Featured
-
-
Article
| Open Access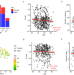 Consistent signatures in the human gut microbiome of old- and young-onset colorectal cancer
Consistent signatures in the human gut microbiome of old- and young-onset colorectal cancer
The rising incidence of young-onset sporadic colorectal cancer (yCRC) is global concern. Here, leveraging a substantial number of deep sequencing metagenomes, the authors show striking similarities in gut microbial patterns at both the taxonomic and selected gene marker levels between yCRC and old-onset CRC.
- Youwen Qin
- , Xin Tong
- & Pei-Rong Ding
-
Article
| Open Access Integrating taxonomic signals from MAGs and contigs improves read annotation and taxonomic profiling of metagenomes
Integrating taxonomic signals from MAGs and contigs improves read annotation and taxonomic profiling of metagenomes
Metagenomic taxonomic profiling usually relies either on reads or assembled contigs/MAGs. Here, authors present RAT, a tool that integrates taxonomic signals from reads, contigs, and MAGs into one profile with high precision and sensitivity. RAT provides a comprehensive view of the microbiome.
- Ernestina Hauptfeld
- , Nikolaos Pappas
- & F. A. Bastiaan von Meijenfeldt
-
Article
| Open Access Survival and rapid resuscitation permit limited productivity in desert microbial communities
Survival and rapid resuscitation permit limited productivity in desert microbial communities
Prompt physiological reactivation after rainfall pulses may be key for microbial survival in arid ecosystems. Here, the authors use stable isotope tracers, single-cell NanoSIMS and metatranscriptomics to shed light on how desert biocrust microbial communities respond to rewetting.
- Stefanie Imminger
- , Dimitri V. Meier
- & Dagmar Woebken
-
Comment
| Open Access All-inclusive nitrifiers in Antarctic soils
All-inclusive nitrifiers in Antarctic soils
Multidisciplinary culture-dependent and -independent techniques elucidate the unique microbial nitrogen cycle in nutrient-poor coastal Antarctica soils and reveal the contribution of novel key microbes to their nitrogen budget.
- Maximiliano Ortiz
-
Article
| Open Access Electrochemically coupled CH4 and CO2 consumption driven by microbial processes
Electrochemically coupled CH4 and CO2 consumption driven by microbial processes
The microbial valorisation of greenhouse gases could offer promising approaches climate change mitigation. Here, authors demonstrate the coupling of methane oxidation and carbon dioxide reduction by microbial consortia, facilitated by the redox cycling of iron minerals.
- Yue Zheng
- , Huan Wang
- & Feng Zhao
-
Article
| Open Access Comparative characterization of the infant gut microbiome and their maternal lineage by a multi-omics approach
Comparative characterization of the infant gut microbiome and their maternal lineage by a multi-omics approach
Here, the authors employ multi-omics on a cohort comprising three generations of family members, showing that fecal microbiota populations, functions, and metabolome of infants vary greatly from their maternal lineage, exhibiting a less diverse microbiota and differences in various metabolite classes including short- and branched-chain fatty acids.
- Tomás Clive Barker-Tejeda
- , Elisa Zubeldia-Varela
- & Alma Villaseñor
-
Article
| Open Access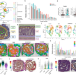 Dermal injury drives a skin to gut axis that disrupts the intestinal microbiome and intestinal immune homeostasis in mice
Dermal injury drives a skin to gut axis that disrupts the intestinal microbiome and intestinal immune homeostasis in mice
The microbial community in the intestine can affect other organs such as the skin but it is not clear if the opposite can occur. Here the authors show that skin wounding affects the microbial composition of the intestinal flora which then enhances DSS induced colitis and intestinal inflammation.
- Tatsuya Dokoshi
- , Yang Chen
- & Richard L. Gallo
-
Article
| Open Access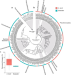 A genome-centric view of the role of the Acropora kenti microbiome in coral health and resilience
A genome-centric view of the role of the Acropora kenti microbiome in coral health and resilience
This study provides insights into the functional roles of microbial symbionts within the reef-building coral Acropora kenti. The findings reveal molecular mechanisms underpinning coral health and adaptation to local environmental stressors, which may support host resilience in the face of anthropogenic climate change and pollution.
- Lauren F. Messer
- , David G. Bourne
- & Gene W. Tyson
-
Article
| Open Access Legume rhizodeposition promotes nitrogen fixation by soil microbiota under crop diversification
Legume rhizodeposition promotes nitrogen fixation by soil microbiota under crop diversification
Sustainability in agriculture can be improved harnessing biological N2 fixation in legumes. Here, the authors combine different crops with peanut plants finding that maize and oilseed rape are the most successful combinations which have potential to enhance rhizosphere microbiota N2 fixation.
- Mengjie Qiao
- , Ruibo Sun
- & Yan Chen
-
Article
| Open Access Genome-scale community modelling reveals conserved metabolic cross-feedings in epipelagic bacterioplankton communities
Genome-scale community modelling reveals conserved metabolic cross-feedings in epipelagic bacterioplankton communities
Identifying the metabolic interactions that underlie microbial communities is challenging. Here, the authors combine Tara Oceans -omics data with co-activity networks and genome-scale metabolic models to predict biotic interactions among planktonic prokaryotes in the upper ocean.
- Nils Giordano
- , Marinna Gaudin
- & Samuel Chaffron
-
Article
| Open Access Profiling the colonic mucosal response to fecal microbiota transplantation identifies a role for GBP5 in colitis in humans and mice
Profiling the colonic mucosal response to fecal microbiota transplantation identifies a role for GBP5 in colitis in humans and mice
Faecal microbiota transplantation (FMT) can be used to treat established colitis. Here the authors profile transcriptional changes in humans after FMT and how this relates to colitis remission identifying a role for GBP5, and this protein is validated in a loss-of-function mouse model.
- Laurence D. W. Luu
- , Abhimanu Pandey
- & Nadeem O. Kaakoush
-
Article
| Open Access Multi-omic integration of microbiome data for identifying disease-associated modules
Multi-omic integration of microbiome data for identifying disease-associated modules
Here, Muller et al. introduce MintTea, a method for analyzing multi-omic microbiome data and identifying disease-associated modules comprising mixed sets of features that collectively shift in disease, offering insights into microbiome-disease interactions.
- Efrat Muller
- , Itamar Shiryan
- & Elhanan Borenstein
-
Article
| Open Access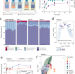 Niche availability and competitive loss by facilitation control proliferation of bacterial strains intended for soil microbiome interventions
Niche availability and competitive loss by facilitation control proliferation of bacterial strains intended for soil microbiome interventions
Bioremediation via microbial inoculation often performs poorly in real-world conditions. Here, the authors show that bacterial inoculants may fail to establish in complex soil microbiomes because they open new niches that facilitate growth of resident microbes.
- Senka Čaušević
- , Manupriyam Dubey
- & Jan Roelof van der Meer
-
Article
| Open Access Targeted metagenomics reveals association between severity and pathogen co-detection in infants with respiratory syncytial virus
Targeted metagenomics reveals association between severity and pathogen co-detection in infants with respiratory syncytial virus
The impact of other pathogens on disease outcome was studied in European infants with RSV infection. Additional viruses were commonly co-detected during infection but were weakly linked to severity. However, presence of Haemophilus bacteria strongly associated with severe cases.
- Gu-Lung Lin
- , Simon B. Drysdale
- & Andrew J. Pollard
-
Article
| Open Access Data-driven prediction of colonization outcomes for complex microbial communities
Data-driven prediction of colonization outcomes for complex microbial communities
Predicting the colonization of exogenous species in complex communities is a challenge in ecology. Here, the authors propose a data-driven approach to predict colonization outcomes and perform validation experiments in human gut microbial communities.
- Lu Wu
- , Xu-Wen Wang
- & Lei Dai
-
Article
| Open Access The defensome of complex bacterial communities
The defensome of complex bacterial communities
Bacteria have evolved numerous innate and adaptive defence mechanisms. Here, Beavogui et al characterise the impact of biogeography, genetic mobility, and clustering in defense islands, on the defence systems of soil, marine, and human gut bacterial populations genomes.
- Angelina Beavogui
- , Auriane Lacroix
- & Pedro H. Oliveira
-
Article
| Open Access A metagenomic catalog of the early-life human gut virome
A metagenomic catalog of the early-life human gut virome
Here, the authors present a metagenomic catalogue of the early-life human gut virome including 160,478 non-redundant DNA and RNA viral sequences from 8,130 gut virus-like particles enriched or bulk metagenomes in the first three years of life.
- Shuqin Zeng
- , Alexandre Almeida
- & Shaopu Wang
-
Article
| Open Access Viral potential to modulate microbial methane metabolism varies by habitat
Viral potential to modulate microbial methane metabolism varies by habitat
The role of viruses in environmental methane cycling is still largely unclear. Here, Zhong et al. analyse metagenomics data to identify auxiliary metabolic genes related to methane metabolism within viral contigs. They found that the potential viral impacts on methane production and oxidation varies by habitat.
- Zhi-Ping Zhong
- , Jingjie Du
- & Matthew B. Sullivan
-
Article
| Open Access Fusobacterium nucleatum promotes tumor progression in KRAS p.G12D-mutant colorectal cancer by binding to DHX15
Fusobacterium nucleatum promotes tumor progression in KRAS p.G12D-mutant colorectal cancer by binding to DHX15
Several studies have shown that Fusobacterium nucleatum aggravates colorectal cancer (CRC) development and chemoresistance. Here the authors show that F. nucleatum is enriched preferentially in patients with KRAS p.G12D mutant CRC and that it promotes colorectal tumorigenesis in preclinical models by binding DHX15 on tumor cells.
- Huiyuan Zhu
- , Man Li
- & Huanlong Qin
-
Article
| Open Access Dynamic root microbiome sustains soybean productivity under unbalanced fertilization
Dynamic root microbiome sustains soybean productivity under unbalanced fertilization
Root-associated microbiomes contribute to plant growth and health. Here, the authors unveil the quantitative development of the root microbiome under unbalanced fertilization and highlight a key microbial cluster for soybean productivity.
- Mingxing Wang
- , An-Hui Ge
- & Ertao Wang
-
Article
| Open Access Enhancing phosphate-solubilising microbial communities through artificial selection
Enhancing phosphate-solubilising microbial communities through artificial selection
Phosphate-solubilising microorganisms can contribute to reduce the use of P fertiliser. Here, the authors use two artificial selection methods, environmental perturbation and propagation, to build phosphate-solubilising communities that retain P-solubilising capacity in hydroponic systems.
- Lena Faller
- , Marcio F. A. Leite
- & Eiko E. Kuramae
-
Article
| Open Access Microbes translocation from oral cavity to nasopharyngeal carcinoma in patients
Microbes translocation from oral cavity to nasopharyngeal carcinoma in patients
Oral microbes in non-oral locations are noted across various cancers. This study highlights the abnormal translocation of oral microbes to the nasopharynx, raising the risk of nasopharyngeal cancer by remodeling the tumor microenvironment and potentially influencing EBV infection.
- Ying Liao
- , Yan-Xia Wu
- & Wei-Hua Jia
-
Article
| Open Access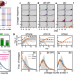 Quantifying the adaptive landscape of commensal gut bacteria using high-resolution lineage tracking
Quantifying the adaptive landscape of commensal gut bacteria using high-resolution lineage tracking
Here, the authors characterize the fine-scale dynamics of genome-wide insertion libraries across four human Bacteroides strains in gnotobiotic mice, revealing rapid adaptation and fitness tradeoffs when commensal gut bacteria adapt to a new host.
- Daniel P. G. H. Wong
- & Benjamin H. Good
-
Article
| Open Access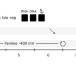 Mechanisms of extracellular electron transfer in anaerobic methanotrophic archaea
Mechanisms of extracellular electron transfer in anaerobic methanotrophic archaea
Anaerobic methanotrophic (ANME) archaea are uncultivated microbes that oxidize the greenhouse gas methane and engage in extracellular electron transfer with other microbes, metal oxides, and electrodes. Here, Ouboter et al. observe strong methane-dependent current associated with high enrichment of ANME archaea on the anode, and provide insights into the mechanisms underlying extracellular electron transfer.
- Heleen T. Ouboter
- , Rob Mesman
- & Cornelia U. Welte
-
Article
| Open Access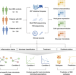 Mucosal host-microbe interactions associate with clinical phenotypes in inflammatory bowel disease
Mucosal host-microbe interactions associate with clinical phenotypes in inflammatory bowel disease
Here, through parallel profiling of the mucosal transcriptome and microbiome of intestinal biopsies derived from patients with IBD and from non-IBD controls, the authors characterize interactions between gene expression and microbiota composition associated with traits of patients with inflammatory bowel diseases. Peer Review Information: Nature Communications thanks Robert Häsler, and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
- Shixian Hu
- , Arno R. Bourgonje
- & Rinse K. Weersma
-
Article
| Open Access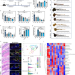 Dubosiella newyorkensis modulates immune tolerance in colitis via the L-lysine-activated AhR-IDO1-Kyn pathway
Dubosiella newyorkensis modulates immune tolerance in colitis via the L-lysine-activated AhR-IDO1-Kyn pathway
Here, Zhang et al. identify a metabolic axis by which Lys-producing commensal bacterium Dubosiella newyorkensis mediates a Treg-mediated immunosuppressive microenvironment by activating AhR-IDO1-Kyn metabolic circuitry in dendritic cells.
- Yanan Zhang
- , Shuyu Tu
- & Shu Jeffrey Zhu
-
Article
| Open Access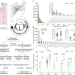 Eco-evolutionary dynamics of gut phageome in wild gibbons (Hoolock tianxing) with seasonal diet variations
Eco-evolutionary dynamics of gut phageome in wild gibbons (Hoolock tianxing) with seasonal diet variations
The significance of gut phageome for wild animals with seasonal diets remains unexplored. Here, the authors use complementary metagenomics to analyze the phage-host dynamics and its implications for diet variations in wild skywalker hoolock gibbons.
- Shao-Ming Gao
- , Han-Lan Fei
- & Peng-Fei Fan
-
Article
| Open Access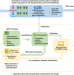 Experimental warming accelerates positive soil priming in a temperate grassland ecosystem
Experimental warming accelerates positive soil priming in a temperate grassland ecosystem
Soil priming could release large amounts of soil C into the atmosphere. Here the authors show that experimental warming boosts soil priming and CO2 emissions in grasslands potentially due to microbial changes. Model accuracy could be improved by incorporating these mechanisms.
- Xuanyu Tao
- , Zhifeng Yang
- & Jizhong Zhou
-
Article
| Open Access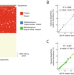 Longitudinal quantification of Bifidobacterium longum subsp. infantis reveals late colonization in the infant gut independent of maternal milk HMO composition
Longitudinal quantification of Bifidobacterium longum subsp. infantis reveals late colonization in the infant gut independent of maternal milk HMO composition
Here, the authors develop a high-throughput method to quantify Bifidobacterium longum subsp. infantis (BL. infantis), a proficient HMO-utilizer, from metagenomic sequencing, and applied it to a longitudinal cohort consisting of 21 mother-infant dyads, suggesting BL. infantis colonization to start late in the breast-feeding period.
- Dena Ennis
- , Shimrit Shmorak
- & Moran Yassour
-
Article
| Open Access A nanoparticle-based sonodynamic therapy reduces Helicobacter pylori infection in mouse without disrupting gut microbiota
A nanoparticle-based sonodynamic therapy reduces Helicobacter pylori infection in mouse without disrupting gut microbiota
Here, the authors develop a non-antibiotic approach using sonodynamic therapy mediated by a lecithin bilayer-coated poly(lactic-co-glycolic) nanoparticle preloaded with verteporfin, Ver-PLGA@Lecithin, to treat Helicobacter pylori.
- Tao Liu
- , Shuang Chai
- & Lihua Yang
-
Article
| Open Access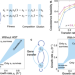 Horizontal gene transfer is predicted to overcome the diversity limit of competing microbial species
Horizontal gene transfer is predicted to overcome the diversity limit of competing microbial species
Combining theoretical derivation and numerical simulations, this study predicts an ecological mechanism for maintaining microbial coexistence via dynamic neutrality enabled by horizontal gene transfer. This mechanism allows the emergence of microbial diversity beyond the limit predicted by previous theories.
- Shiben Zhu
- , Juken Hong
- & Teng Wang
-
Article
| Open Access Predatory protists reduce bacteria wilt disease incidence in tomato plants
Predatory protists reduce bacteria wilt disease incidence in tomato plants
Soil organisms are affected by the presence of predatory protists. Here, the authors predatory protists are negatively associated with bacteria wilt disease incidence in tomato plants and that fertilisation enhances the abundance of predatory protists
- Sai Guo
- , Zixuan Jiao
- & Stefan Geisen
-
Article
| Open Access Compositional and temporal division of labor modulates mixed sugar fermentation by an engineered yeast consortium
Compositional and temporal division of labor modulates mixed sugar fermentation by an engineered yeast consortium
Synthetic microbial communities are suitable for mixed substrates fermentation and long metabolic pathway engineering. Here, the authors combine fermentation experiments with mathematical modeling to reveal the effect of compositional and temporal changes on division of labor in cellulosic ethanol production using two yeast strains.
- Jonghyeok Shin
- , Siqi Liao
- & Yong-Su Jin
-
Article
| Open Access Microbial decomposition of biodegradable plastics on the deep-sea floor
Microbial decomposition of biodegradable plastics on the deep-sea floor
It is unclear whether microbes can efficiently degrade biodegradable plastics in the extreme environmental conditions of the seafloor. Here, Omura et al. show that biodegradable plastics can be degraded by the action of microorganisms on the deep-sea floor, although with much less efficiency than in coastal settings.
- Taku Omura
- , Noriyuki Isobe
- & Tadahisa Iwata
-
Article
| Open Access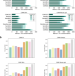 Effective binning of metagenomic contigs using contrastive multi-view representation learning
Effective binning of metagenomic contigs using contrastive multi-view representation learning
Here, the authors present COMEBin, a metagenomics binning method based on contrastive multi-view representation learning that uses data augmentation to generate multiple fragments (views) of each contig, resulting in high-quality embeddings of heterogeneous features. COMEBin outperforms state-of-the art binning methods, particularly in recovering near-complete genomes from real environmental samples.
- Ziye Wang
- , Ronghui You
- & Shanfeng Zhu
-
Article
| Open Access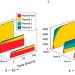 Host- plasmid network structure in wastewater is linked to antimicrobial resistance genes
Host- plasmid network structure in wastewater is linked to antimicrobial resistance genes
Authors apply theory and microbial ecology modelling to a wastewater sample, and show that antimicrobial resistance carrying plasmids interact with a higher number and more diverse range of bacteria than plasmids that do not carry resistance genes.
- Alice Risely
- , Arthur Newbury
- & Dirk Sanders
-
Article
| Open Access Towards estimating the number of strains that make up a natural bacterial population
Towards estimating the number of strains that make up a natural bacterial population
What a microbial strain is and how many strains make up a natural bacterial population remain elusive concepts. Here, Viver et al. analyse Salinibacter ruber isolates and metagenomes from two solar salterns, revealing gaps within the species sequence space that they use to define and quantify sub-species categories, such as genomovars and strains, that co-exist in a saltern pond.
- Tomeu Viver
- , Roth E. Conrad
- & Ramon Rossello-Mora
-
Article
| Open Access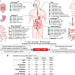 Defining the biogeographical map and potential bacterial translocation of microbiome in human ‘surface organs’
Defining the biogeographical map and potential bacterial translocation of microbiome in human ‘surface organs’
Given that the human body is composed of many microbial niches, and there have been few reports on the biogeography of the microbiome, the authors analyse the intra-individual inter-organ and intra-organ microbiome of seven surface organs of deceased individuals.
- Jun-Jun She
- , Wei-Xin Liu
- & Jun Yu
-
Article
| Open Access Functional host-specific adaptation of the intestinal microbiome in hominids
Functional host-specific adaptation of the intestinal microbiome in hominids
Here, Rühlemann et al. analyze the gut microbiome of wild-living African great apes (Gorillas, Bonobos, Chimpanzees) in comparison to that of humans, identifying host specific patterns and shared evolutionary conserved traits disrupted in humans.
- M. C. Rühlemann
- , C. Bang
- & A. Franke
-
Article
| Open Access Spatial structure, chemotaxis and quorum sensing shape bacterial biomass accumulation in complex porous media
Spatial structure, chemotaxis and quorum sensing shape bacterial biomass accumulation in complex porous media
Pores and channels within complex porous structures, such as the soil or the human gut, influence fluid flow and thus bacterial colonization. Here, Scheidweiler et al. study bacterial colonization of a model complex porous structure and show how the interactions between fluid flow, microscale structure, chemotaxis, and gradients of a quorum-sensing signaling molecule control the heterogenous accumulation of bacterial biomass.
- David Scheidweiler
- , Ankur Deep Bordoloi
- & Pietro de Anna
-
Article
| Open Access Gut microbiota facilitate chronic spontaneous urticaria
Gut microbiota facilitate chronic spontaneous urticaria
Chronic spontaneous urticarial is an inflammatory skin disease which has been linked to intestinal dysbiosis. Here the authors implicate intestinal dysbiosis with the inflammatory response in a murine model of urticaria.
- Lei Zhu
- , Xingxing Jian
- & Jie Li
-
Article
| Open Access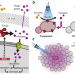 Optogenetic spatial patterning of cooperation in yeast populations
Optogenetic spatial patterning of cooperation in yeast populations
Microbial communities are the siege of complex metabolic interactions including cooperation and competition. Here, the authors report the utilization of optogenetics and spatial light-patterning to activate the expression of the invertase SUC2 at selected locations and selectively switch cooperation and competition roles of the yeast cells.
- Matthias Le Bec
- , Sylvain Pouzet
- & Pascal Hersen
-
Article
| Open Access The antibiotic resistance reservoir of the lung microbiome expands with age in a population of critically ill patients
The antibiotic resistance reservoir of the lung microbiome expands with age in a population of critically ill patients
Here, by performing tracheal aspirate RNA sequencing of critically ill patients, the authors find that older age associates with a greater number of detectably expressed antimicrobial resistance genes in the lower respiratory tract microbiome.
- Victoria T. Chu
- , Alexandra Tsitsiklis
- & Charles R. Langelier
-
Article
| Open Access Microbiome homeostasis on rice leaves is regulated by a precursor molecule of lignin biosynthesis
Microbiome homeostasis on rice leaves is regulated by a precursor molecule of lignin biosynthesis
The underlying mechanisms of host-driven assembly of phyllosphere microbiota remain largely unknown. Here, 4-hydroxycinnamic acid synthesized by the rice plant’s PAL02 in the phenylpropanoid biosynthesis pathway is shown to be the main driver for enrichment of Pseudomonadales bacteria.
- Pin Su
- , Houxiang Kang
- & Yong Liu
-
Article
| Open Access Microbial co-occurrences on catheters from long-term catheterized patients
Microbial co-occurrences on catheters from long-term catheterized patients
The authors examine temporal polymicrobial community composition in patients with long-term urinary catheters to identify species co-occurrences and demonstrate uropathogenic Escherichia coli augments growth of a prevalent opportunistic uropathogen in urine.
- Taylor M. Nye
- , Zongsen Zou
- & Scott J. Hultgren
-
Article
| Open Access Differential responses of the gut microbiome and resistome to antibiotic exposures in infants and adults
Differential responses of the gut microbiome and resistome to antibiotic exposures in infants and adults
Knowledge on how the gut microbiome and resistome responds to antibiotics across age remains limited. Here, using metagenomics data from Danish infants and young adults, the authors show that antibiotics have a more lasting impact on adults compared to infants.
- Xuanji Li
- , Asker Brejnrod
- & Søren Johannes Sørensen
-
Article
| Open Access Microbial interactions shape cheese flavour formation
Microbial interactions shape cheese flavour formation
Cheese fermentation and flavour formation are the result of complex biochemical reactions driven by the activity of multiple microorganisms. Here, the authors identify microbial interactions as a mechanism underlying flavour formation in Cheddar cheese.
- Chrats Melkonian
- , Francisco Zorrilla
- & Ahmad A. Zeidan
-
Article
| Open Access Identification of inulin-responsive bacteria in the gut microbiota via multi-modal activity-based sorting
Identification of inulin-responsive bacteria in the gut microbiota via multi-modal activity-based sorting
Here, Riva et al. employ a multi-modal approach to identify gut microbes stimulated by the popular dietary supplement inulin and reveal that inulin binding and metabolic stimulation are widespread in the microbiome, making the framework a suitable way to study key microbes that perform specific functions in the microbiome.
- Alessandra Riva
- , Hamid Rasoulimehrabani
- & David Berry
-
Article
| Open Access Genome-resolved metatranscriptomics reveals conserved root colonization determinants in a synthetic microbiota
Genome-resolved metatranscriptomics reveals conserved root colonization determinants in a synthetic microbiota
The identification of processes activated by specific microbes during microbiota colonization of plant roots is hampered by technical issues in metatranscriptomics. Here, Vannier et al. colonized germ-free plants with a defined root microbiota comprising over 100 microbial isolates, and addressed those issues in various ways to identify strain-specific processes as well as common gene sets activated by microbes during root colonization.
- Nathan Vannier
- , Fantin Mesny
- & Stéphane Hacquard

 LSD1 drives intestinal epithelial maturation and controls small intestinal immune cell composition independent of microbiota in a murine model
LSD1 drives intestinal epithelial maturation and controls small intestinal immune cell composition independent of microbiota in a murine model