, the authors use protein structural similarity and gene co-occurrence analyses for identification of new anti-phage and counter-defense systems.
- Ning Duan
- , Emily Hand
- & Akintunde Emiola
-
Article
| Open Access Biophysical basis of filamentous phage tactoid-mediated antibiotic tolerance in P. aeruginosa
Biophysical basis of filamentous phage tactoid-mediated antibiotic tolerance in P. aeruginosa
Filamentous phages can assemble into mesoscale structures termed tactoids that protect bacteria in biofilms from antibiotics. Here, the authors dissect the biophysical factors influencing this protection using two model phages, Pf4 and fd.
- Jan Böhning
- , Miles Graham
- & Tanmay A. M. Bharat
-
Article
| Open Access Nearly complete structure of bacteriophage DT57C reveals architecture of head-to-tail interface and lateral tail fibers
Nearly complete structure of bacteriophage DT57C reveals architecture of head-to-tail interface and lateral tail fibers
The authors present the nearly-complete structure of the DT57C bacteriophage of the Siphovirus family, revealing the molecular architecture of its capsid, neck, tail and tail tip, and providing insights into the process of DNA ejection.
- Rafael Ayala
- , Andrey V. Moiseenko
- & Matthias Wolf
-
Article
| Open Access Ongoing shuffling of protein fragments diversifies core viral functions linked to interactions with bacterial hosts
Ongoing shuffling of protein fragments diversifies core viral functions linked to interactions with bacterial hosts
Proteins are composed of distinct functional domains, each serving a specific role. Here, Smug et al. show that phages are able to shuffle fragments of their proteins and this predominantly occurs in proteins involved in bacterial host interactions.
- Bogna J. Smug
- , Krzysztof Szczepaniak
- & Rafał J. Mostowy
-
Article
| Open Access The ClpX protease is essential for inactivating the CI master repressor and completing prophage induction in Staphylococcus aureus
The ClpX protease is essential for inactivating the CI master repressor and completing prophage induction in Staphylococcus aureus
Prophage induction is a fundamental process in the phage life cycle. Here the authors show that the final stage of prophage induction in Staphylococcus aureus is controlled by the ClpX protease, unveiling and unexpected role for ClpX in phage biology.
- Mohammed A. Thabet
- , José R. Penadés
- & Andreas F. Haag
-
Article
| Open Access Distantly related Alteromonas bacteriophages share tail fibers exhibiting properties of transient chaperone caps
Distantly related Alteromonas bacteriophages share tail fibers exhibiting properties of transient chaperone caps
Receptor binding proteins in bacteriophage tails mediate host recognition and thus infectivity. Here Gonzalez et al. identify a receptor binding module shared by different phages that all use receptor binding tail fibers that are transiently capped by their chaperones.
- Rafael Gonzalez-Serrano
- , Riccardo Rosselli
- & Matthew Dunne
-
Article
| Open Access A systematic analysis of marine lysogens and proviruses
A systematic analysis of marine lysogens and proviruses
Viruses are ubiquitous in the oceans, exhibiting high abundance and diversity. Here, Yi et al. present a systematic catalogue and analysis of genomic sequences from marine prokaryotes and their proviruses, thus contributing to a better understanding of the ecology of these microorganisms.
- Yi Yi
- , Shunzhang Liu
- & Huahua Jian
-
Article
| Open Access Next-generation proteomics for quantitative Jumbophage-bacteria interaction mapping
Next-generation proteomics for quantitative Jumbophage-bacteria interaction mapping
In this study Fossati et al. demonstrate how co-fractionation mass spectrometry can be applied to systematically investigate pathogen proteome organization and host interactome plasticity upon Jumbophage infection of Pseudomonas aeruginosa.
- Andrea Fossati
- , Deepto Mozumdar
- & Danielle L. Swaney
-
Article
| Open Access Enhancing bacteriophage therapeutics through in situ production and release of heterologous antimicrobial effectors
Enhancing bacteriophage therapeutics through in situ production and release of heterologous antimicrobial effectors
Du et al. genetically engineer bacteriophages into heterologous effector phage therapeutics, enabling dual phage- and effector-mediated targeting for a two-pronged attack against bacterial pathogens.
- Jiemin Du
- , Susanne Meile
- & Matthew Dunne
-
Article
| Open Access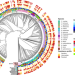 Roving methyltransferases generate a mosaic epigenetic landscape and influence evolution in Bacteroides fragilis group
Roving methyltransferases generate a mosaic epigenetic landscape and influence evolution in Bacteroides fragilis group
Here, Tisza, Dekker, and colleagues perform large scale analysis of genome methylation in the gut commensal and pathogen, Bacteroides fragilis group, revealing immense methyl motif diversity and evidence of widespread methyltransferase exchange among phages.
- Michael J. Tisza
- , Derek D. N. Smith
- & John P. Dekker
-
Article
| Open Access High-resolution cryo-EM structure of the Pseudomonas bacteriophage E217
High-resolution cryo-EM structure of the Pseudomonas bacteriophage E217
E217 is a Myoviridae used in an experimental phage cocktail to eradicate Pseudomonas aeruginosa. Here, the authors utilize cryo-EM and functional analysis to delineate E217 structural proteins, tail dynamics, and mechanisms of host recognition.
- Fenglin Li
- , Chun-Feng David Hou
- & Gino Cingolani
-
Article
| Open Access Structure and proposed DNA delivery mechanism of a marine roseophage
Structure and proposed DNA delivery mechanism of a marine roseophage
Tailed bacteriophages account for the majority of all phages. Here, the authors employ cryo-EM and structure prediction techniques to investigate the atomic structure of the R4C phage capsid and the in- situ structure of its unique long rigid tail.
- Yang Huang
- , Hui Sun
- & Ningshao Xia
-
Article
| Open Access Design of bacteriophage T4-based artificial viral vectors for human genome remodeling
Design of bacteriophage T4-based artificial viral vectors for human genome remodeling
Safe delivery of genes is needed for gene therapy. Here the authors build “artificial viral vectors” (AVVs) by engineering the well-characterised structural components of bacteriophage T4: the large capacity, all-in-one, multiplex, programmable, and phage-based AVV nanomaterials have potential for gene therapy.
- Jingen Zhu
- , Himanshu Batra
- & Venigalla B. Rao
-
Article
| Open Access Cryo-electron microscopy of the f1 filamentous phage reveals insights into viral infection and assembly
Cryo-electron microscopy of the f1 filamentous phage reveals insights into viral infection and assembly
In this work, the authors report a system for production of short versions of a filamentous phage enables the structure to be determined by cryo-electron microscopy. Structure combined with mutagenesis allows the identification of phage domains that are important in bacterial attack and for release of new viral progeny.
- Rebecca Conners
- , Rayén Ignacia León-Quezada
- & Vicki A. M. Gold
-
Article
| Open Access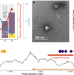 Mutation-induced infections of phage-plasmids
Mutation-induced infections of phage-plasmids
Phage-plasmids are bacterial extrachromosomal elements that act both as plasmids and as viruses. Here, Shan et al. show that segregational drift and loss-of-function mutations play key roles in the infection dynamics of a cosmopolitan phage-plasmid, allowing it to create continuous productive infections in marine bacteria.
- Xiaoyu Shan
- , Rachel E. Szabo
- & Otto X. Cordero
-
Article
| Open Access Topical phage therapy in a mouse model of Cutibacterium acnes-induced acne-like lesions
Topical phage therapy in a mouse model of Cutibacterium acnes-induced acne-like lesions
Bacteriophage therapy is evolving as a promising approach to tackling bacterial infection, even in the case of emerging antibiotic resistance. In this work, authors present the topical application of numerous Cutibacterium acnes phage in an in vivo mouse model of acne vulgaris.
- Amit Rimon
- , Chani Rakov
- & Ronen Hazan
-
Article
| Open Access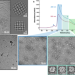 The ϕPA3 phage nucleus is enclosed by a self-assembling 2D crystalline lattice
The ϕPA3 phage nucleus is enclosed by a self-assembling 2D crystalline lattice
To protect from host attack, numerous jumbo bacteriophages establish a micron-scale, protein-based structure to enclose their replicating DNA. Using cryoEM, the authors show that the 2D crystal enclosing this so-called phage nucleus is an assembly of tetramers linked by flexible loops and tails.
- Eliza S. Nieweglowska
- , Axel F. Brilot
- & David A. Agard
-
Article
| Open Access The coordination of anti-phage immunity mechanisms in bacterial cells
The coordination of anti-phage immunity mechanisms in bacterial cells
Bacteria are equipped with diverse immune strategies to fight bacteriophage infections, including restriction nucleases, abortive infection and CRISPR-Cas systems. Here, Arias et al. use mathematical models of immune responses in individual bacterial cells to highlight the importance of the timing and coordination of different antiviral systems, and present hypotheses that may inspire future research.
- Clemente F. Arias
- , Francisco J. Acosta
- & Cristina Fernández-Arias
-
Article
| Open Access CryoEM structure and assembly mechanism of a bacterial virus genome gatekeeper
CryoEM structure and assembly mechanism of a bacterial virus genome gatekeeper
Numerous viruses use a portal system for dsDNA entry and exit from their capsid. Here the authors report the atomic structure of phage SPP1 portal DNA gatekeeper and its mechanism of assembly. They also identify evolution breakpoints between different tailed bacteriophages morphotypes and herpesviruses.
- Igor Orlov
- , Stéphane Roche
- & Elena V. Orlova
-
Article
| Open Access High-resolution reconstruction of a Jumbo-bacteriophage infecting capsulated bacteria using hyperbranched tail fibers
High-resolution reconstruction of a Jumbo-bacteriophage infecting capsulated bacteria using hyperbranched tail fibers
The jumbo contractile bacteriophage, Kp24, has been isolated from clinical strains of Klebsiella pneumonia. Here, the authors present structural and functional insight into the capsid, tail and tail fibres and how this impacts infectivity.
- Ruochen Ouyang
- , Ana Rita Costa
- & Ariane Briegel
-
Article
| Open Access Experimental validation that human microbiome phages use alternative genetic coding
Experimental validation that human microbiome phages use alternative genetic coding
Previous bioinformatic analyses have indicated that bacteriophages can use genetic codes different from those of their host bacteria. Here, Peters et al. use metaproteomics to provide experimental evidence of reassignment of stop codon TAG to glutamine in phages found in the human gut microbiome.
- Samantha L. Peters
- , Adair L. Borges
- & Robert L. Hettich
-
Article
| Open Access Tail proteins of phage SU10 reorganize into the nozzle for genome delivery
Tail proteins of phage SU10 reorganize into the nozzle for genome delivery
E. coli phage SU10 has a short non-contractile tail. Here, the authors show that after cell binding, nozzle proteins and tail fibers of SU10 change conformation to form a nozzle that enables the delivery of the phage DNA into the bacterial cytoplasm.
- Marta Šiborová
- , Tibor Füzik
- & Pavel Plevka
-
Article
| Open Access Insights into the mechanism of action of the arbitrium communication system in SPbeta phages
Insights into the mechanism of action of the arbitrium communication system in SPbeta phages
The arbitrium system is a peptide-based communication system to coordinate the lysis-lysogenic cycle of phages infecting bacteria. Here Gallego del Sol et al. provide the crystal structure of Aim-receptor of Katmira phage infecting B. subtilis in apo, peptide-bound and DNA-bound form.
- Francisca Gallego del Sol
- , Nuria Quiles-Puchalt
- & Alberto Marina
-
Article
| Open Access Gut virome profiling identifies a widespread bacteriophage family associated with metabolic syndrome
Gut virome profiling identifies a widespread bacteriophage family associated with metabolic syndrome
Here, the authors characterize gut viromes in a cohort of individuals with metabolic syndrome, which they find associated with a highly widespread family of gut bacteriophages they name Candidatus Heliusviridae.
- Patrick A. de Jonge
- , Koen Wortelboer
- & Hilde Herrema
-
Article
| Open Access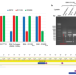 A truncated anti-CRISPR protein prevents spacer acquisition but not interference
A truncated anti-CRISPR protein prevents spacer acquisition but not interference
Phages can use ACR proteins that inhibit the adaptive immunity activities of bacterial CRISPR-Cas systems. Here, Philippe et al. show that these systems can block ACR-containing phages by targeting the acr gene, and this can select for phage mutants carrying a deletion within acr that does not block DNA cleavage (interference) but prevents the addition of new immunity (spacer acquisition).
- Cécile Philippe
- , Carlee Morency
- & Sylvain Moineau
-
Article
| Open Access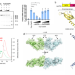 Insights into the inhibition of type I-F CRISPR-Cas system by a multifunctional anti-CRISPR protein AcrIF24
Insights into the inhibition of type I-F CRISPR-Cas system by a multifunctional anti-CRISPR protein AcrIF24
Phages use anti-CRISPR proteins (Acrs) to counteract the bacterial CRISPR-Cas systems. Here, the authors characterize AcrIF24, which functions as an Aca (Acr-associated) to repress and regulate its own transcription, dimerizes the Csy complex, blocks the hybridization of target DNA, and tethers non-sequence-specific DNA to the Csy complex.
- Lingguang Yang
- , Laixing Zhang
- & Yue Feng
-
Article
| Open Access Genome binning of viral entities from bulk metagenomics data
Genome binning of viral entities from bulk metagenomics data
Here, Johansen et al. develop an approach, Phages from Metagenomics Binning (PHAMB), that allows the binning of thousands of viral genomes directly from bulk metagenomics data, while simultaneously enabling clustering of viral genomes into accurate taxonomic viral populations, unveiling viral-microbial host interactions in the gut.
- Joachim Johansen
- , Damian R. Plichta
- & Simon Rasmussen
-
Article
| Open Access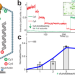 A viral genome packaging ring-ATPase is a flexibly coordinated pentamer
A viral genome packaging ring-ATPase is a flexibly coordinated pentamer
In viruses, multi-subunit ring-ATPases are involved in genome packaging. Here, using single-molecule techniques, the authors determine that the active bacteriophage T4 DNA packaging motor is a pentamer and show that the motor can tolerate inactive subunits, suggesting that strict coordination between the subunits is not crucial.
- Li Dai
- , Digvijay Singh
- & Venigalla B. Rao
-
Comment
| Open Access Is the bacterial chromosome a mobile genetic element?
Is the bacterial chromosome a mobile genetic element?
An outcome of phage infection, lateral transduction, has been shown to mobilize chromosomal genes between bacterial cells at rates that exceed those of mobile genetic elements such as plasmids. Does this mean that the bacterial chromosome should be considered a mobile genetic element?
- James P. J. Hall
-
Article
| Open Access CryoEM structure of the outer membrane secretin channel pIV from the f1 filamentous bacteriophage
CryoEM structure of the outer membrane secretin channel pIV from the f1 filamentous bacteriophage
New virions of Ff bacteriophages are extruded from the host cell via the channel built from phage protein pIV, homologous to bacterial secretins. Here, the authors report the structure of this channel from the f1 filamentous bacteriophage and propose its use as an adjuvant to increase the uptake and efficacy of antibiotics.
- Rebecca Conners
- , Mathew McLaren
- & Vicki A. M. Gold
-
Article
| Open Access Ecology and molecular targets of hypermutation in the global microbiome
Ecology and molecular targets of hypermutation in the global microbiome
Here, the authors report a large-scale comparative analysis of <30,000 Diversity-Generating Retroelements (DGRs) across ~9000 metagenomes (representing diverse taxa and biomes), to identify patterns in terms of prevalence and activity. Combined with examination of longitudinal data on <100 metagenomes part of time series, they demonstrate that DGRs are broadly and consistently active, implying an important role in microbiota ecology and evolution.
- Simon Roux
- , Blair G. Paul
- & Emiley A. Eloe-Fadrosh
-
Article
| Open Access Analysis of metagenome-assembled viral genomes from the human gut reveals diverse putative CrAss-like phages with unique genomic features
Analysis of metagenome-assembled viral genomes from the human gut reveals diverse putative CrAss-like phages with unique genomic features
Here, the authors analyze 4907 Circular Metagenome Assembled Genomes from human microbiomes and identify and characterize nearly 600 diverse genomes of crAss-like phages, finding two putative families with unusual genomic features, including high density of self-splicing introns and inteins.
- Natalya Yutin
- , Sean Benler
- & Eugene V. Koonin
-
Article
| Open Access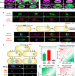 Viral speciation through subcellular genetic isolation and virogenesis incompatibility
Viral speciation through subcellular genetic isolation and virogenesis incompatibility
Virus speciation cannot be fully explained by the evolution of different host specificities. Here, Chaikeeratisak et al. identify ways viruses can remain genetically isolated despite co-infecting the same cell, providing insight into how new virus species evolve.
- Vorrapon Chaikeeratisak
- , Erica A. Birkholz
- & Joe Pogliano
-
Article
| Open Access Flexible genes establish widespread bacteriophage pan-genomes in cryoconite hole ecosystems
Flexible genes establish widespread bacteriophage pan-genomes in cryoconite hole ecosystems
Bacteriophages and their hosts are involved in a constant evolutionary arms race that should lead to divergence between phage genes over time. Here, the authors recruit metagenomic reads to virus reference genomes and genome fragments in samples from cryoconite holes and show that phages with near-identical core genomes maintain diversity by possession of numerous flexible gene modules, where homologous genes present in the pan-genome interchange to create new phage variants.
- Christopher M. Bellas
- , Declan C. Schroeder
- & Alexandre M. Anesio
-
Article
| Open Access The architecture and stabilisation of flagellotropic tailed bacteriophages
The architecture and stabilisation of flagellotropic tailed bacteriophages
Flagellotropic phages spin down flagella to reach the bacterial surface and must withstand remarkable drag forces. Here authors show how two nested sets of chainmail stabilise the viral head and a beta-hairpin regulates the formation of the robust yet pliable tail, characteristic of siphoviruses.
- Joshua M. Hardy
- , Rhys A. Dunstan
- & Fasséli Coulibaly
-
Article
| Open Access Structure and mechanism of DNA delivery of a gene transfer agent
Structure and mechanism of DNA delivery of a gene transfer agent
Gene transfer agents (GTAs) are phage-like particles that mediate lateral gene exchange. Here, the authors provide the structure of the GTA of Rhodobacter capsulatus (RcGTA), which resembles a tailed phage, and describe the conformational changes required for DNA ejection.
- Pavol Bárdy
- , Tibor Füzik
- & Pavel Plevka
-
Article
| Open Access Structural morphing in a symmetry-mismatched viral vertex
Structural morphing in a symmetry-mismatched viral vertex
In icosahedral viruses, a symmetry-mismatched portal vertex is assembled by inserting a 12-fold-symmetric portal complex into a 5-fold-symmetric capsid environment. Here, the authors report a near-atomic-resolution in situ cryo-electron microscopy structure of this symmetrically mismatched viral vertex from bacteriophage T4.
- Qianglin Fang
- , Wei-Chun Tang
- & Venigalla B. Rao
-
Article
| Open Access Virulent coliphages in 1-year-old children fecal samples are fewer, but more infectious than temperate coliphages
Virulent coliphages in 1-year-old children fecal samples are fewer, but more infectious than temperate coliphages
The impact of bacteriophages in the human gut microbiome remains poorly understood. Here, the authors characterize coliphages isolated from a large cohort of 1-year-old infants and show that temperate coliphages dominate, while virulent ones have greater infectivity and broader host range.
- Aurélie Mathieu
- , Moïra Dion
- & Marie-Agnès Petit
-
Article
| Open Access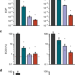 Type I-F CRISPR-Cas resistance against virulent phages results in abortive infection and provides population-level immunity
Type I-F CRISPR-Cas resistance against virulent phages results in abortive infection and provides population-level immunity
Bacteria use CRISPR-Cas systems to protect themselves against viral infections. Here, Watson et al. show that a type I CRISPR-Cas system can induce abortive viral infection, where infected cells do not survive but viral propagation is decreased, thus protecting the bacterial population.
- Bridget N. J. Watson
- , Reuben B. Vercoe
- & Peter C. Fineran
-
Article
| Open Access Coordination of cohabiting phage elements supports bacteria–phage cooperation
Coordination of cohabiting phage elements supports bacteria–phage cooperation
Bacterial pathogens often carry multiple phage-derived elements within their genome. Here, the authors show that two phage elements are co-regulated in Listeria monocytogenes, the first one controlling the induction of the second one, which in turn regulates virulence of their bacterial host.
- Tal Argov
- , Shai Ran Sapir
- & Anat A. Herskovits
-
Article
| Open Access Prophages and satellite prophages are widespread in Streptococcus and may play a role in pneumococcal pathogenesis
Prophages and satellite prophages are widespread in Streptococcus and may play a role in pneumococcal pathogenesis
Prophages are viral genomes integrated within bacterial genomes. Here, Rezaei Javan et al. identify nearly 800 prophages and satellite prophages in > 1300 Streptococcus genomes, and show that a satellite prophage is associated with virulence in a mouse model of pneumococcal infection.
- Reza Rezaei Javan
- , Elisa Ramos-Sevillano
- & Angela B. Brueggemann
-
Article
| Open Access The structure of a polygamous repressor reveals how phage-inducible chromosomal islands spread in nature
The structure of a polygamous repressor reveals how phage-inducible chromosomal islands spread in nature
Staphylococcus aureus pathogenicity islands (SaPIs) encode the master repressor Stl and after bacteriophage infection Stl interacts with specific phage proteins leading to a derepression of SaPIs. Here the authors provide structural insights into this family of repressors by determining the crystal structures of SaPIbov1 Stl alone and in complex with two structurally unrelated phage dUTPases.
- J. Rafael Ciges-Tomas
- , Christian Alite
- & Alberto Marina

 Diverse and abundant phages exploit conjugative plasmids
Diverse and abundant phages exploit conjugative plasmids