Abstract
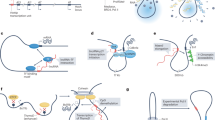
The central dogma of molecular biology states that the flow of genetic information moves from DNA to RNA to protein. However, in the last decade this dogma has been challenged by new findings on non-coding RNAs (ncRNAs) such as microRNAs (miRNAs). More recently, long non-coding RNAs (lncRNAs) have attracted much attention due to their large number and biological significance. Many lncRNAs have been identified as mapping to regulatory elements including gene promoters and enhancers, ultraconserved regions and intergenic regions of protein-coding genes. Yet, the biological function and molecular mechanisms of lncRNA in human diseases in general and cancer in particular remain largely unknown. Data from the literature suggest that lncRNA, often via interaction with proteins, functions in specific genomic loci or use their own transcription loci for regulatory activity. In this review, we summarize recent findings supporting the importance of DNA loci in lncRNA function and the underlying molecular mechanisms via cis or trans regulation, and discuss their implications in cancer. In addition, we use the 8q24 genomic locus, a region containing interactive SNPs, DNA regulatory elements and lncRNAs, as an example to illustrate how single-nucleotide polymorphism (SNP) located within lncRNAs may be functionally associated with the individual’s susceptibility to cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Djebali S, Davis CA, Merkel A, Dobin A, Lassmann T, Mortazavi A et al. Landscape of transcription in human cells. Nature 2012; 489: 101–108.
Comings DE . The structure and function of chromatin. Adv Hum Genet 1972; 3: 237–431.
Calin GA, Croce CM . MicroRNA signatures in human cancers. Nat Rev Cancer 2006; 6: 857–866.
Ling H, Fabbri M, Calin GA . MicroRNAs and other non-coding RNAs as targets for anticancer drug development. Nat Rev Drug Discov 2013; 12: 847–865.
Esteller M . Non-coding RNAs in human disease. Nat Rev Genet 2011; 12: 861–874.
Derrien T, Johnson R, Bussotti G, Tanzer A, Djebali S, Tilgner H et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res 2012; 22: 1775–1789.
Wang KC, Chang HY . Molecular mechanisms of long noncoding RNAs. Mol Cell 2011; 43: 904–914.
Schmitz KM, Mayer C, Postepska A, Grummt I . Interaction of noncoding RNA with the rDNA promoter mediates recruitment of DNMT3b and silencing of rRNA genes. Gene Dev 2010; 24: 2264–2269.
Chu C, Qu K, Zhong FL, Artandi SE, Chang HY . Genomic maps of long noncoding RNA occupancy reveal principles of RNA-chromatin interactions. Mol Cell 2011; 44: 667–678.
Hindorff LA, Sethupathy P, Junkins HA, Ramos EM, Mehta JP, Collins FS et al. Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc Natl Acad Sci USA 2009; 106: 9362–9367.
Ulitsky I, Shkumatava A, Jan CH, Sive H, Bartel DP . Conserved function of lincRNAs in vertebrate embryonic development despite rapid sequence evolution. Cell 2011; 147: 1537–1550.
Fatica A, Bozzoni I . Long non-coding RNAs: new players in cell differentiation and development. Nat Rev Genet 2014; 15: 7–21.
Batista PJ, Chang HY . Long noncoding RNAs: cellular address codes in development and disease. Cell 2013; 152: 1298–1307.
Wapinski O, Chang HY . Long noncoding RNAs and human disease. Trends Cell Biol 2011; 21: 354–361.
Mercer TR, Mattick JS . Structure and function of long noncoding RNAs in epigenetic regulation. Nat Struct Mol Biol 2013; 20: 300–307.
Geisler S, Coller J . RNA in unexpected places: long non-coding RNA functions in diverse cellular contexts. Nat Rev Mol Cell Biol 2013; 14: 699–712.
Yang L, Froberg JE, Lee JT . Long noncoding RNAs: fresh perspectives into the RNA world. Trends Biochem Sci 2014; 39: 35–43.
Cech TR, Steitz JA . The noncoding RNA revolution-trashing old rules to forge new ones. Cell 2014; 157: 77–94.
Ulitsky I, Bartel DP . lincRNAs: genomics, evolution, and mechanisms. Cell 2013; 154: 26–46.
Lee JT, Bartolomei MS . X-inactivation, imprinting, and long noncoding RNAs in health and disease. Cell 2013; 152: 1308–1323.
Kim HS, Minna JD, White MA . GWAS meets TCGA to illuminate mechanisms of cancer predisposition. Cell 2013; 152: 387–389.
Calin GA, Dumitru CD, Shimizu M, Bichi R, Zupo S, Noch E et al. Frequent deletions and down-regulation of micro- RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci USA 2002; 99: 15524–15529.
Gong C, Maquat LE . lncRNAs transactivate STAU1-mediated mRNA decay by duplexing with 3' UTRs via Alu elements. Nature 2011; 470: 284–288.
Mariner PD, Walters RD, Espinoza CA, Drullinger LF, Wagner SD, Kugel JF et al. Human Alu RNA is a modular transacting repressor of mRNA transcription during heat shock. Mol Cell 2008; 29: 499–509.
Levine M, Tjian R . Transcription regulation and animal diversity. Nature 2003; 424: 147–151.
Core LJ, Waterfall JJ, Lis JT . Nascent RNA sequencing reveals widespread pausing and divergent initiation at human promoters. Science 2008; 322: 1845–1848.
Seila AC, Calabrese JM, Levine SS, Yeo GW, Rahl PB, Flynn RA et al. Divergent transcription from active promoters. Science 2008; 322: 1849–1851.
Hung T, Wang Y, Lin MF, Koegel AK, Kotake Y, Grant GD et al. Extensive and coordinated transcription of noncoding RNAs within cell-cycle promoters. Nat Genet 2011; 43: 621–629.
Blackwood EM, Kadonaga JT . Going the distance: a current view of enhancer action. Science 1998; 281: 60–63.
Orom UA, Shiekhattar R . Long noncoding RNAs usher in a new era in the biology of enhancers. Cell 2013; 154: 1190–1193.
Kim TK, Hemberg M, Gray JM, Costa AM, Bear DM, Wu J et al. Widespread transcription at neuronal activity-regulated enhancers. Nature 2010; 465: 182–187.
Orom UA, Derrien T, Beringer M, Gumireddy K, Gardini A, Bussotti G et al. Long noncoding RNAs with enhancer-like function in human cells. Cell 2010; 143: 46–58.
Koch F, Fenouil R, Gut M, Cauchy P, Albert TK, Zacarias-Cabeza J et al. Transcription initiation platforms and GTF recruitment at tissue-specific enhancers and promoters. Nat Struct Mol Biol 2011; 18: 956–963.
Feng J, Bi C, Clark BS, Mady R, Shah P, Kohtz JD . The Evf-2 noncoding RNA is transcribed from the Dlx-5/6 ultraconserved region and functions as a Dlx-2 transcriptional coactivator. Gene Dev 2006; 20: 1470–1484.
Wang KC, Yang YW, Liu B, Sanyal A, Corces-Zimmerman R, Chen Y et al. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature 2011; 472: 120–124.
Yang L, Lin C, Jin C, Yang JC, Tanasa B, Li W et al. lncRNA-dependent mechanisms of androgen-receptor-regulated gene activation programs. Nature 2013; 500: 598–602.
Bejerano G, Pheasant M, Makunin I, Stephen S, Kent WJ, Mattick JS et al. Ultraconserved elements in the human genome. Science 2004; 304: 1321–1325.
Calin GA, Liu CG, Ferracin M, Hyslop T, Spizzo R, Sevignani C et al. Ultraconserved regions encoding ncRNAs are altered in human leukemias and carcinomas. Cancer Cell 2007; 12: 215–229.
Braconi C, Valeri N, Kogure T, Gasparini P, Huang N, Nuovo GJ et al. Expression and functional role of a transcribed noncoding RNA with an ultraconserved element in hepatocellular carcinoma. Proc Natl Acad Sci USA 2011; 108: 786–791.
Ferdin J, Nishida N, Wu X, Nicoloso MS, Shah MY, Devlin C et al. HINCUTs in cancer: hypoxia-induced noncoding ultraconserved transcripts. Cell Death Differ 2013; 20: 1675–1687.
Ling H, Spizzo R, Atlasi Y, Nicoloso M, Shimizu M, Redis RS et al. CCAT2, a novel noncoding RNA mapping to 8q24, underlies metastatic progression and chromosomal instability in colon cancer. Genome Res 2013; 23: 1446–1461.
Liz J, Portela A, Soler M, Gomez A, Ling H, Michlewski G et al. Regulation of pri-miRNA processing by a long noncoding RNA transcribed from an ultraconserved region. Mol Cell 2014; 55: 138–147.
Katayama S, Tomaru Y, Kasukawa T, Waki K, Nakanishi M, Nakamura M et al. Antisense transcription in the mammalian transcriptome. Science 2005; 309: 1564–1566.
Wang XJ, Gaasterland T, Chua NH . Genome-wide prediction and identification of cis-natural antisense transcripts in Arabidopsis thaliana. Genome Biol 2005; 6: R30.
Faghihi MA, Wahlestedt C . Regulatory roles of natural antisense transcripts. Nat Rev Mol Cell Biol 2009; 10: 637–643.
Yu W, Gius D, Onyango P, Muldoon-Jacobs K, Karp J, Feinberg AP et al. Epigenetic silencing of tumour suppressor gene p15 by its antisense RNA. Nature 2008; 451: 202–206.
Pasmant E, Laurendeau I, Heron D, Vidaud M, Vidaud D, Bieche I . Characterization of a germ-line deletion, including the entire INK4/ARF locus, in a melanoma-neural system tumor family: identification of ANRIL, an antisense noncoding RNA whose expression coclusters with ARF. Cancer Res 2007; 67: 3963–3969.
Yap KL, Li S, Munoz-Cabello AM, Raguz S, Zeng L, Mujtaba S et al. Molecular interplay of the noncoding RNA ANRIL and methylated histone H3 lysine 27 by polycomb CBX7 in transcriptional silencing of INK4a. Mol Cell 2010; 38: 662–674.
Beltran M, Puig I, Pena C, Garcia JM, Alvarez AB, Pena R et al. A natural antisense transcript regulates Zeb2/Sip1 gene expression during Snail1-induced epithelial-mesenchymal transition. Gene Dev 2008; 22: 756–769.
Rinn JL, Kertesz M, Wang JK, Squazzo SL, Xu X, Brugmann SA et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 2007; 129: 1311–1323.
Tsai MC, Manor O, Wan Y, Mosammaparast N, Wang JK, Lan F et al. Long noncoding RNA as modular scaffold of histone modification complexes. Science 2010; 329: 689–693.
Huarte M, Guttman M, Feldser D, Garber M, Koziol MJ, Kenzelmann-Broz D et al. A large intergenic noncoding RNA induced by p53 mediates global gene repression in the p53 response. Cell 2010; 142: 409–419.
Lee JT . Lessons from X-chromosome inactivation: long ncRNA as guides and tethers to the epigenome. Gene Dev 2009; 23: 1831–1842.
Bertani S, Sauer S, Bolotin E, Sauer F . The noncoding RNA Mistral activates Hoxa6 and Hoxa7 expression and stem cell differentiation by recruiting MLL1 to chromatin. Mol Cell 2011; 43: 1040–1046.
Di Ruscio A, Ebralidze AK, Benoukraf T, Amabile G, Goff LA, Terragni J et al. DNMT1-interacting RNAs block gene-specific DNA methylation. Nature 2013; 503: 371–376.
Jeon Y, Lee JT . YY1 tethers Xist RNA to the inactive X nucleation center. Cell 2011; 146: 119–133.
Simon MD, Pinter SF, Fang R, Sarma K, Rutenberg-Schoenberg M, Bowman SK et al. High-resolution Xist binding maps reveal two-step spreading during X-chromosome inactivation. Nature 2013; 504: 465–469.
Maenner S, Blaud M, Fouillen L, Savoye A, Marchand V, Dubois A et al. 2-D structure of the A region of Xist RNA and its implication for PRC2 association. PLoS Biol 2010; 8: e1000276.
Nagano T, Mitchell JA, Sanz LA, Pauler FM, Ferguson-Smith AC, Feil R et al. The Air noncoding RNA epigenetically silences transcription by targeting G9a to chromatin. Science 2008; 322: 1717–1720.
Lee JT, Davidow LS, Warshawsky D . Tsix, a gene antisense to Xist at the X-inactivation centre. Nat Genet 1999; 21: 400–404.
Sun BK, Deaton AM, Lee JT . A transient heterochromatic state in Xist preempts X inactivation choice without RNA stabilization. Mol Cell 2006; 21: 617–628.
Ogawa Y, Sun BK, Lee JT . Intersection of the RNA interference and X-inactivation pathways. Science 2008; 320: 1336–1341.
Lai F, Orom UA, Cesaroni M, Beringer M, Taatjes DJ, Blobel GA et al. Activating RNAs associate with Mediator to enhance chromatin architecture and transcription. Nature 2013; 494: 497–501.
Melo CA, Drost J, Wijchers PJ, van de Werken H, de Wit E, Oude Vrielink JA et al. eRNAs are required for p53-dependent enhancer activity and gene transcription. Mol Cell 2013; 49: 524–535.
Khalil AM, Guttman M, Huarte M, Garber M, Raj A, Rivea Morales D et al. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc Natl Acad Sci USA 2009; 106: 11667–11672.
Sanford JR, Wang X, Mort M, Vanduyn N, Cooper DN, Mooney SD et al. Splicing factor SFRS1 recognizes a functionally diverse landscape of RNA transcripts. Genome Res 2009; 19: 381–394.
Zhang X, Rice K, Wang Y, Chen W, Zhong Y, Nakayama Y et al. Maternally expressed gene 3 (MEG3) noncoding ribonucleic acid: isoform structure, expression, and functions. Endocrinology 2010; 151: 939–947.
Novikova IV, Hennelly SP, Sanbonmatsu KY . Structural architecture of the human long non-coding RNA, steroid receptor RNA activator. Nucleic Acids Res 2012; 40: 5034–5051.
Plath K, Fang J, Mlynarczyk-Evans SK, Cao R, Worringer KA, Wang H et al. Role of histone H3 lysine 27 methylation in X inactivation. Science 2003; 300: 131–135.
Allen TA, Von Kaenel S, Goodrich JA, Kugel JF . The SINE-encoded mouse B2 RNA represses mRNA transcription in response to heat shock. Nat Struct Mol Biol 2004; 11: 816–821.
Yakovchuk P, Goodrich JA, Kugel JF . B2 RNA and Alu RNA repress transcription by disrupting contacts between RNA polymerase II and promoter DNA within assembled complexes. Proc Natl Acad Sci USA 2009; 106: 5569–5574.
Ning S, Zhao Z, Ye J, Wang P, Zhi H, Li R et al. LincSNP: a database of linking disease-associated SNPs to human large intergenic non-coding RNAs. BMC Bioinformatics 2014; 15: 152.
Pasmant E, Sabbagh A, Vidaud M, Bieche I . ANRIL, a long, noncoding RNA, is an unexpected major hotspot in GWAS. FASEB J 2011; 25: 444–448.
Pasmant E, Sabbagh A, Masliah-Planchon J, Ortonne N, Laurendeau I, Melin L et al. Role of noncoding RNA ANRIL in genesis of plexiform neurofibromas in neurofibromatosis type 1. J Natil Cancer Inst 2011; 103: 1713–1722.
Matouk IJ, DeGroot N, Mezan S, Ayesh S, Abu-lail R, Hochberg A et al. The H19 non-coding RNA is essential for human tumor growth. PloS one 2007; 2: e845.
Zhang L, Yang F, Yuan JH, Yuan SX, Zhou WP, Huo XS et al. Epigenetic activation of the MiR-200 family contributes to H19-mediated metastasis suppression in hepatocellular carcinoma. Carcinogenesis 2013; 34: 577–586.
Verhaegh GW, Verkleij L, Vermeulen SH, den Heijer M, Witjes JA, Kiemeney LA . Polymorphisms in the H19 gene and the risk of bladder cancer. Eur Urol 2008; 54: 1118–1126.
Easton DF, Pooley KA, Dunning AM, Pharoah PD, Thompson D, Ballinger DG et al. Genome-wide association study identifies novel breast cancer susceptibility loci. Nature 2007; 447: 1087–1093.
Liu Y, Pan S, Liu L, Zhai X, Liu J, Wen J et al. A genetic variant in long non-coding RNA HULC contributes to risk of HBV-related hepatocellular carcinoma in a Chinese population. PloS one 2012; 7: e35145.
Kallioniemi A, Kallioniemi OP, Sudar D, Rutovitz D, Gray JW, Waldman F et al. Comparative genomic hybridization for molecular cytogenetic analysis of solid tumors. Science 1992; 258: 818–821.
Beroukhim R, Mermel CH, Porter D, Wei G, Raychaudhuri S, Donovan J et al. The landscape of somatic copy-number alteration across human cancers. Nature 2010; 463: 899–905.
Sur I, Tuupanen S, Whitington T, Aaltonen LA, Taipale J . Lessons from functional analysis of genome-wide association studies. Cancer Res 2013; 73: 4180–4184.
Huppi K, Pitt JJ, Wahlberg BM, Caplen NJ . The 8q24 gene desert: an oasis of non-coding transcriptional activity. Front Genet 2012; 3: 69.
Jia L, Landan G, Pomerantz M, Jaschek R, Herman P, Reich D et al. Functional enhancers at the gene-poor 8q24 cancer-linked locus. PLoS Genet 2009; 5: e1000597.
Xiang JF, Yin QF, Chen T, Zhang Y, Zhang XO, Wu Z et al. Human colorectal cancer-specific CCAT1-L lncRNA regulates long-range chromatin interactions at the MYC locus. Cell Res 2014; 24: 513–531.
Kim T, Cui R, Jeon YJ, Lee JH, Lee JH, Sim H et al. Long-range interaction and correlation between MYC enhancer and oncogenic long noncoding RNA CARLo-5. Proc Natl Acad Sci USA 2014; 111: 4173–4178.
Tseng YY, Moriarity BS, Gong W, Akiyama R, Tiwari A, Kawakami H et al. PVT1 dependence in cancer with MYC copy-number increase. Nature 2014; 512: 82–86.
Prensner JR, Iyer MK, Balbin OA, Dhanasekaran SM, Cao Q, Brenner JC et al. Transcriptome sequencing across a prostate cancer cohort identifies PCAT-1, an unannotated lincRNA implicated in disease progression. Nat Biotechnol 2011; 29: 742–749.
Li L, Sun R, Liang Y, Pan X, Li Z, Bai P et al. Association between polymorphisms in long non-coding RNA PRNCR1 in 8q24 and risk of colorectal cancer. J Exp Clin Cancer Res 2013; 32: 104.
Gaudet MM, Kirchhoff T, Green T, Vijai J, Korn JM, Guiducci C et al. Common genetic variants and modification of penetrance of BRCA2-associated breast cancer. PLoS Genet 2010; 6: e1001183.
Eeles RA, Kote-Jarai Z, Giles GG, Olama AA, Guy M, Jugurnauth SK et al. Multiple newly identified loci associated with prostate cancer susceptibility. Nat Genet 2008; 40: 316–321.
Chung S, Nakagawa H, Uemura M, Piao L, Ashikawa K, Hosono N et al. Association of a novel long non-coding RNA in 8q24 with prostate cancer susceptibility. Cancer Sci 2011; 102: 245–252.
Tuupanen S, Turunen M, Lehtonen R, Hallikas O, Vanharanta S, Kivioja T et al. The common colorectal cancer predisposition SNP rs6983267 at chromosome 8q24 confers potential to enhanced Wnt signaling. Nat Genet 2009; 41: 885–890.
Pomerantz MM, Ahmadiyeh N, Jia L, Herman P, Verzi MP, Doddapaneni H et al. The 8q24 cancer risk variant rs6983267 shows long-range interaction with MYC in colorectal cancer. Nat Genet 2009; 41: 882–884.
Ghoussaini M, Song H, Koessler T, Al Olama AA, Kote-Jarai Z, Driver KE et al. Multiple loci with different cancer specificities within the 8q24 gene desert. J Natl Cancer Inst 2008; 100: 962–966.
Bertucci F, Lagarde A, Ferrari A, Finetti P, Charafe-Jauffret E, Van Laere S et al. 8q24 Cancer risk allele associated with major metastatic risk in inflammatory breast cancer. PloS one 2012; 7: e37943.
Sur IK, Hallikas O, Vaharautio A, Yan J, Turunen M, Enge M et al. Mice lacking a Myc enhancer that includes human SNP rs6983267 are resistant to intestinal tumors. Science 2012; 338: 1360–1363.
Cancer Genome Atlas N. Comprehensive molecular characterization of human colon and rectal cancer. Nature 2012; 487: 330–337.
Hnisz D, Abraham BJ, Lee TI, Lau A, Saint-Andre V, Sigova AA et al. Super-enhancers in the control of cell identity and disease. Cell 2013; 155: 934–947.
Kumar V, Westra HJ, Karjalainen J, Zhernakova DV, Esko T, Hrdlickova B et al. Human disease-associated genetic variation impacts large intergenic non-coding RNA expression. PLoS Genet 2013; 9: e1003201.
Prensner JR, Chen W, Iyer MK, Cao Q, Ma T, Han S et al. PCAT-1, a long noncoding RNA, regulates BRCA2 and controls homologous recombination in cancer. Cancer Res 2014; 74: 1651–1660.
Gupta RA, Shah N, Wang KC, Kim J, Horlings HM, Wong DJ et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 2010; 464: 1071–1076.
Gutschner T, Diederichs S . The hallmarks of cancer: a long non-coding RNA point of view. RNA Biol 2012; 9: 703–719.
Yildirim E, Kirby JE, Brown DE, Mercier FE, Sadreyev RI, Scadden DT et al. Xist RNA is a potent suppressor of hematologic cancer in mice. Cell 2013; 152: 727–742.
Ji P, Diederichs S, Wang W, Boing S, Metzger R, Schneider PM et al. MALAT-1, a novel noncoding RNA, and thymosin beta4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene 2003; 22: 8031–8041.
Yang Z, Zhou L, Wu LM, Lai MC, Xie HY, Zhang F et al. Overexpression of long non-coding RNA HOTAIR predicts tumor recurrence in hepatocellular carcinoma patients following liver transplantation. Ann Surgical Oncol 2011; 18: 1243–1250.
Kogo R, Shimamura T, Mimori K, Kawahara K, Imoto S, Sudo T et al. Long noncoding RNA HOTAIR regulates polycomb-dependent chromatin modification and is associated with poor prognosis in colorectal cancers. Cancer Res 2011; 71: 6320–6326.
Niinuma T, Suzuki H, Nojima M, Nosho K, Yamamoto H, Takamaru H et al. Upregulation of miR-196a and HOTAIR drive malignant character in gastrointestinal stromal tumors. Cancer Res 2012; 72: 1126–1136.
Kim K, Jutooru I, Chadalapaka G, Johnson G, Frank J, Burghardt R et al. HOTAIR is a negative prognostic factor and exhibits pro-oncogenic activity in pancreatic cancer. Oncogene 2013; 32: 1616–1625.
Redis RS, Sieuwerts AM, Look MP, Tudoran O, Ivan C, Spizzo R et al. CCAT2, a novel long non-coding RNA in breast cancer: expression study and clinical correlations. Oncotarget 2013; 4: 1748–1762.
Du Z, Fei T, Verhaak RG, Su Z, Zhang Y, Brown M et al. Integrative genomic analyses reveal clinically relevant long noncoding RNAs in human cancer. Nat Struct Mol Biol 2013; 20: 908–913.
Ren S, Wang F, Shen J, Sun Y, Xu W, Lu J et al. Long non-coding RNA metastasis associated in lung adenocarcinoma transcript 1 derived miniRNA as a novel plasma-based biomarker for diagnosing prostate cancer. Eur J Cancer 2013; 49: 2949–2959.
Rittenhouse H, Blase A, Shamel B, Schalken J, Groskopf J . The long and winding road to FDA approval of a novel prostate cancer test: our story. Clin Chem 2013; 59: 32–34.
Wojcik SE, Rossi S, Shimizu M, Nicoloso MS, Cimmino A, Alder H et al. Non-codingRNA sequence variations in human chronic lymphocytic leukemia and colorectal cancer. Carcinogenesis 2010; 31: 208–215.
Hafner M, Landthaler M, Burger L, Khorshid M, Hausser J, Berninger P et al. Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell 2010; 141: 129–141.
Martin L, Meier M, Lyons SM, Sit RV, Marzluff WF, Quake SR et al. Systematic reconstruction of RNA functional motifs with high-throughput microfluidics. Nat Methods 2012; 9: 1192–1194.
Paige JS, Wu KY, Jaffrey SR . RNA mimics of green fluorescent protein. Science 2011; 333: 642–646.
Srikantan V, Zou Z, Petrovics G, Xu L, Augustus M, Davis L et al. PCGEM1, a prostate-specific gene, is overexpressed in prostate cancer. Proc Natl Acad Sci USA 2000; 97: 12216–12221.
Fu X, Ravindranath L, Tran N, Petrovics G, Srivastava S . Regulation of apoptosis by a prostate-specific and prostate cancer-associated noncoding gene, PCGEM1. DNA Cell Biol 2006; 25: 135–141.
Xue Y, Wang M, Kang M, Wang Q, Wu B, Chu H et al. Association between lncrna PCGEM1 polymorphisms and prostate cancer risk. Prost Cancer Prost Dis 2013; 16: 139–144, S131.
Du Y, Kong G, You X, Zhang S, Zhang T, Gao Y et al. Elevation of highly up-regulated in liver cancer (HULC) by hepatitis B virus X protein promotes hepatoma cell proliferation via down-regulating p18. J Biol Chem 2012; 287: 26302–26311.
Tomlinson I, Webb E, Carvajal-Carmona L, Broderick P, Kemp Z, Spain S et al. A genome-wide association scan of tag SNPs identifies a susceptibility variant for colorectal cancer at 8q24.21. Nat Genet 2007; 39: 984–988.
Yeager M, Orr N, Hayes RB, Jacobs KB, Kraft P, Wacholder S et al. Genome-wide association study of prostate cancer identifies a second risk locus at 8q24. Nat Genet 2007; 39: 645–649.
Chen D, Zhang Z, Mao C, Zhou Y, Yu L, Yin Y et al. ANRIL inhibits p15(INK4b) through the TGFbeta1 signaling pathway in human esophageal squamous cell carcinoma. Cell Immunol 2014; 289: 91–96.
Gutschner T, Hammerle M, Eissmann M, Hsu J, Kim Y, Hung G et al. The noncoding RNA MALAT1 is a critical regulator of the metastasis phenotype of lung cancer cells. Cancer Res 2013; 73: 1180–1189.
Acknowledgements
HL is an Odyssey Fellow, and his work is supported in part by the Odyssey Program at The University of Texas MD Anderson Cancer Center. GAC is The Alan M. Gewirtz Leukemia & Lymphoma Society Scholar. Work in Dr Calin's laboratory is supported in part by the NIH/NCI grants 1UH2TR00943-01 and 1 R01 CA182905-01, the UT MD Anderson Cancer Center SPORE in Melanoma grant from NCI (P50 CA093459), Aim at Melanoma Foundation and the Miriam and Jim Mulva research funds, the Brain SPORE (2P50CA127001), the Center for radiation Oncology Research Project, the Center for Cancer Epigenetics Pilot project, a 2014 Knowledge GAP MDACC grant, a CLL Moonshot pilot project, the UT MD Anderson Cancer Center Duncan Family Institute for Cancer Prevention and Risk Assessment, a SINF grant in colon cancer, the Laura and John Arnold Foundation, the RGK Foundation and the Estate of C. G. Johnson, Jr. MP is supported by an Erwin-Schroedinger Scholarship of the Austrian Science Funds (project no. J3389-B23). IBN is a Fulbright Scholar at MD Anderson Cancer Center. FS was supported by NIH grants CA131301 and CA157749. We apologize to all colleagues whose work was not cited because of space restrictions.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Ling, H., Vincent, K., Pichler, M. et al. Junk DNA and the long non-coding RNA twist in cancer genetics. Oncogene 34, 5003–5011 (2015). https://doi.org/10.1038/onc.2014.456
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2014.456
This article is cited by
-
Amino acid metabolism regulated by lncRNAs: the propellant behind cancer metabolic reprogramming
Cell Communication and Signaling (2023)
-
Estimating residual undifferentiated cells in human chemically induced pluripotent stem cell derived islets using lncRNA as biomarkers
Scientific Reports (2023)
-
Long Non-coding RNA UCA1a Promotes Proliferation via PKM2 in Cervical Cancer
Reproductive Sciences (2023)
-
Establishment and validation of a prognostic model based on HRR-related lncRNAs in colon adenocarcinoma
World Journal of Surgical Oncology (2022)
-
Prognostic risk assessment model and drug sensitivity analysis of colon adenocarcinoma (COAD) based on immune-related lncRNA pairs
BMC Bioinformatics (2022)