Abstract
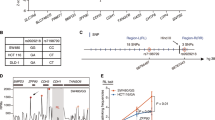
Common genetic variation at human 14q22.2 tagged by rs4444235 is significantly associated with colorectal cancer (CRC) risk. Re-sequencing was used to comprehensively annotate the 17kb region of strong linkage disequilibrium encompassing rs4444235. Through bioinformatic analyses using H3K4Me1, H3K4Me3, and DNase-I hypersensitivity chromatin signatures and evolutionary conservation we identified seven candidate disease-causing single-nucleotide polymorphisms mapping to six regions within the 17-kb region predicted to have regulatory potential. Reporter gene studies of these regions demonstrated that the element to which rs4444235 maps acts as an allele-specific transcriptional enhancer. Allele-specific expression studies in CRC cell lines heterozygous for rs4444235 showed significantly increased expression of bone morphogenetic protein-4 (BMP4) associated with the risk allele (P<0.001). These data provide evidence for a functional basis for the non-coding risk variant rs4444235 at 14q22.2 and emphasizes the importance of genetic variation in the BMP pathway genes as determinants of CRC risk.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Aaltonen L, Johns L, Jarvinen H, Mecklin JP, Houlston R . (2007). Explaining the familial colorectal cancer risk associated with mismatch repair (MMR)-deficient and MMR-stable tumors. Clin Cancer Res 13: 356–361.
Abdel-Rahman WM, Katsura K, Rens W, Gorman PA, Sheer D, Bicknell D et al. (2001). Spectral karyotyping suggests additional subsets of colorectal cancers characterized by pattern of chromosome rearrangement. Proc Natl Acad Sci USA 98: 2538–2543.
Broderick P, Carvajal-Carmona L, Pittman AM, Webb E, Howarth K, Rowan A et al. (2007). A genome-wide association study shows that common alleles of SMAD7 influence colorectal cancer risk. Nat Genet 39: 1315–1317.
Deng H, Makizumi R, Ravikumar TS, Dong H, Yang W, Yang WL . (2007). Bone morphogenetic protein-4 is overexpressed in colonic adenocarcinomas and promotes migration and invasion of HCT116 cells. Exp Cell Res 313: 1033–1044.
Eisen T, Matakidou A, Houlston R . (2008). Identification of low penetrance alleles for lung cancer: the GEnetic Lung CAncer Predisposition Study (GELCAPS). BMC Cancer 8: 244.
Fried M, Crothers DM . (1981). Equilibria and kinetics of lac repressor–operator interactions by polyacrylamide gel electrophoresis. Nucleic Acids Res 9: 6505–6525.
Gaasenbeek M, Howarth K, Rowan AJ, Gorman PA, Jones A, Chaplin T et al. (2006). Combined array–comparative genomic hybridization and single-nucleotide polymorphism–loss of heterozygosity analysis reveals complex changes and multiple forms of chromosomal instability in colorectal cancers. Cancer Res 66: 3471–3479.
Garner MM, Revzin A . (1981). A gel electrophoresis method for quantifying the binding of proteins to specific DNA regions: application to components of the Escherichia coli lactose operon regulatory system. Nucleic Acids Res 9: 3047–3060.
Gomez-Skarmeta JL, Lenhard B, Becker TS . (2006). New technologies, new findings, and new concepts in the study of vertebrate cis-regulatory sequences. Dev Dyn 235: 870–885.
He XC, Zhang J, Tong WG, Tawfik O, Ross J, Scoville DH et al. (2004). BMP signaling inhibits intestinal stem cell self-renewal through suppression of Wnt–beta-catenin signaling. Nat Genet 36: 1117–1121.
Heinemeyer T, Wingender E, Reuter I, Hermjakob H, Kel AE, Kel OV et al. (1998). Databases on transcriptional regulation: TRANSFAC, TRRD and COMPEL. Nucleic Acids Res 26: 362–367.
Houlston RS, Cheadle J, Dobbins SE, Tenesa A, Jones AM, Howarth K et al. (2010). Meta-analysis of three genome-wide association studies identifies susceptibility loci for colorectal cancer at 1q41, 3q26.2, 12q13.13 and 20q13.33. Nat Genet 42: 973–977.
Houlston RS, Webb E, Broderick P, Pittman AM, Di Bernardo MC, Lubbe S et al. (2008). Meta-analysis of genome-wide association data identifies four new susceptibility loci for colorectal cancer. Nat Genet 40: 1426–1435.
Jaeger E, Webb E, Howarth K, Carvajal-Carmona L, Rowan A, Broderick P et al. (2008). Common genetic variants at the CRAC1 (HMPS) locus on chromosome 15q13.3 influence colorectal cancer risk. Nat Genet 40: 26–28.
Kawai K, Viars C, Arden K, Tarin D, Urquidi V, Goodison S . (2002). Comprehensive karyotyping of the HT-29 colon adenocarcinoma cell line. Genes Chromosomes Cancer 34: 1–8.
Kosinski C, Li VS, Chan AS, Zhang J, Ho C, Tsui WY et al. (2007). Gene expression patterns of human colon tops and basal crypts and BMP antagonists as intestinal stem cell niche factors. Proc Natl Acad Sci USA 104: 15418–15423.
Lichtenstein P, Holm NV, Verkasalo PK, Iliadou A, Kaprio J, Koskenvuo M et al. (2000). Environmental and heritable factors in the causation of cancer—analyses of cohorts of twins from Sweden, Denmark, and Finland. N Engl J Med 343: 78–85.
Liu C, Calogero A, Ragona G, Adamson E, Mercola D . (1996). EGR-1, the reluctant suppression factor: EGR-1 is known to function in the regulation of growth, differentiation, and also has significant tumor suppressor activity and a mechanism involving the induction of TGF-beta1 is postulated to account for this suppressor activity. Crit Rev Oncog 7: 101–125.
Lo HS, Wang Z, Hu Y, Yang HH, Gere S, Buetow KH et al. (2003). Allelic variation in gene expression is common in the human genome. Genome Res 13: 1855–1862.
Lubbe SJ, Webb EL, Chandler IP, Houlston RS . (2009). Implications of familial colorectal cancer risk profiles and microsatellite instability status. J Clin Oncol 27: 2238–2244.
Milani L, Gupta M, Andersen M, Dhar S, Fryknas M, Isaksson A et al. (2007). Allelic imbalance in gene expression as a guide to cis-acting regulatory single nucleotide polymorphisms in cancer cells. Nucleic Acids Res 35: e34.
Palin K, Taipale J, Ukkonen E . (2006). Locating potential enhancer elements by comparative genomics using the EEL software. Nat Protoc 1: 368–374.
Penegar S, Wood W, Lubbe S, Chandler I, Broderick P, Papaemmanuil E et al. (2007). National study of colorectal cancer genetics. Br J Cancer 97: 1305–1309.
Pittman AM, Naranjo S, Jalava SE, Twiss P, Ma Y, Olver B et al. (2010). Allelic variation at the 8q23.3 colorectal cancer risk locus functions as a cis-acting regulator of EIF3H. PLoS Genet 6: e1001126.
Pittman AM, Naranjo S, Webb E, Broderick P, Lips EH, van Wezel T et al. (2009). The colorectal cancer risk at 18q21 is caused by a novel variant altering SMAD7 expression. Genome Res 19: 987–993.
Polakis P . (2000). Wnt signaling and cancer. Genes Dev 14: 1837–1851.
Pomerantz MM, Ahmadiyeh N, Jia L, Herman P, Verzi MP, Doddapaneni H et al. (2009). The 8q24 cancer risk variant rs6983267 shows long-range interaction with MYC in colorectal cancer. Nat Genet 41: 882–884.
Pregizer S, Mortlock DP . (2009). Control of BMP gene expression by long-range regulatory elements. Cytokine Growth Factor Rev 20: 509–515.
Takash W, Canizares J, Bonneaud N, Poulat F, Mattei MG, Jay P et al. (2001). SOX7 transcription factor: sequence, chromosomal localisation, expression, transactivation and interference with Wnt signalling. Nucleic Acids Res 29: 4274–4283.
Tenesa A, Farrington SM, Prendergast JG, Porteous ME, Walker M, Haq N et al. (2008). Genome-wide association scan identifies a colorectal cancer susceptibility locus on 11q23 and replicates risk loci at 8q24 and 18q21. Nat Genet 40: 631–637.
Tomlinson I, Carvajal-Carmona L, Dobbins SE, Tenesa A, Jones A, Howarth K et al. (2011). Multiple common susceptibility variants near BMP pathway loci GREM1, BMP4 and BMP2 explain part of the missing heritability of colorectal cancer. PLoS Genet 7: e1002105.
Tomlinson I, Webb E, Carvajal-Carmona L, Broderick P, Kemp Z, Spain S et al. (2007). A genome-wide association scan of tag SNPs identifies a susceptibility variant for colorectal cancer at 8q24.21. Nat Genet 39: 984–988.
Tomlinson IP, Webb E, Carvajal-Carmona L, Broderick P, Howarth K, Pittman AM et al. (2008). A genome-wide association study identifies colorectal cancer susceptibility loci on chromosomes 10p14 and 8q23.3. Nat Genet 40: 623–630.
Tsushimi T, Noshima S, Oga A, Esato K, Sasaki K . (2001). DNA amplification and chromosomal translocations are accompanied by chromosomal instability: analysis of seven human colon cancer cell lines by comparative genomic hybridization and spectral karyotyping. Cancer Genet Cytogenet 126: 34–38.
Tuupanen S, Turunen M, Lehtonen R, Hallikas O, Vanharanta S, Kivioja T et al. (2009). The common colorectal cancer predisposition SNP rs6983267 at chromosome 8q24 confers potential to enhanced Wnt signaling. Nat Genet 41: 885–890.
Zanke BW, Greenwood CM, Rangrej J, Kustra R, Tenesa A, Farrington SM et al. (2007). Genome-wide association scan identifies a colorectal cancer susceptibility locus on chromosome 8q24. Nat Genet 39: 989–994.
Acknowledgements
We acknowledge the National Health Service (NHS) funding to the National Institute for Health Research (NIHR) Biomedical Research Centre. Finally, we are grateful to all patients and individuals for participation. Cancer Research UK (C1298/A8362 supported by the Bobby Moore Fund) provided principal funding for the study. JLG-S acknowledges grants from the Spanish Ministry of Education and Science (BFU2010-14839 and CSD2007-00008) and Junta de Andalucía (CVI-3488). SJL is in receipt of a PhD studentship from Cancer Research UK. The research leading to these results has received funding from the European Union Seventh Framework Programme (FP7/207-2013) under grant no. 258236, FP7 collaborative project SYSCOL.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Lubbe, S., Pittman, A., Olver, B. et al. The 14q22.2 colorectal cancer variant rs4444235 shows cis-acting regulation of BMP4. Oncogene 31, 3777–3784 (2012). https://doi.org/10.1038/onc.2011.564
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2011.564
Keywords
This article is cited by
-
Promoter/enhancer-based controllability of regulatory networks
Scientific Reports (2022)
-
Evaluation of gene-environment interactions for colorectal cancer susceptibility loci using case-only and case-control designs
BMC Cancer (2019)
-
Inhibition of bone morphogenetic protein signaling reduces viability, growth and migratory potential of non-small cell lung carcinoma cells
Journal of Cancer Research and Clinical Oncology (2019)
-
A subset of genetic susceptibility variants for colorectal cancer also has prognostic value
The Pharmacogenomics Journal (2016)