Abstract
Mathematical modeling can provide unique insights and predictions about a signaling pathway. Parameter variations allow identification of key reactions that govern signaling features such as the response time that may have a direct impact on the functional outcome. The effect of varying one parameter, however, may depend on values of another. To address the issue, we performed multi-parameter variations of an experimentally validated mathematical model of NF-κB regulatory network and analyzed the inter-relationships of the parameters in shaping key dynamic features. We find that nonlinear dependencies are ubiquitous among parameters. Such phenomena may underlie the emergence of cell type-specific behaviors from essentially the same molecular network. Our results from a multivariate ensemble of models highlight the hypothesis that cell type specificity in signaling phenotype can arise from quantitatively altered strength of reactions in the pathway, in the absence of tissue-specific factors that re-wire the network for a new topology.
Similar content being viewed by others
Introduction
Mathematical modeling of cell signaling pathways is recognized as an important component of molecular systems biology1,2,3,4,5,6. However, it is still a long way before the approach is widely accepted and utilized in mainstream cell biology. This could be attributed to several things. Models are often represented by time-dependent equations that contain kinetic parameters and most of these rate constants are unknown. One can attempt to estimate some of the constants by in vitro assays, but it is not clear how they approximate the in vivo values. Other rate constants are simply not feasible to measure directly and need to be inferred. Therefore, quite often it is judged that mathematical modeling of a pathway is likely to produce a ‘wrong’ model, because it is impossible to determine all rate constants accurately.
So how can one avoid using wrong models? A most relevant clue may come from the experimental counterpart: biological results do not come from studying the behavior of one cell. Even in single cell experiments, a finding is confirmed to be definitive if it is reproduced in a large number of cells. Thus, it would be more appropriate to consider an ensemble of models that occupy a ‘cloud’ of multi-parameter space and correspond to the natural variability of the biological system, rather than looking for ‘the correct model’ (with a single set of parameter values). Exploration of a range of possible parameter values is necessary not only because of the uncertainty in the model parameter values that were inferred or compiled from diverse sources. But also, individual cells are likely to have slightly variable rate constants for any molecular process in the model7. Moreover, studying the parameter space helps understand all the possible behaviors that could be realized under certain pathological or distinct situations.
Here we applied these principles to the widely studied NF-κB pathway and considered an ensemble of models and their signaling characteristics. We present theoretical evidence that context-specific signaling behavior can emerge from parameter dependencies inherent in the nonlinear network of molecular interactions. Our results also imply the existence of situations where reaction kinetics can have discrepant signaling roles in different cell contexts.
Results
NF-κB as a prototypic signaling system within a complex network
NF-κB is an example of latent transcription factors that respond to cell stress and operate in a feedback-controlled network8,9,10. It regulates numerous cell signaling processes and its activity is controlled in part by the level of nuclear translocation. In resting cells, the predominant dimer p65:p50 exists mostly as a cytoplasmic complex bound to its inhibitor IκB proteins. Numerous upstream signals induce degradation of the IκB proteins following phosphorylation by the IκB kinase complex (IKK). This release from latency allows NF-κB to translocate into the nucleus and activate expression of target genes, including several feedback genes8,11.
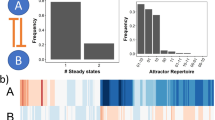
We used a previously published model of NF-κB that captures experimentally observed behaviors reasonably well12,13,14. The mathematical model includes the core NF-κB regulatory network that operates in virtually all cell types (Fig. 1, Tables 1 and 2). Diverse upstream signals that activate the canonical NF-κB pathway converge at the IKK complex. It consists of catalytic subunits IKKα and IKKβ and the regulatory subunit NEMO. The ‘IKK’ in the model corresponds to the active form of IKK, as it has different kinase activities depending on many factors such as its phosphorylation status. The input ‘IKK’ is introduced as an approximate step function in later simulations. All the processes that negatively regulate IKK activity are combined into a first order term with rate constant neg. The IKK-initiated processes, IκBα phosphorylation at serines by IKK, ubiquitination and degradation of IκBα by the proteasome is lumped into a single catalytic reaction with constant r1. r2 is for a similar but less efficient reaction that targets free IκBα. IκBα binding to NF-κB and IKK binding to NF-κB-bound or free IκBα, are all reversible reactions with association and dissociation rates that are roughly based on their binding affinities. dg2 and dg1 denote parameters for the constitutive degradation of NF-κB bound and free IκBα, respectively. The nuclear import and export of NF-κB, IκBα and the complex are included as first order terms. Finally, the induction of IκBα gene by NF-κB is represented by a first order process with a rate constant s and a time delay τ. The delay allows one to incorporate multiple processes during the de novo IκBα synthesis (gene transcription, mRNA processing and export, translation, folding, etc.) into a single term, thereby avoiding unnecessary model complexities that would arise from numerous unknown kinetics involving the intermediates.
A mathematical model of the core regulatory network for NF-κB.
The process diagram represents the individual reactions included in our model. It includes essential regulatory events such as IKK activation, inducible/constitutive degradation of IκBα, nuclear import/export, inducible synthesis of IκBα and post-stimulus attenuation of IKK activity. The quantitative model is described in full by the differential equations in Table 1. The arrows are color-coded based on the reaction type (black: transport, red: complex formation, gray: degradation, purple: multiple molecular processes).
A multivariate ensemble of mathematical models for NF-κB in the high-dimensional parameter space
We explored our differential equations model with an extensive multi-parameter sampling approach. Instead of varying one parameter at a time while fixing all the others, which results in an extremely limited investigation of the system properties, we employed a large set of random parameter combinations for model simulations. Such a high-dimensional ensemble of parameter states better recapitulate the true variability within a population of single cells, because individual cells are unlikely to have identical values for any rate constant. In fact, typical single cell measurements from flow cytometry or quantitative microscopy result in a distribution, not a single value. Each parameter was allowed to vary by two orders of magnitude and 1000 random combinations of parameters were generated by Latin Hypercube sampling method for computational efficiency (see Methods for details).
The randomly generated parameter sets were used to solve the delay differential equations where IKK is activated at t = 0 and to obtain our multi-parameter variation results. Because of the significant coverage of the high dimensional parameter space, the simulated time course profiles consisted of remarkably diverse response patterns (Fig. 2A), providing numerous signaling dynamics that are possible and may be realized in some cellular and microenvironmental contexts.
Multiparameter variation analysis and characteristic measures of signaling dynamics.
(A) Six example time course plots from the simulations. The dynamical model was numerically solved for a random sample of 1000 parameter combinations. Each kinetic parameter was varied by two orders of magnitude around the reference value (see Methods). (B) Four defining characteristics of a temporal profile: F1, F2, F3 and F4 (the period of oscillation if the temporal profile is periodic).
Control parameters that influence characteristic features of signaling dynamics
To identify the parameters that influence NF-κB signaling dynamics, we examined the sensitivity of four defining characteristics in a temporal profile of free nuclear NF-κB (see Fig. 2B), against variations in parameter values. We will consider F1, the integrated activity, which is the area under the time course curve divided by the time interval. It is also mathematically equivalent to the time average response. The first response magnitude F2 is simply the height of the first peak. F3, the response time, is the time from the onset of stimulation to the first peak. F4 is the period of oscillation if the temporal profile is periodic. These features capture some essential aspects of a temporal profile.
To assess sensitivity of feature Fk (k = 1, …, 4) against variations in parameter pi (i = 1, …, 18, as ordered in Table 2), we binned the parameter vectors in the high dimensional parameter space, according to their pi values (regardless of the other parameter values). Bin-average Fk values were obtained and the standard deviation of these values across the bins was taken to be our sensitivity measure Δi Fk. Table S1 shows the parameters sorted by this measure, i.e. how much each parameter influences Fk.
Parameter dependencies are prevalent
Next we addressed our main question by determining whether the influence of a parameter on Fk depends on another parameter. We first illustrate some cases of control parameter pairs with a strong interaction in Fig. 3 (using InterF below; see Methods). Panel A shows how the integrated activity F1 was influenced by rates of IKK association to NF-κB:IκBα complex (a2) and IKK-induced phosphorylation/degradation of NF-κB bound IκBα (r1). Their relationship represented by the best-fit surface indicates that F1 is a decreasing function of r1 for low a2 values, but F1 is roughly a parabola for high a2. This can be interpreted in biological terms as follows. First, the integrated activity of NF-κB over time is a most likely determinant of the transcriptional output of direct NF-κB-dependent genes12. Then panel A implies that the gene output can be greater for slower signal-dependent degradation of IκBα when IKK binding to substrate is relatively slow. But in a different cellular context where substrate recognition of IKK is faster (due to local tethering, for example), the transcriptional output may be generally elevated with a slight moderation at a mid-range degradation rate of IκBα.
Nonlinearities in sensitivity of NF-κB signaling characteristics to kinetic parameter values.
(A) The surface plot shows the coordinate effect of varying a2 and r1 upon F1. The smooth surface was obtained by locally fitting the individual simulated results (see Methods). The x and y axes are in log10 scale. (B) A similar plot for F1 against iI and a3. (C) A plot for F1 against dg2 and τ. (D) A plot for F3 against d1 and s. (E) A plot for F3 against s and dg1.
It is also to be noted that the parameter dependencies are not necessarily symmetric, i.e. the effect of r1 depended on a2, but a2 did not depend on r1(Fig. 3A). Our simulations also found F1 to depend on iI and a3 in a nonlinear fashion (Fig. 3B). If the import rate iI was low, F1 increased with the association rate a3, but if iI was high, a3 had little effect on F1.
Figure 3C shows another pair of inter-dependent parameters caused by a more complex nonlinearity in their influence on F1. When the constitutive degradation of NF-κB bound IκBα (dg2) was minimal, shorter time delays involved in IκBα re-synthesis (τ) resulted in higher integrated NF-κB activity. However, this trend switched to a completely different outcome when the degradation of NF-κB bound IκBα was constitutively higher: There was an optimal time delay that produced maximal transcriptional activity in such a condition. Therefore we conclude that the effect of τ depends on dg2.
The response time F3 had differential dependence on d1 and s in that F3 was minimized for a distinct value of IκBα synthesis rate s only if d1, the dissociation rate of NF-κB:IκBα was high (Fig. 3D). Figure 3E indicates that the constitutive degradation of free IκBα (dg1) affected how the IκBα re-synthesis rate (s) influenced the response time, F3. If the constitutive degradation was low, the response was fastest at an optimal IκBα induction rate. If, on the other hand, the degradation was constitutively fast, the response was generally fast regardless of the synthesis rate.
Finally, we examined all the inter-dependencies systematically in the following way. For each combination (i, j, k), the coordinate effect of pi and pj on Fk was extracted by fitting the data with a smooth surface as shown in Fig. 3. We defined a quantity InterFk(i, j) to capture the deviation from the independence of pi effect on Fk from pj (see Methods). Nonzero InterFk(i, j) values indicate the presence of an inter-dependency for the two parameters, where the parameter pi had a different qualitative effect on Fk depending on the value range of pj. There were numerous such incidences and some pairs (i, j) corresponded to parameters that had weak influences on Fk, where any dependencies would impose an insignificant effect. So, for cases in Fig. 3, we chose those pairs that had significant influence as single control parameters and had high interF values.
We summarize all the results in the ‘parameter dependency map’ in Fig. 4 (strong to weak interF in yellow to red) which shows the prevalence of inter-dependencies among parameters in shaping the signaling dynamics. Most parameters had differential effects on Fk depending on the values of one or more parameters. On the other extreme, some vertical stretches of red are discernable from the map and correspond to parameters that did not depend on the other parameters. Most of these exert strong control over the relevant Fk. For example, the time delay (τ) and the inactivation rate of IKK (neg) were critical parameters that determine the period F4 (see Table S1) and their influence on the period were not affected by other parameters.
Parameter dependency map.
The color-coded matrix plots display the extent of interaction between all parameter pairs, using the measure InterF (see Methods) for each Fi. The order of model parameters on the axes is the same as in Table 2. The color scale from red to yellow corresponds to low to high interF values.
We explore a possible manifestation of our findings by illustrating a scenario that corresponds to Fig. 3A in more detail (Fig. 5). In mathematical terms, we found that F1(r1) is a decreasing function for low a2 (lower arrow in the surface plot) and is an increasing function for a higher range of a2 (upper arrow). In biological terms, this implies that the transcriptional consequence of inhibiting signal-induced degradation of IκBα can vary depending on the association rate of IKK to its substrate, NF-κB bound IκBα. This rate, in turn, may well depend on the cell type under study. Cellular features such as the organization and volume of the cytoplasm, or local tethering of kinase scaffolds, are different for distinct cell types. Smaller cytoplasm and local clustering can endow the cells with faster substrate recognition with little need for diffusion-based association. Cell type A represents such a situation. When such cells are treated with an inhibitor that reduces the induced degradation of IκBα, the integrated activity of NF-κB, therefore target gene output, is decreased (indicated by the direction of the upper arrow on the surface plot). However, just the opposite outcome is expected for the same perturbation in another cell type B, where the IKK recognition of its substrate is relatively slow. Other dependencies we found can similarly be elaborated with concrete biological interpretations.
Biological manifestations: a plausible scenario.
Implications from the dependence of r1, signal-induced degradation of IκBα, on a2, the association rate of IKK to its substrate, in affecting F1, the integrated activity of NF-κB. The surface plot is from the result in Fig. 3A. Assuming F1 is a primary determinant of transcriptional output of NF-κB dependent genes, inhibition of signal-dependent degradation of IκBα (that lowers r1) has opposite effect on transcriptional output in two cell types A and B, where the IKK substrate recognition is fast and slow (with distinct ranges of parameter a2), respectively.
Inter-dependencies of reaction kinetics may well explain, perhaps to a significant extent, the cell type specificity of the signaling roles of numerous factors that seem to have context-dependent actions15,16. We note that most signaling pathways possess feedback structures and that the ensuing system nonlinearity is likely to cause inter-dependencies of parameters. To this end, we have looked into another signaling pathway, Wnt/β-catenin and found a similar extent of parameter dependencies (Myong-Hee Sung, Songjoon Baek, Kwang-Hyun Cho, unpublished data).
Discussion
A significant hindrance in translating the knowledge from a particular quantitative signaling model to a real-world molecular system is the lack of in vivo measurements of the kinetic parameters from the relevant context, such as particular cell lines or primary tissues. We have looked into the effects of varying kinetic parameters upon signaling characteristics and their dependencies on other parameters. For example, suppose that a higher degradation rate of a signaling protein A has the effect of shortening the response time. But this effect may depend on whether the synthesis rate of protein B is within a certain range. The effect of A on response time may even be opposite in other conditions or cellular contexts. By extensive simulations of an NF-κB model, we demonstrate that such a phenomenon can be widespread. This may be a source of apparently discrepant behaviors of the same cellular signaling system in different biological contexts.
The phenomenon seen here may underlie the differential effect of a given molecular process/reaction that is dependent upon distinct levels or efficiencies of another reaction. In general, different cell types are thought to have differences in splice variants17,18, organization of the genome into accessible chromatin domains19, basal turnover of signaling factors, subunit composition of holoenzymes that may affect catalysis rates20 and more. Here our results explain how quantitative differences in such key molecular systems can lead to qualitatively distinct signaling behaviors.
Methods
A mathematical model of NF-κB signaling network
We used a published mathematical model12. Briefly, the delay differential equations (DDE) model described in21 was modified by including a term (neg in Fig. 1) to represent the inactivation of IKK by various mechanisms including A20, CYLD and IKK autophosphorylation11. These IKK inactivation mechanisms lack single cell data on their kinetic parameters and could not be represented individually. The 9-variable DDE model is shown in Table 1 and the reference parameter values are listed in Table 2.
Multi-parameter variation and model simulations
Each kinetic parameter was varied by 2 orders of magnitude around the reference value (from 0.1- to 10-fold) and was randomly combined with others by the Latin Hypercube sampling method to limit the total number of simulations. The time delay parameter for IκBα synthesis was constrained to vary between 30 and 55 minutes to avoid an unrealistic range. Specifically, parameter combinations were generated by the following procedure. For the j-th parameter, we subdivide the range of the parameter into n ( = 5) subintervals of equal size. Then randomly sample n values (pij, i = 1, …, n), one from each subinterval, for the j-th parameter. The uniform sampling was done on the logscale for all parameters. To combine these values of individual parameters to generate sets of parameter values, we randomly permute the n values for each parameter to get the parameter vectors, i.e. we permute the elements of each column of the matrix pij separately and use the rows as the parameter vectors. This sampling method was implemented by the MATLAB function ‘lhsdesign’ to produce 1000 sets of parameter values. These parameter vectors were used for DDE simulations.
The initial condition for numerical solutions of DDE was provided by specifying a constant history of I = 0.03 µM; NF:I = 0.04 µM; In = 0.03 µM; the other variables = 0.
The time delay was given by the parameter τ. All simulations were run by using the MATLAB solver ‘dde23’ on the time interval [−10 h, 12 h]. IKK activation was introduced at t = 0 by a sharp Gaussian k(t) (standard deviation = 5 min) multiplied by 0.025 µM/min. Evaluation of NFn from each numerical solution was obtained at the 5 min resolution grid of time points spanning [0 h, 12 h]. The evaluated series NFn(t) for each simulation was taken as its time course profile for subsequent analyses.
Dynamic measures Fi
For each time course profile, Fi (i = 1, …, 4) values were calculated as follows.

where T is the time course interval (12 h) and [total NF] is the amount of all NF-containing molecular species (determined by the initial condition and fixed at 0.04 µM). F2 and F3 were obtained by finding the first time point t* (> 0) where the time series becomes decreasing, i.e. NFn(t*+dt) − NFn(t*) ≤ 0. Then

For the period F4, all time course profiles were analyzed by Fourier analysis to sort for the oscillating profiles. NFn was considered oscillating if the periodogram from the fast Fourier transform had a detectable peak between 0 h and 5 h either as a global maximum or a local maximum which is at least 0.7 of the global maximum. 345 cases (among 1000) passed the criteria and were used for the analysis of F4.
Identification of single parameters that control Fi
Given Fi, its sensitivity against varying parameter pj was measured by Δj Fi = σ(mean {Fi(p) | pj in k-th bin of j-th parameter}) where σ is the standard deviation of bin-averaged Fi values. In particular, pj was subdivided into 10 bins in log scale and then all the parameter vectors were binned according to their value range in the j-th dimension. The standard deviation of the bin-average values across the bins was taken as our sensitivity measure Δj Fi.
Assessment of pairwise parameter interactions upon Fi
For each Fk - pi - pj combination, we extracted the predominant effect of the parameter pairs (pi, pj) on Fk by considering a smooth surface fit zk(pi, pj) of the data distribution. This was implemented by Loess fitting Fk against (pi, pj) in log scale with a span of 0.5 and by evaluating on a fixed grid of 20 by 20 points over the parameter domain. The pj-conditional effect of pi, dFk, was set to be (zk(pimax, pj) – zk(pimin, pj))/zk(pimid, pj), where pimax, pimin, pimid are maximum, minimum and center grid values of pi, respectively. The pairwise interaction measure InterFk(i, j) was then defined as

All Loess-fitting and interaction measures were computed in R (http://www.r-project.org).
References
Suel, G. M., Garcia-Ojalvo, J., Liberman, L. M. & Elowitz, M. B. An excitable gene regulatory circuit induces transient cellular differentiation. Nature 440, 545–550 (2006).
Mellman, I. & Misteli, T. Computational cell biology. J Cell Biol 161, 463–464 (2003).
Sible, J. C. & Tyson, J. J. Mathematical modeling as a tool for investigating cell cycle control networks. Methods 41, 238–247 (2007).
von Dassow, G., Meir, E., Munro, E. M. & Odell, G. M. The segment polarity network is a robust developmental module. Nature 406, 188–192 (2000).
Geva-Zatorsky, N. et al. Oscillations and variability in the p53 system. Mol Syst Biol 2, 2006 0033 (2006).
Longabaugh, W. J., Davidson, E. H. & Bolouri, H. Computational representation of developmental genetic regulatory networks. Dev Biol 283, 1–16 (2005).
Voss, T. C. et al. Combinatorial probabilistic chromatin interactions produce transcriptional heterogeneity. J Cell Sci 122, 345–356 (2009).
O'Dea, E. & Hoffmann, A. The regulatory logic of the NF-kappaB signaling system. Cold Spring Harb Perspect Biol 2, a000216.
Escoubet-Lozach, L. et al. Mechanisms establishing TLR4-responsive activation states of inflammatory response genes. PLoS Genet 7, e1002401.
Siggers, T. et al. Principles of dimer-specific gene regulation revealed by a comprehensive characterization of NF-kappaB family DNA binding. Nat Immunol 13, 95–102.
Hayden, M. S. & Ghosh, S. Shared principles in NF-kappaB signaling. Cell 132, 344–362 (2008).
Sung, M. H. et al. Sustained oscillations of NF-kappaB produce distinct genome scanning and gene expression profiles. PLoS One 4, e7163 (2009).
Nelson, D. E. et al. Oscillations in NF-kappaB signaling control the dynamics of gene expression. Science 306, 704–708 (2004).
Hoffmann, A., Levchenko, A., Scott, M. L. & Baltimore, D. The IkappaB-NF-kappaB signaling module: temporal control and selective gene activation. Science 298, 1241–1245 (2002).
Chen, F., Beezhold, K. & Castranova, V. Tumor promoting or tumor suppressing of NF-kappa B, a matter of cell context dependency. Int Rev Immunol 27, 183–204 (2008).
Dutta, J., Fan, Y., Gupta, N., Fan, G. & Gelinas, C. Current insights into the regulation of programmed cell death by NF-kappaB. Oncogene 25, 6800–6816 (2006).
Phair, R. D. et al. Global nature of dynamic protein-chromatin interactions in vivo: three-dimensional genome scanning and dynamic interaction networks of chromatin proteins. Mol Cell Biol 24, 6393–6402 (2004).
Hoppler, S. & Kavanagh, C. L. Wnt signalling: variety at the core. J Cell Sci 120, 385–393 (2007).
John, S. et al. Interaction of the glucocorticoid receptor with the chromatin landscape. Mol Cell 29, 611–624 (2008).
Burke, P. V. & Poyton, R. O. Structure/function of oxygen-regulated isoforms in cytochrome c oxidase. J Exp Biol 201, 1163–1175 (1998).
Sung, M. H. & Simon, R. In silico simulation of inhibitor drug effects on nuclear factor-kappaB pathway dynamics. Mol Pharmacol 66, 70–75 (2004).
Acknowledgements
We thank Alessandra Agresti for helpful discussions. This work was supported in part by the Intramural Research Program of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
M.H.S. designed the study, performed the computation, analyzed results and wrote the paper. G.L.H. supported the study and commented on the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Table S1
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Sung, MH., Hager, G. Nonlinear Dependencies of Biochemical Reactions for Context-specific Signaling Dynamics. Sci Rep 2, 616 (2012). https://doi.org/10.1038/srep00616
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00616
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.