Featured
-
-
Comment
| Open Access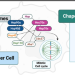 Epichaperomics reveals dysfunctional chaperone protein networks
Epichaperomics reveals dysfunctional chaperone protein networks
Molecular chaperones establish essential protein-protein interaction networks. Modified versions of these assemblies are generally enriched in certain maladies. A study published in Nature Communications used epichaperomics to identify unique changes occurring in chaperone-formed protein networks during mitosis in cancer cells.
- Mark R. Woodford
- , Dimitra Bourboulia
- & Mehdi Mollapour
-
Article
| Open Access Systems-level analyses of protein-protein interaction network dysfunctions via epichaperomics identify cancer-specific mechanisms of stress adaptation
Systems-level analyses of protein-protein interaction network dysfunctions via epichaperomics identify cancer-specific mechanisms of stress adaptation
Epichaperomics allow the study of protein-protein interactions and their alterations, but probes have been limited to capturing HSP90 epichaperomes. Here, the authors introduce and validate a toolset of HSP70 epichaperome ligands, and use them in epichaperomics to identify a mechanism with which cancer cells can enhance the fitness of mitotic protein networks.
- Anna Rodina
- , Chao Xu
- & Gabriela Chiosis
-
Article
| Open Access Disrupting the α-synuclein-ESCRT interaction with a peptide inhibitor mitigates neurodegeneration in preclinical models of Parkinson’s disease
Disrupting the α-synuclein-ESCRT interaction with a peptide inhibitor mitigates neurodegeneration in preclinical models of Parkinson’s disease
ESCRT-III is involved in the endolysosomal system and disturbed in neurodegenerative diseases. Here the authors show that disruption of an interaction between ESCRT-III member CHMP2B and α-synuclein by a peptide inhibitor mitigates neurodegeneration in Parkinson’s disease models.
- Satra Nim
- , Darren M. O’Hara
- & Philip M. Kim
-
Article
| Open Access Signal processing and generation of bioactive nitric oxide in a model prototissue
Signal processing and generation of bioactive nitric oxide in a model prototissue
A challenge for synthetic biology is the design and construction of prototissue. Here, the authors spatially segregate layers of enzyme-decorated coacervate protocells as a model prototissue capable of chemical signal processing and modulating outputs of nitric oxide to inhibit blood clot formation.
- Songyang Liu
- , Yanwen Zhang
- & Jianbo Liu
-
Article
| Open Access De novo biosynthesis of rubusoside and rebaudiosides in engineered yeasts
De novo biosynthesis of rubusoside and rebaudiosides in engineered yeasts
Rubusoside and rebaudiosides are considered the next generation of sugar substitutes. In this article, the authors report the engineering of Saccharomyces cerevisiae, remodelling the complex metabolic networks by a modular engineering approach, obtaining rubusoside and rebaudiosides at titers of around 1.4 g/L and 100 mg/L, respectively.
- Yameng Xu
- , Xinglong Wang
- & Long Liu
-
Article
| Open Access Bioactivity descriptors for uncharacterized chemical compounds
Bioactivity descriptors for uncharacterized chemical compounds
Small molecules bioactivity descriptors are enriched representations of compounds, reaching beyond chemical structures and capturing their known biological properties. Here the authors present a collection of deep neural networks able to infer bioactivity signatures for any compound of interest, even when little or no experimental information is available for them.
- Martino Bertoni
- , Miquel Duran-Frigola
- & Patrick Aloy
-
Article
| Open Access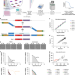 Quantitative and multiplexed chemical-genetic phenotyping in mammalian cells with QMAP-Seq
Quantitative and multiplexed chemical-genetic phenotyping in mammalian cells with QMAP-Seq
Identifying chemical-genetic interactions in mammalian cells is limited to low-throughput or computational methods. Here, the authors present QMAP-Seq, a broadly accessible and scalable approach that uses NGS for pooled high-throughput chemical-genetic profiling in mammalian cells.
- Sonia Brockway
- , Geng Wang
- & Marc L. Mendillo
-
Article
| Open Access DNA-based artificial molecular signaling system that mimics basic elements of reception and response
DNA-based artificial molecular signaling system that mimics basic elements of reception and response
Cells communicate with the outside world to maintain homeostasis. Here the authors design a synthetic biology DNA-based signalling system AMSsys that responds to the presence of ATP.
- Ruizi Peng
- , Liujun Xu
- & Weihong Tan
-
Article
| Open Access Genome-scale reconstructions of the mammalian secretory pathway predict metabolic costs and limitations of protein secretion
Genome-scale reconstructions of the mammalian secretory pathway predict metabolic costs and limitations of protein secretion
The secretory pathway is used in the production of most biopharmaceuticals, but the associated biosynthetic costs are little understood. Here, the authors integrate the core secretory pathway into genome-scale metabolic models of human, mouse, and CHO cells, enabling in silico analysis.
- Jahir M. Gutierrez
- , Amir Feizi
- & Nathan E. Lewis
-
Article
| Open Access Mapping the perturbome network of cellular perturbations
Mapping the perturbome network of cellular perturbations
Our understanding of the mechanisms of drug interactions remains limited. Here the authors introduce a framework to study how complex cellular perturbations induced by different drugs affect each other in morphological feature space.
- Michael Caldera
- , Felix Müller
- & Jörg Menche
-
Article
| Open Access Breast cancer quantitative proteome and proteogenomic landscape
Breast cancer quantitative proteome and proteogenomic landscape
Gene expression profiles can classify breast cancer into five clinically relevant subtypes. Here, the authors perform an in-depth quantitative profiling of the proteome of 45 breast tumors, and show they can recapitulate the transcriptome-based classifications and identify many potentially antigenic tumour-specific peptides.
- Henrik J. Johansson
- , Fabio Socciarelli
- & Janne Lehtiö
-
Article
| Open Access Cycles of external dependency drive evolution of avian carotenoid networks
Cycles of external dependency drive evolution of avian carotenoid networks
The mechanisms that accommodate variable external dependencies in evolution are not clear. Here, the authors show that switches between external and internal metabolic controls of carotenoid-producing networks in birds are linked to shifts in evolutionary rates, with internalization of control resulting in bursts of evolutionary diversification.
- Alexander V. Badyaev
- , Alexander B. Posner
- & Dawn M. Higginson
-
Article
| Open Access Network-based prediction of drug combinations
Network-based prediction of drug combinations
Combination therapy holds great promise, but discovery remains challenging. Here, the authors propose a method to identify efficacious drug combinations for specific diseases, and find that successful combinations tend to target separate neighbourhoods of the disease module in the human interactome.
- Feixiong Cheng
- , István A. Kovács
- & Albert-László Barabási
-
Article
| Open Access Combined chemical genetics and data-driven bioinformatics approach identifies receptor tyrosine kinase inhibitors as host-directed antimicrobials
Combined chemical genetics and data-driven bioinformatics approach identifies receptor tyrosine kinase inhibitors as host-directed antimicrobials
Multidrug resistance necessitates novel approaches to treating bacterial infections. Here, the authors apply their high-throughput screening and in silico prediction approaches to show that host receptor tyrosine kinases are good targets for host-directed therapies against intracellular bacteria.
- Cornelis J. Korbee
- , Matthias T. Heemskerk
- & Tom H. M. Ottenhoff
-
Article |
 Topological data analysis of contagion maps for examining spreading processes on networks
Topological data analysis of contagion maps for examining spreading processes on networks
The spreading dynamics of a contagion depend on the structure of an underlying network, and long-range edges due to airline transportation or media communication can significantly alter such dynamics. Here the authors use contagion dynamics on networks to produce point clouds for this analysis.
- Dane Taylor
- , Florian Klimm
- & Peter J. Mucha
-
Article
| Open Access Navigable networks as Nash equilibria of navigation games
Navigable networks as Nash equilibria of navigation games
Connections in networks are organized to fulfil a function, and a common one is targeted transport or navigation. Here the authors use game theory to show how networks designed to maximize navigation efficiency at minimal cost share basic structural properties, which are also found in real cases.
- András Gulyás
- , József J. Bíró
- & Dmitri Krioukov
-
Article |
 Structural reducibility of multilayer networks
Structural reducibility of multilayer networks
Multilayer networks have been used to capture the structure of complex systems with different types of interactions, but often contain redundant information. Here, De Domenico et al. present a method based on quantum information, to identify the minimal configuration of layers to retain.
- Manlio De Domenico
- , Vincenzo Nicosia
- & Vito Latora
-
Article
| Open Access A scaling law for random walks on networks
A scaling law for random walks on networks
Random walks on a network describe the dynamics of many natural and artificial systems. Here, Perkins et al.study the path distribution—characterizing how the walker moves—and find that it is either finite, stretched exponential or power law for any random walk on a finite network.
- Theodore J. Perkins
- , Eric Foxall
- & Roderick Edwards
-
Article |
 Human symptoms–disease network
Human symptoms–disease network
Unravelling the relationships between disease symptoms and underlying molecular origins is an important task in biomedical research. Here, Zhou et al.link diseases via their symptom overlap, and show that similar phenotypes are mirrored in networks that connect diseases with common genes or protein interactions.
- XueZhong Zhou
- , Jörg Menche
- & Amitabh Sharma
-
Article |
 Epithelial organisation revealed by a network of cellular contacts
Epithelial organisation revealed by a network of cellular contacts
Differences in the arrangement of cells is a fundamental precursor to the establishment of different organs. In this study, network theory is applied at the level of individual cells to map patterns in cell-to-cell contacts, creating a new approach to objectively characterise epithelia.
- Luis M. Escudero
- , Luciano da F. Costa
- & M. Madan Babu

 Improved in situ characterization of protein complex dynamics at scale with thermal proximity co-aggregation
Improved in situ characterization of protein complex dynamics at scale with thermal proximity co-aggregation