« Prev Next »
Small genome sequences, such as those of viruses and bacteria, have shed light on the basic principles of genomics. Thanks to such sequence information, researchers now know how small a genome can be and how few genes are required for self-sufficient life. Furthermore, genomes from organisms like E. coli and baker's yeast are helpful tools in the effort to define the basic biochemical networks within more complex organisms.
Sequencing Small Genomes
The bacterium that most commonly causes ear infections in children is Haemophilus influenza, and in 1995, the H. influenza genome was the first bacterial genome to be fully sequenced (Fleischmann et al., 1995). It was also the first completed sequence for a free-living organism. This landmark effort proved the utility of the then-novel technique of whole-genome shotgun (WGS) sequencing. Basically, the researchers cut the bacterial DNA into many small, easily sequenced pieces and then relied upon computer algorithms to align overlapping segments and thus assemble the entire genome. Despite early concerns, WGS has since become the method of choice for most sequencing projects large and small.
Of average size and DNA composition for a bacterium, H. influenza contains 1.8 million bases and about 1,700 genes, with very little repetitive DNA. Interestingly, more than 1,000 of the organism's genes have been identified as orthologs to known genes from other organisms, including humans (Davies, 2001). The gene families present in H. influenza, including genes for such functions as translation, transport, energy production, and envelope structure, emphasize the commonalities of all life and the lessons that can be learned from comparative genomics. The H. influenza genome also served as an early description of the minimal set of genes required for free-living life, and it strongly affirmed the relevance of spending time and money to sequence the genomes of even simple organisms.
Shortly after work on the H. influenza genome was completed, researcher J. Craig Venter next sequenced the Mycoplasma genitalium genome as part of his continued search for the minimal set of genes required for life (Fraser et al., 1995). Unlike other organisms on the to-be-sequenced lists of the 1990s, M. genitalium (its name refers to its preferred habitat in the human body) is neither a laboratory model to be studied nor a pathogen to be conquered. Rather, it is one of the smallest free-living organisms known to man, with only half a million bases and a mere 500 or so genes. Scientists continue to be interested in this bacterium because of its miniscule size, which poses a fundamental question: How small and simple can an organism be while still satisfying the definition of life? It is hoped that in the years to come, disrupting gene functions in M. genitalium one-by-one might shed light upon this inquiry, thus revealing the essential genetic components of cellular viability.
Presently, thousands of viral, bacterial, archaeal, and single-celled eukaryotic genomes are being sequenced to completion as part of large-scale metagenomic sequencing efforts, such as the Human Microbiome Project (Turnbaugh et al., 2007) and other environmental sequencing projects (Tyson et al., 2004). Sequences from microorganisms that endure extreme or toxic environmental conditions give researchers insight into useful genes that could be exploited to clean up pollutants or digest organic materials for biofuels, for example. Although using naturally occurring microbes in such ways might be feasible, much greater attention continues to be focused on engineering organisms that can carry out customized biochemistry using synthetic genomics. But what does this field entail?
Synthetic Genomics
These findings challenge the current hypothesis regarding the pathogenicity of the Spanish flu virus, which argues that the virus was so very deadly because humans were more susceptible to infection and illness in 1918 (Lamb & Jackson, 2005). In particular, the experiments of Tumpey and colleagues (2005) indicate that the severity of the Spanish flu is based upon the virus rather than upon the vulnerability of human hosts. Interestingly, the 1918 strain is sensitive to today's antiviral drugs, suggesting that modern technology might have helped prevent this pandemic.

From One Organism to Many
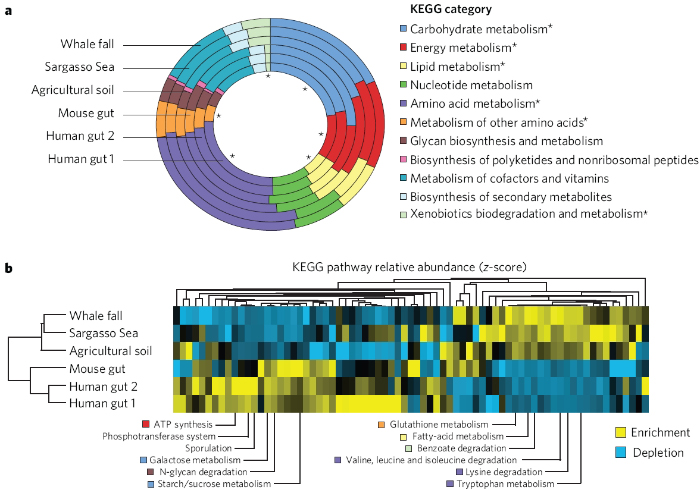
Recent en masse sequencing of various bacteria that colonize humans, such as intestinal flora, has greatly expanded researchers' understanding of our relationship with and dependence upon beneficial microbes. Although the specific functions of each of the bacterial species that inhabit our digestive tracts are not yet known, our microbiota are estimated to outnumber our own body cells by a factor of 10. The Human Microbiome Project seeks to sequence and characterize these microbes—but as whole communities rather than individual species (Turnbaugh et al., 2007). The overriding goals of the Human Microbiome Project include identifying new ways we might be predisposed to disease, developing approaches with which to manipulate human microbiota, and optimizing the function of these microbes in the context of an individual's physiology. Some combinations of microbes, for example, might be found to influence metabolism, thus making them helpful in controlling obesity (Figure 3).
In the near future, synthetic genomes promise to allow scientists to create increasingly tailored and specialized microbes that can be used to produce pharmaceuticals, degrade pollutants, and perform other complex biochemistry for biotechnological purposes. Despite these promises, the wealth of sequence data from human pathogens coupled with advances in synthetic genomics also poses certain ethical and safety-related challenges. What if artificial life-forms contaminate the biosphere? What if unethical governments or terrorist groups synthesize new biological agents with multi-drug resistance or new toxins that can be used as weapons? Sequencing the smallest organisms reveals the essential elements of life and provides opportunities in genetics and biotechnology. At the same time, however, building life from its core components necessitates an ongoing public discussion of the potential impacts of this practice upon society.
References and Recommended Reading
Davies, K. Cracking the Genome: Inside the Race to Unlock Human DNA (New York, Free Press, 2001)
Endy, D. Reconstruction of the Genomes. Science, 319, 1197-1197 (2008)
Fleischmann, R. D., et al. Whole-genome random sequencing and assembly of Haemophilus influenzae Rd. Science 269, 496-512 (1995)
Fraser, C. M., et al. The minimal gene complement of Mycoplasma genitalium. Science 270, 397-403 (1995)
Gibson, D. G., et al. Complete chemical synthesis, assembly, and cloning of a Mycoplasma genitalium genome. Science 319, 1215-1220 (2008) doi:10.1126/science.1151721
Goffeau, A., et al. Life with 6000 genes. Science 274, 546-567 (1996) doi:10.1126/science.274.5287.546
Lamb, R. A., & Jackson, D. Extinct 1918 virus comes alive. Nature Medicine 11, 1154-1156 (2005) (link to article)
Taubenberger, J. K., et. al. Characterization of the 1918 influenza virus polymerase genes. Nature 437, 889-893 (2005), doi:10.1038/nature04230 (link to article)
Tumpey, T. M., et al. Characterization of the reconstructed 1918 Spanish influenza pandemic virus. Science 310, 77-80 (2005)
Turnbaugh, P. J., et al. The Human Microbiome Project. Nature 449, 804-810 (2007) doi:10.1038/nature06244 (link to article)
Tyson, G. W., et al. Community structure and metabolism through reconstruction of microbial genomes from the environment. Nature 428, 37-43 (2004) (link to article)



 Figure 1
Figure 1


























