Abstract
RF pulse schemes for the simultaneous acquisition of heteronuclear multi-dimensional chemical shift correlation spectra, such as {HA(CA)NH & HA(CACO)NH}, {HA(CA)NH & H(N)CAHA} and {H(N)CAHA & H(CC)NH}, that are commonly employed in the study of moderately-sized protein molecules, have been implemented using dual sequential 1H acquisitions in the direct dimension. Such an approach is not only beneficial in terms of the reduction of experimental time as compared to data collection via two separate experiments but also facilitates the unambiguous sequential linking of the backbone amino acid residues. The potential of sequential 1H data acquisition procedure in the study of RNA is also demonstrated here.
Similar content being viewed by others
Introduction
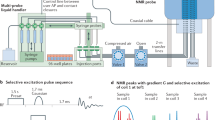
NMR spectroscopy is a powerful technique for the study of biomolecular structure and dynamics, both in solution as well as in the solid state. In protein NMR a variety of multi-dimensional heteronuclear chemical shift correlation experiments are typically used for resonance assignment and for extraction of 1H-1H distance restraints, e.g. HACANH1,2, HNCAHA3,4,5, HACACONH2, HNCOCAHA4,5, HCCNH1,6 and 15N-edited 1H-1H NOESY, respectively. These multi-dimensional spectra are based on different magnetisation transfer pathways and are customarily collected individually. As a result, the time for the acquisition of all required data sets can in many cases become exceedingly long. In this context, a variety of techniques are currently being explored for reducing data acquisition times7,8,9,10. One of the approaches that has received considerable attention in the study of proteins, both in solution11,12,13,14 and in the solid state15,16,17, involves the simultaneous collection of different chemical shift correlation spectra. For example, using dual receivers with 1H and13C acquisition in the direct dimension, the simultaneous collection of chemical shift correlation spectra, e.g. {HACANH & HACACO} and {HACACO & HACACONH}, has been demonstrated employing both parallel12 and sequential13 data acquisition procedures. However, as noted already in the literature11,12, all other aspects being equal, e.g. magnetisation transfer and relaxation characteristics, the signal intensities seen in the 1H and 13C detected data sets would differ because of the difference in the gyromagnetic ratios of the two nuclei. Furthermore, the utility of such experiments is limited as the solution state NMR probes are typically optimised either for 1H or for 13C detection only. In addition, many of the chemical shift correlation experiments of interest do not require dual receivers and instead may involve 1H or 13C acquisition only. In this context, RF pulse schemes enabling the collection of multi-dimensional data sets with a single receiver is of great interest18,19,20,21. Recently we have reported RF pulse schemes involving dual sequential 1H acquisition with only amide proton detection and making use of dual 15N-13C mixing steps for achieving protein resonance assignment22. While such sequences can be easily adapted to the study of large 2H-labeled protein samples, here we present RF pulse schemes that were developed in the context of moderately sized proteins. In such systems the relaxation losses during 15N-13C mixing periods are not expected to be significant even in a fully protonated sample. The pulse sequences presented here make use of the availability of both the HA and HN protons and employ only a single15N-13C mixing step to achieve sequential resonance assignments in protonated protein samples. The efficacy of the approach is experimentally demonstrated by the ‘one-shot’ collection of representative protein NMR spectra. We also show that this approach is useful for the study of RNA.
Results and discussion
{HA(CA)NH & HA(CACO)NH}
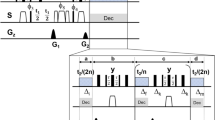
The triple resonance HACANH experiment is frequently used for sequential resonance assignment of the backbone 13Cα,15N,1HN and 1Hα nuclei and involves through-bond magnetisation transfer between the directly coupled nuclei via the pathway 1Hα->13Cα->15N->1HN. Interresidue cross peaks arising from transfer of magnetisation from the 13Cα spin of residue (i) to the 15N spin of residue of (i + 1), resulting from 2JCaN couplings, are observed in this experiment. Interresidue cross peaks can usually be distinguished from intraresidue cross peaks based on their respective signal intensity. However, to achieve unambiguous resonance assignment, the HACACONH experiment involving magnetisation transfers via the pathway 1Hα->13Cα->13CO->15N->1HN and leading only to interresidue cross peaks is generally carried out in addition. In the HACANH experiment, however, the 13Cα->15N magnetisation transfer is the critical step as it relies on weak intraresidue 1JCaN couplings (~11 Hz). Although heteronuclear magnetisation transfers are typically carried out via INEPT type transfers, in-phase magnetisation transfers via heteronuclear cross polarization schemes have also been successfully used to enhance sensitivity in triple resonance NMR experiments such as in HACANH1. Here, we have implemented RF pulse schemes for the sequential collection of different correlation spectra using 15N<->13C cross polarization schemes. The RF pulse scheme given in Fig. 1a permits the ‘one-shot’ acquisition of 3D HA(CA)NH and 3D HA(CACO)NH data sets. The initial transverse 1H magnetisation generated by the first 90° pulse is allowed to evolve under its chemical shift during the t1(HA)/t1′(HA) period and under the one bond heteronuclear 13C-1H coupling for a period of 2Δ0 to generate antiphase 1H magnetisation. The anti-phase 1H magnetisation is then converted into antiphase carbon magnetisation by the 90° pulses applied to the two nuclei. The antiphase 13Cα polarisation is allowed to refocus during the interval 2Δ1 to generate (13Cα)x magnetisation and then subjected to a period of 13Cα->15N magnetisation exchange via the application of a band-selective het-TOCSY mixing sequence. The residual13C transverse magnetisation remaining after the 13Cα->15N transfer step is flipped to the z axis and the 15N magnetisation generated after 13Cα->15N mixing is allowed to evolve under its chemical shift during the t2(N) period and transferred to the attached proton via an INEPT step and is observed in the t3 period under 15N decoupling to generate the 3D HA(CA)NH spectrum. The WATERGATE sequence29 is used for water suppression. After the completion the first 1H acquisition, the residual 13Cα magnetisation is brought to the transverse plane and subjected to 13Cα->13CO cross polarisation. This transverse 13CO magnetisation is then subjected to a period of 13CO->15N magnetisation exchange via the application of a band-selective het-TOCSY mixing sequence. The resulting transverse 15N magnetisation is allowed to evolve during the t2′ (N) period and then transferred to the attached proton via the INEPT procedure. The 1H signals are acquired in t3′, under 15N decoupling to generate the 3D-HA(CACO)NH data. The cross peak intensities observed in the HA(CACO)NH spectrum is dependent on the amount of residual 13Cα magnetisation present after the 13Cα->15N mixing period and hence related to the duration of the mixing period and the performance characteristics of the mixing sequence employed. The residual 13Cα magnetisation has to be kept along the z axis until the first data acquisition is completed and this may affect the signal intensities observed in the HA(CACO)NH spectrum due to relaxation losses. However, in the systems studied here, significant variation in signal intensities were not observed when the residence time of the 13Cα magnetisation along the z axis was varied over a range of 0–100 ms (Fig. S1). The optimal length of the 15N-13C het-TOCSY mixing period was found to be ~50 ms (Fig. S2). With this approach we have successfully acquired {3D HA(CA)NH & 3D HA(CACO)NH} spectra of the MCM C-terminal winged helix domain (Fig. 2). Representative spectral cross sections taken from these 3D data sets are given in the supplementary material (Fig. S3, S4) to indicate spectral quality.
RF pulse schemes for the simultaneous acquisition of (a) 3D {HA(CA)NH & HA(CACO)NH} (b) 3D {H(N)CAHA & HA(CA)NH} and (c) 3D {H(N)CAHA & H(CC)NH} chemical shift correlation spectra of proteins with dual sequential 1H acquisitions in the direct dimension.
Open and filled rectangles represent 180° and 90° pulses, respectively. Phase cycling is as follows: (a) φ1 = x, − x; φ2 = 8(x), 8(−x); φ3 = 2(x), 2(−x); φ4 = 4(x), 4(−x); φ5 = 2(y), 2(−y); φ6 = 4(x), 4(−x); φR1 = φR2 = x, 2(−x), x, −x, 2(x), 2(−x), 2(x), −x, x, 2(−x), x. Gradients with a sine bell amplitude profile were used. Durations and strength with respect to the maximum strength of 50 G/cm are: G1 = 1 ms (60%), G2 = 1 ms (80%). (b) φ1 = x, − x; φ2 = 4(x), 4(−x); φ3 = φ4 = 2(y), 2(−y); φ5 = 8(y), 8(−y); φ6 = 4(y), 4(−y); φR1 = x, 2(−x), x, − x, 2(x), 2(−x), 2(x), −x, x, 2(−x), x; φR2 = x, 2(−x), x, −x, 2(x), −x; G1 = 1 ms (60%), G2,3 = 5 ms (60%), G4 = 4.4 ms (60%), G5 = 1 ms (80%). (c) φ1 = x, − x; φ2 = 4(x), 4(−x); φ3 = 16(y), 16(−y); φ4 = 2(y), 2(−y); φ5 = 8(y), 8(−y); φ6 = 4(y), 4(−y); φ7 = 2(x), 2(−x); φR1 = x, 2(−x), x, −x, 2(x), 2(−x), 2(x), −x, x, 2(−x), x; φR2 = x, 2(−x), x, −x, 2(x), −x, x, 2(−x), x, −x, 2(x), −x, −x, 2(x), −x, x, 2(−x), x, −x, 2(x), −x, x, 2(−x), x; G1 = 1 ms (60%), G2,3 = 5 ms (60%), G4 = 4.4 ms (60%), G5 = 1 ms (80%).
Simultaneously acquired (a) 3D HA(CA)NH and (b) 3D HA(CACO)NH spectra of the MCM C-terminal winged helix domain of Sulfolobus solfataricus recorded at 600 MHz with 16 transients per t1 increment, 41 t1 increments, 45 t2 increments, spectral widths of 1559 Hz (1Ha) and 1945 Hz (15N) in the indirect dimensions, respectively, a recycle time of 1.0 s and a proton acquisition time of 60 ms in the direct dimension.
The total experimental time was 44 h. The AK2-JCHaniso1 and AK2-JCaC'aniso sequences were used for 15N-13C and 13Ca-13CO anisotropic cross polarisation, respectively. 13Ca-13CO mixing was carried out keeping the 13C RF carrier at 115 ppm, with a peak RF power level of ~11 kHz and for a total duration of 17.92 ms by repeating the basic sequence twice (8.96 ms * 2). 15N-13C mixing was carried out by keeping the 13C RF carrier either at 55 ppm or at 175 ppm for achieving band-selective 15N-13CA and 15N-13CO cross polarisations for durations of 25 ms and 50 ms (25 ms * 2), respectively. The 15N-13C mixing sequence with the basic cycle duration of 25 ms was applied employing 15N/13C peak RF power level of ~3.6 kHz, keeping the 15N RF carrier at 121 ppm. The 1H RF carrier was kept at 4.3 ppm during t1 and subsequently switched back to the water position (4.7 ppm). Δ0,1,2 = 1.56, 1.56, 2.38 ms were used for INEPT transfers. (c)1Hα-1HN spectral cross-sections from the HA(CA)NH (blue) and HA(CACO)NH (red) spectra taken at the 15N chemical shifts positions indicated and showing the sequential walk along the backbone residues spanning the region E641-K650.
{HA(CA)NH & H(N)CAHA}
The RF pulse scheme given in Fig. 1b allows to simultaneously collect data from both the HA(CA)NH and H(N)CAHA experiments. Here, unlike the case in the RF pulse schemes given in Fig. 1a, the initial transverse magnetisation generated from both15N and 13C attached protons by the first 90° pulse is allowed to undergo chemical shift evolution during the t1(HN)/t1′(HA) period. These evolve under the one bond heteronuclear 15N-1H and 13C-1H scalar couplings during the periods 2Δ0 and (Δ0 + Δ1 − Δ2), respectively (taking into account the different one bond heteronuclear scalar couplings), to generate the relevant antiphase 1H magnetisation. The antiphase 1H magnetisation are then converted into the corresponding antiphase nitrogen and carbon magnetisation by the 90° pulses applied to the different nuclei. The antiphase 15N and 13C polarisation is then allowed to refocus during the interval 2τ1 and 2τ2 to generate (15N/13C)x magnetisation and then subjected to 15N<->13Cα magnetisation exchange via the application of a band-selective het-TOCSY mixing sequence. Both, the 15N and 13C transverse magnetisation present after the 15N<->13Cα transfer step is flipped to the z axis. First, the data from the 1HN->15N->13Cα->1Hα pathway [H(N)CAHA] is collected, followed by the acquisition of the signals from the 1Hα->13Cα->15N->1HN pathway. Sufficient solvent suppression was accomplished by 1H x- and y-purge pulses in combination with gradient pulses30 just before the 13C-1H cross polarisation step and with the 13Cα magnetisation along the z axis. In both data sets, intra- and inter-residue peaks arising, respectively, due to 1JCaN and 2JCaN couplings are observed.


With a single15N-13C mixing step in the RF pulse sequence (Fig. 1b), the simultaneous collection of H(N)CAHA and HA(CA)NH spectra delivers the chemical shifts of the backbone 13Cα;i,15Ni,1HN;i and 1Hα;i nuclei. Additionally, the (15N,1H) backbone chemical shifts of the adjacent i+1 residue and the (13CA,1HA) chemical shifts of the preceding i−1 residue are also obtained. This facilitates the unambiguous linking of three amino acid residues, i.e. i-1, i and i+1. With this approach we have successfully acquired a combined data set comprising the HA(CA)NH and H(N)CAHA experiment (Fig. 3). Data collected with a cryoprobe are provided in the supplementary material (Fig. S5, S6) to illustrate the performance of the sequence at lower protein concentrations.
Simultaneously acquired 1H-1H correlation spectra via the (a) 3D H(N)CAHA and (b) 3D HA(CA)NH experiments.
These (ω1–ω3) spectra of the MCM C-terminal winged helix domain of Sulfolobus solfataricus recorded at 600 MHz with 16 transients per t1 increment, 105 t1 increments, spectral widths in the indirect dimensions of 3598 Hz (1H), a recycle time of 1.0 s and a proton acquisition time of 60 ms in the direct dimension. Total experimental time was ~1 h. The AK2-JCHaniso1 sequence was used for both 15N->13CA and 13C->1H anisotropic cross polarisation transfers. The 15N-13CA mixing was carried out by keeping the 13C RF carrier at 55 ppm, employing 15N/13C peak RF power level of ~3.6 kHz and for a duration of 50 ms by repeating the basic sequence twice (25 ms * 2). The 13C->1H het-TOCSY was carried out with one cycle of the AK2-JCHaniso1 sequence having a duration of 7.2 ms, employing 1H/13CA peak RF power level of ~12.5 kHz. The 1H RF carrier was kept at 4.7 ppm. The 1H RF carrier was kept at 3 ppm during t1 and subsequently switched back to the water position (4.7 ppm). Δ0,1,2 = 2.58, 1.79, 0.79 ms, 2τ1 = 2Δ0 and 2τ2 = 3 ms.
{H(N)CAHA & H(CC)NH}
In addition to resonance assignment of backbone nuclei a modification of the RF pulse scheme given in Fig. 1b allows to simultaneously acquire the 3D H(CC)NH and H(N)CAHA correlation spectra and to obtain chemical shift information on the protein side chain as well as on the backbone nuclei. In a simple HACANH experiment the 13Cα magnetisation used for 13Cα->15N mixing arises only from the magnetisation transfer from directly attached 1Hα protons. However, the 13Cα magnetisation in the HCCNH experiment is also generated starting from the side chain protons via the 1Hsc->13Csc->13Cα magnetisation transfer pathway. This is achieved by introducing a 13C-13C longitudinal TOCSY mixing period just before the heteronuclear cross polarisation step (Fig. 1c), with the remainder of the pulse sequence essentially the same as in Fig. 1b. Obviously, one can design the RF pulse scheme to obtain either 1H or 13C side chain chemical shift information. The spectral widths in the indirect dimension in the two data sets can also be independently adjusted by appropriate scaling of the t2 (CA)/t2′ (N) increments and spectral folding in one data set does not lead to resonance overlaps in the other, as the two data sets are effectively independent. As in the case of HACANH experiment, interresidue side chain cross peaks arising from transfer of magnetisation from 13Cα spin of residue (i) to the 15N spin of residue of (i + 1) are observed in the HCCNH spectrum. The HCCNH and HNCAHA spectra (Fig. 4) were acquired in one shot via the pulse scheme given in Fig. 1c.
Simultaneously acquired 3D correlation spectra via the (a) 3D H(N)CAHA and (b) 3D H(CC)NH experiments.
These spectra of the MCM C-terminal winged helix domain of Sulfolobus solfataricus recorded at 600 MHz with 16 transients per t1 increment, 36 t1 increments, 50 t2 increments, spectral widths in the indirect dimensions of 3598 Hz (1H), 5278 Hz (13C), 2639 Hz (15N), a recycle time of 1.0 s and a proton acquisition time of 60 ms in the direct dimension. Total experimental time was 42 h. The AK2-JCHaniso1 sequence was used for both15N->13CA and 13C->1H anisotropic cross polarisation transfers. The 15N-13CA mixing was carried out by keeping the 13C RF carrier at 55 ppm, employing15N/13C peak RF power level of ~3.6 kHz and for a duration of 50 ms by repeating the basic sequence twice (25 ms * 2). The 13C->1H het-TOCSY was carried out with one cycle of the AK2-JCHaniso1 sequence having a duration of 7.2 ms, employing 1H/13CA peak RF power level of ~12.5 kHz. Longitudinal 13C-13C mixing in the aliphatic region was carried out employing the AK2-JCC sequence, with a peak 13C RF power level of 10 kHz and for a duration of 9.6 ms by repeating two times the basic cycle of duration 4.8 ms (4.8 ms *2). The RF carrier was kept at 35 ppm during 13C-13C mixing and at 55 ppm for 13CA-15N band-selective mixing. The 1H RF carrier was kept at 3 ppm during t1 and subsequently switched back to the water position at 4.7 ppm. Δ0,1,2 = 2.58, 1.79, 0.79 ms, 2τ1 = 2Δ0 and 2τ2 = 3 ms were used for INEPT transfers. (c)1HN-1Hα spectral cross-sections from the H(N)CAHA spectrum taken at the 13C chemical shifts positions indicated and showing the sequential walk along the backbone residues spanning the region E641-K650 (d)1Hα-1HN spectral cross-sections from the H(CC)NH spectrum taken at the15N chemical shifts positions indicated and showing the connectivities between the adjacent backbone residues.
The results presented here clearly demonstrate that it is possible to achieve simultaneous acquisition of multidimensional data sets in solution using the sequential data acquisition procedure, akin to recently reported solid state NMR studies of proteins15,16,17. Additionally, the unambiguous sequential linking of backbone nuclei (i − 1, i, i + 1) is achieved in one shot. The basic strategy with sequential data acquisition procedure is that two different experiments leading to correlation spectra arising from different magnetisation transfer pathways are simultaneously started and at a defined intermediate stage the relevant magnetisation belonging to one of the pathways is kept along the z axis. Depending on the type of data to be sequentially collected, it can be either 15N or the 13C nuclei that are to be kept as longitudinal polarisation. After this, magnetisation transfers followed by the first data acquisition are carried out to complete the experiment via the first pathway. Subsequently, appropriate magnetisation transfers allow for the second data acquisition to complete the correlation experiment via the second pathway. Central to this approach is, that the magnetisation which is stored along the z axis for usage in the second experiment should not be disturbed during the completion of the first experiment. The order in which both multi-dimensional data sets are sequentially acquired has to be chosen appropriately in this context. For medium sized molecules, which are the focus of the present study and with short acquisition times on the order of ~50 ms in the direct dimension, sequential acquisition of correlation spectra do not suffer from significant relaxation losses irrespective of whether 15N or 13C nuclei are kept along the z axis. Although the acquisitions of only a few representative spectra are demonstrated here, the approach outlined can be extended to acquire simultaneously other types of correlation spectra. For example, utilizing the HNCAHA experiment for the sequential assignment of the backbone 13Cα,15N,1HN and 1Hα nuclei allow to exploit the residual 15N magnetisation after the 15N->13Cα transfer for generating simultaneously a 15N edited 1H-1H NOESY spectrum (Fig. S7, S8, S9). As the application of het-TOCSY mixing schemes over long periods of time may lead to sample heating and hence might pose problems in the study of temperature sensitive samples, one may take recourse to the INEPT procedure to effect 15N-13C magnetisation transfers. Although the relative merits in the context of sequential data acquisitions are yet to be fully assessed, good quality spectra are obtained by implementing INEPT type transfers31 for 15N<->13C mixing (Fig. S10).
The sequential data acquisition procedure can also be effectively used for simultaneously generating heteronuclear correlation spectra of RNA32,33. For example, triple resonance NMR experiments such as HCNCH/HCNH are often used for achieving intra- nucleotide correlation of the sugar protons with the base protons. The HCNCH experiment involves through-bond magnetisation transfers between the directly coupled nuclei via the pathway 1H1′->13C1′->15N1,9->13C6,8->1H6,8 and makes use of 1JCN couplings (~12 Hz). The HCNH experiment involves the magnetisation transfer pathway 1H1′->3C1′->15N1,9->1H6,8 and makes use of 1JCN and 2JNH couplings. In both experiments, that are typically carried out in D2O, the residual13C1′ magnetisation after the 13C1′->15N1,9 transfer step can efficiently be exploited to obtain simultaneously COSY/TOCSY data for sugar protons (Fig. S11, S12). For RNA samples dissolved in H2O, the through-bond HNCCH experiment is often used for correlating the H3 imino protons of uridines with the H5/H6 base protons via the pathway 1Him->15Nim->13C4->13C5,6->1H5,6. Employing the residual magnetisation after the 15Nim->13C4 transfer step, one can simultaneously obtain the NOE correlation spectrum of the imino protons in RNA (Fig. S11, S13). The sequential data acquisition strategy presented here may also be combined with other approaches for further reducing the data acquisition time, e.g. sparse sampling in the indirect dimension34. Furthermore, such dual sequential acquisition procedure may also be applied to collect 3D data with direct 13C detection.
Methods
Uniformly (13C,15N)-labelled samples of the 82 amino acid MCM C-terminal winged helix domain of Sulfolobus solfataricus and the 94 amino acid N-terminal region of human hnRNP C proteins were expressed and purified as reported earlier23,24. For development, a room temperature triple resonance probe was used and the final protein concentrations were 7 mM and 3 mM, respectively. In addition studies were also undertaken with a cryoprobe utilizing protein samples at lower concentrations (0.8 mM and 1.2 mM, supplementary material). The uniformly (13C,15N)-labelled RNA (5′-GGCGUUCGCUUAGAACGUC-3′), referred to as BEVSLD5, was prepared as described25 and a final concentration of 0.9 mM was used here. Multi-dimensional chemical shift correlation experiments for proteins were carried out with a Bruker 600 MHz narrow-bore Avance III NMR spectrometer equipped with pulse field gradient accessories, pulse shaping units and a triple resonance probe. Sample temperature was set to 303 K. For the BEVSLD5 RNA a triple resonance cryoprobe was used; sample temperature was set to 293 K for experiments on non-exchangeable and 288 K for experiments on exchangeable protons, respectively. Homo- and heteronuclear magnetisation transfers were achieved using amplitude and phase-modulated mixing sequences (AK2-JCHanisol, AK2-JCαC'aniso:26; AK2-JCC27). Where required, RF field strength and duration of the mixing period were scaled appropriately. The States procedure28 was applied for phase-sensitive detection in the indirect dimensions. Standard phase cycling procedures were employed to select signals arising from desired magnetisation transfer pathways.
References
Richardson, J. M. et al. The Use of Heteronuclear Cross Polarization to Enhance the Sensitivity of Triple-Resonance NMR Experiments. Improved 4D HCNNH Pulse Sequences. J. Magn. Reson. 101, 223–227 (1993).
Boucher, W., Laue, E. D., Campbell-Burk, S. & Domaille, P. J. Four-dimensional heteronuclear triple resonance NMR methods for the assignment of backbone nuclei in proteins. J. Am. Chem. Soc. 114, 2262–2264 (1992).
Clubb, R. T., Thanabal, V. & Wagner, G. A new 3D HN (CA) HA experiment for obtaining fingerprint HN-Hα cross peaks in15N-and13C-labeled proteins. J. Biomol. NMR 2, 203–210 (1992).
Olejniczak, E. T., Xu, R. X., Petros, A. M. & Fesik, S. W. Optimized constant-time 4D HNCAHA and HN(CO)CAHA experiments. Applications to the backbone assignments of the FKBP/ascomycin complex. J. Magn. Reson. 100, 444–450 (1992).
Kay, L. E., Wittekind, M., McCoy, M. A., Friedrichs, M. S. & Mueller, L. 4D NMR triple-resonance experiments for assignment of protein backbone nuclei using shared constant-time evolution periods. J. Magn. Reson. 98, 443–450 (1992).
Lyons, B. A. & Montelione, G. T. An HCCNH triple-resonance experiment using carbon-13 isotropic mixing for correlating backbone amide and side-chain aliphatic resonances in isotopically enriched proteins. J. Magn. Reson. 101, 206–209 (1993).
Hyberts, S. G., Arthanari, H. & Wagner, G. Applications of Non-Uniform Sampling and Processing. Top. Curr. Chem. 316, 125–148 (2011).
Hiller, S. & Wider, G. Automated Projection Spectroscopy and Its Applications. Top. Curr. Chem. 316, 21–47 (2011).
Maciejewski, M. W., Mobli, M., Schuyler, A. D., Stern, A. S. & Hoch, J. C. Data Sampling in Multidimensional NMR: Fundamentals and Strategies. Top. Curr. Chem. 316, 49–77 (2011).
Schanda, P. Fast-pulsing longitudinal relaxation optimized techniques: Enriching the toolbox of fast biomolecular NMR spectroscopy. Prog. Nucl. Magn. Reson. Spectrosc. 55, 238–265 (2009).
Kupče, E. E. & Kay, L. E. L. Parallel acquisition of multi-dimensional spectra in protein NMR. J. Biomol. NMR 54, 1–7 (2012).
Reddy, J. G. & Hosur, R. V. Parallel acquisition of 3D-HA(CA)NH and 3D-HACACO spectra. J. Biomol. NMR 56, 77–84 (2013).
Kupče, E., Kay, L. E. & Freeman, R. Detecting the “Afterglow” of 13C NMR in Proteins Using Multiple Receivers. J. Am. Chem. Soc. 132, 18008–18011 (2010).
Chakraborty, S., Paul, S. & Hosur, R. V. Simultaneous acquisition of 13Cα-15N and 1H-15N-15N sequential correlations in proteins: application of dual receivers in 3D HNN. J. Biomol. NMR 52, 5–10 (2011).
Banigan, J. R. & Traaseth, N. J. Utilizing afterglow magnetization from cross-polarization magic-angle-spinning solid-state NMR spectroscopy to obtain simultaneous heteronuclear multidimensional spectra. J. Phys. Chem. B 116, 7138–7144 (2012).
Gopinath, T. & Veglia, G. Dual acquisition magic-angle spinning solid-state NMR-spectroscopy: simultaneous acquisition of multidimensional spectra of biomacromolecules. Angew. Chem., Int. Ed. Engl. 51, 2731–2735 (2012).
Bellstedt, P. et al. Solid state NMR of proteins at high MAS frequencies: symmetry-based mixing and simultaneous acquisition of chemical shift correlation spectra. J. Biomol. NMR 54, 325–335 (2012).
Farmer, B. T. & Mueller, L. Simultaneous acquisition of [13C,15N]- and [15N,15N]-separated 4D gradient-enhanced NOESY spectra in proteins. J. Biomol. NMR 4, 673–687 (1994).
Pascal, S. M., Muhandiram, D. R., Yamazaki, T., Formankay, J. D. & Kay, L. E. Simultaneous Acquisition of 15 N-and 13 C-Edited NOE Spectra of Proteins Dissolved in H2O. J. Magn. Reson. 103, 197–201 (1994).
Ramachandran, R. R., Sich, C. C., Grüne, M. M., Soskic, V. V. & Brown, L. R. L. Sequential assignments in uniformly 13C- and 15N-labelled RNAs: the HC(N,P) and HC(N,P)-CCH-TOCSY experiments. J. Biomol. NMR 7, 251–255 (1996).
Haasnoot, C. A. G., van de Ven, F. J. M. & Hilbers, C. W. COCONOSY: combination of 2D correlated and 2D nuclear overhauser enhancement spectroscopy in a single experiment. J. Magn. Reson. 56, 343–349 (1984).
Wiedemann, C. et al. Sequential protein NMR assignments in the liquid state via sequential data acquisition. J. Magn. Reson. 239, 23–28 (2014).
Wiedemann, C., Ohlenschläger, O., Medagli, B., Onesti, S. & Görlach, M. 1H, 15N and 13C chemical shift assignments for the winged helix domains of two archeal MCM C-termini. Biomol. NMR Assign. Epub ahead of print (2013).
Wittekind, M., Görlach, M., Friedrichs, M., Dreyfuss, G. & Mueller, L. 1H, 13C and 15N NMR assignments and global folding pattern of the RNA-binding domain of the human hnRNP C proteins. Biochemistry 31, 6254–6265 (1992).
Ihle, Y. et al. A novel cGUUAg tetraloop structure with a conserved yYNMGg-type backbone conformation from cloverleaf 1 of bovine enterovirus 1 RNA. Nucleic Acids Res. 33, 2003–2011 (2005).
Kirschstein, A. et al. Heteronuclear J cross-polarisation in liquids using amplitude and phase modulated mixing sequences. J. Biomol. NMR 40, 277–283 (2008).
Kirschstein, A. et al. Broadband homonuclear TOCSY with amplitude and phase-modulated RF mixing schemes. J. Biomol. NMR 40, 227–237 (2008).
States, D. J., Haberkorn, R. A. & Ruben, D. J. A two-dimensional nuclear overhauser experiment with pure absorption phase in four quadrants. J. Magn. Reson. 48, 286–292 (1982).
Piotto, M., Saudek, V. & Sklenár, V. Gradient-tailored excitation for single-quantum NMR spectroscopy of aqueous solutions. J. Biomol. NMR. 2, 661–665 (1992).
Kay, L. E., Xu, G. Y., Singer, A. U., Muhandiram, D. R. & Formankay, J. D. A Gradient-Enhanced HCCH-TOCSY Experiment for Recording Side-Chain 1H and 13C Correlations in H2O Samples of Proteins. J. Magn. Reson. 101, 333–337 (1993).
Morris, G. A. & Freeman, R. Enhancement of nuclear magnetic resonance signals by polarization transfer. J. Am. Chem. Soc. 101, 760–762 (1979).
Flinders, J. & Dieckmann, T. NMR spectroscopy of ribonucleic acids. Prog. Nucl. Magn. Reson. Spectrosc. 48, 137–159 (2006).
Wijmenga, S. S. & van Buuren, B. N. M. The use of NMR methods for conformational studies of nucleic acids. Prog. Nucl. Magn. Reson. Spectrosc. 32, 287–387 (1998).
Kupče, E. r. & Freeman, R. Projection−Reconstruction Technique for Speeding up Multidimensional NMR Spectroscopy. J. Am. Chem. Soc. 126, 6429–6440 (2004).
Acknowledgements
The FLI is a member of the Science Association ‘Gottfried Wilhelm Leibniz’ (WGL) and is financially supported by the Federal Government of Germany and the State of Thuringia. PB and CW were supported by the Leibniz Graduate School on Ageing (LGSA).
Author information
Authors and Affiliations
Contributions
P.B., Y.I. and R.R. jointly conceived the study and wrote the manuscript. R.R. implemented the idea and collected data. P.B. evaluated data and prepared the figures. Y.I. prepared the RNA sample and analysed data, C.W. prepared the protein sample and analysed data, A.K. and C.H. designed NMR mixing sequences, M.G. corrected the manuscript and supervised the study. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Material
Rights and permissions
This work is licensed under a Creative Commons Attribution 3.0 Unported License. The images in this article are included in the article's Creative Commons license, unless indicated otherwise in the image credit; if the image is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the image. To view a copy of this license, visit http://creativecommons.org/licenses/by/3.0/
About this article
Cite this article
Bellstedt, P., Ihle, Y., Wiedemann, C. et al. Sequential acquisition of multi-dimensional heteronuclear chemical shift correlation spectra with 1H detection. Sci Rep 4, 4490 (2014). https://doi.org/10.1038/srep04490
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep04490
This article is cited by
-
PD-L1 degradation is regulated by electrostatic membrane association of its cytoplasmic domain
Nature Communications (2021)
-
Parallel nuclear magnetic resonance spectroscopy
Nature Reviews Methods Primers (2021)
-
A suite of pulse sequences based on multiple sequential acquisitions at one and two radiofrequency channels for solid-state magic-angle spinning NMR studies of proteins
Journal of Biomolecular NMR (2016)
-
HN-NCA heteronuclear TOCSY-NH experiment for 1HN and 15N sequential correlations in (13C, 15N) labelled intrinsically disordered proteins
Journal of Biomolecular NMR (2015)
-
An approach to sequential NMR assignments of proteins: application to chemical shift restraint-based structure prediction
Journal of Biomolecular NMR (2014)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.