Abstract
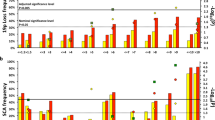
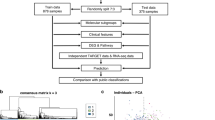
Imbalances in chromosome 11q occur in approximately 30% of primary neuroblastoma and are associated with poor outcome. It has been suggested that 11q loss constitutes a distinct clinico-genetic neuroblastoma subgroup by affecting expression levels of corresponding genes. This study analysed the relationship of 11q loss, clinical phenotype and global transcriptomic profiles in four clinico-genetic subgroups (11q alteration/favourable outcome, n=7; 11q alteration/unfavourable outcome, n=14; no 11q alteration/favourable outcome, n=81; no 11q alteration/unfavourable outcome, n=8; tumours with MYCN amplification and/or 1p loss were excluded). Unsupervised and supervised comparisons of gene expression profiles consistently showed significantly different mRNA patterns between favourable and unfavourable neuroblastomas, both in the subgroups with and without 11q loss. In contrast, favourable tumours with and without 11q loss showed highly similar transcriptomic profiles. Disproportionate downregulation of 11q genes was observed only in unfavourable tumours with 11q loss. The diverging molecular profiles were neither caused by considerable differences in the size of the deleted regions nor by differential methylation patterns of 11q genes. Together, this study shows that neuroblastoma with 11q loss comprises two biological subgroups that differ both in their clinical phenotype and gene expression patterns, indicating that 11q loss is not a primary determinant of neuroblastoma tumour behaviour.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Alaminos M, Mora J, Cheung NK, Smith A, Qin J, Chen L et al. (2003). Genome-wide analysis of gene expression associated with MYCN in human neuroblastoma. Cancer Res 63: 4538–4546.
Ambros PF, Ambros IM . (2001). Pathology and biology guidelines for resectable and unresectable neuroblastic tumors and bone marrow examination guidelines. Med Pediatr Oncol 37: 492–504.
Asgharzadeh S, Pique-Regi R, Sposto R, Wang H, Yang Y, Shimada H et al. (2006). Prognostic significance of gene expression profiles of metastatic neuroblastomas lacking MYCN gene amplification. J Natl Cancer Inst 98: 1193–1203.
Attiyeh EF, London WB, Mosse YP, Wang Q, Winter C, Khazi D et al. (2005). Chromosome 1p and 11q deletions and outcome in neuroblastoma. N Engl J Med 353: 2243–2253.
Berwanger B, Hartmann O, Bergmann E, Bernard S, Nielsen D, Krause M et al. (2002). Loss of a FYN-regulated differentiation and growth arrest pathway in advanced stage neuroblastoma. Cancer Cell 2: 377–386.
Bilke S, Chen QR, Westerman F, Schwab M, Catchpoole D, Khan J . (2005). Inferring a tumor progression model for neuroblastoma from genomic data. J Clin Oncol 23: 7322–7331.
Boon K, Caron HN, van Asperen R, Valentijn L, Hermus MC, van Sluis P et al. (2001). N-myc enhances the expression of a large set of genes functioning in ribosome biogenesis and protein synthesis. EMBO J 20: 1383–1393.
Brodeur GM . (2003). Neuroblastoma: biological insights into a clinical enigma. Nat Rev Cancer 3: 203–216.
Chen W, Salto-Tellez M, Palanisamy N, Ganesan K, Hou Q, Tan LK et al. (2007). Targets of genome copy number reduction in primary breast cancers identified by integrative genomics. Genes Chromosomes Cancer 46: 288–301.
Classen S, Zander T, Eggle D, Chemnitz JM, Brors B, Buchmann I et al. (2007). Human resting CD4+ T cells are constitutively inhibited by TGF beta under steady-state conditions. J Immunol 178: 6931–6940.
Cohn SL, Pearson AD, London WB, Monclair T, Ambros PF, Brodeur GM et al. (2009). The International Neuroblastoma Risk Group (INRG) classification system: an INRG Task Force report. J Clin Oncol 27: 289–297.
Ehrich M, Nelson MR, Stanssens P, Zabeau M, Liloglou T, Xinarianos G et al. (2005). Quantitative high-throughput analysis of DNA methylation patterns by base-specific cleavage and mass spectrometry. Proc Natl Acad Sci USA 102: 15785–15790.
Fischer M, Oberthuer A, Brors B, Kahlert Y, Skowron M, Voth H et al. (2006). Differential expression of neuronal genes defines subtypes of disseminated neuroblastoma with favorable and unfavorable outcome. Clin Cancer Res 12: 5118–5128.
Fischer M, Spitz R, Oberthur A, Westermann F, Berthold F . (2008). Risk estimation of neuroblastoma patients using molecular markers. Klin Padiatr 220: 137–146.
Gallegos Ruiz MI, Floor K, Roepman P, Rodriguez JA, Meijer GA, Mooi WJ et al. (2008). Integration of gene dosage and gene expression in non-small cell lung cancer, identification of HSP90 as potential target. PLoS ONE 3: e0001722.
Henrich KO, Fischer M, Mertens D, Benner A, Wiedemeyer R, Brors B et al. (2006). Reduced expression of CAMTA1 correlates with adverse outcome in neuroblastoma patients. Clin Cancer Res 12: 131–138.
Huang J, Sheng HH, Shen T, Hu YJ, Xiao HS, Zhang Q et al. (2006). Correlation between genomic DNA copy number alterations and transcriptional expression in hepatitis B virus-associated hepatocellular carcinoma. FEBS Lett 580: 3571–3581.
Janoueix-Lerosey I, Schleiermacher G, Michels E, Mosseri V, Ribeiro A, Lequin D et al. (2009). Overall genomic pattern is a predictor of outcome in neuroblastoma. J Clin Oncol 27: 1026–1033.
Lastowska M, Viprey V, Santibanez-Koref M, Wappler I, Peters H, Cullinane C et al. (2007). Identification of candidate genes involved in neuroblastoma progression by combining genomic and expression microarrays with survival data. Oncogene 26: 7432–7444.
Maris JM, Hogarty MD, Bagatell R, Cohn SL . (2007). Neuroblastoma. Lancet 369: 2106–2120.
McArdle L, McDermott M, Purcell R, Grehan D, O’Meara A, Breatnach F et al. (2004). Oligonucleotide microarray analysis of gene expression in neuroblastoma displaying loss of chromosome 11q. Carcinogenesis 25: 1599–1609.
Mosse YP, Diskin SJ, Wasserman N, Rinaldi K, Attiyeh EF, Cole K et al. (2007). Neuroblastomas have distinct genomic DNA profiles that predict clinical phenotype and regional gene expression. Genes Chromosomes Cancer 46: 936–949.
Mosse YP, Laudenslager M, Longo L, Cole KA, Wood A, Attiyeh EF et al. (2008). Identification of ALK as a major familial neuroblastoma predisposition gene. Nature 455: 930–935.
Nigro JM, Misra A, Zhang L, Smirnov I, Colman H, Griffin C et al. (2005). Integrated array-comparative genomic hybridization and expression array profiles identify clinically relevant molecular subtypes of glioblastoma. Cancer Res 65: 1678–1686.
Oberthuer A, Berthold F, Warnat P, Hero B, Kahlert Y, Spitz R et al. (2006). Customized oligonucleotide microarray gene expression-based classification of neuroblastoma patients outperforms current clinical risk stratification. J Clin Oncol 24: 5070–5078.
Oberthuer A, Kaderali L, Kahlert Y, Hero B, Westermann F, Berthold F et al. (2008). Subclassification and individual survival time prediction from gene expression data of neuroblastoma patients by using CASPAR. Clin Cancer Res 14: 6590–6601.
Ohira M, Oba S, Nakamura Y, Isogai E, Kaneko S, Nakagawa A et al. (2005). Expression profiling using a tumor-specific cDNA microarray predicts the prognosis of intermediate risk neuroblastomas. Cancer Cell 7: 337–350.
Platzer P, Upender MB, Wilson K, Willis J, Lutterbaugh J, Nosrati A et al. (2002). Silence of chromosomal amplifications in colon cancer. Cancer Res 62: 1134–1138.
Pollack JR, Sorlie T, Perou CM, Rees CA, Jeffrey SS, Lonning PE et al. (2002). Microarray analysis reveals a major direct role of DNA copy number alteration in the transcriptional program of human breast tumors. Proc Natl Acad Sci USA 99: 12963–12968.
Potter N, Karakoula A, Phipps KP, Harkness W, Hayward R, Thompson DN et al. (2008). Genomic deletions correlate with underexpression of novel candidate genes at six loci in pediatric pilocytic astrocytoma. Neoplasia 10: 757–772.
Savelyeva L, Schwab M . (2001). Amplification of oncogenes revisited: from expression profiling to clinical application. Cancer Lett 167: 115–123.
Spitz R, Hero B, Ernestus K, Berthold F . (2003). Deletions in chromosome arms 3p and 11q are new prognostic markers in localized and 4s neuroblastoma. Clin Cancer Res 9: 52–58.
Spitz R, Hero B, Simon T, Berthold F . (2006a). Loss in chromosome 11q identifies tumors with increased risk for metastatic relapses in localized and 4S neuroblastoma. Clin Cancer Res 12: 3368–3373.
Spitz R, Oberthuer A, Zapatka M, Brors B, Hero B, Ernestus K et al. (2006b). Oligonucleotide array-based comparative genomic hybridization (aCGH) of 90 neuroblastomas reveals aberration patterns closely associated with relapse pattern and outcome. Genes Chromosomes Cancer 45: 1130–1142.
Stallings RL . (2007). Origin and functional significance of large-scale chromosomal imbalances in neuroblastoma. Cytogenet Genome Res 118: 110–115.
Stallings RL, Nair P, Maris JM, Catchpoole D, McDermott M, O’Meara A et al. (2006). High-resolution analysis of chromosomal breakpoints and genomic instability identifies PTPRD as a candidate tumor suppressor gene in neuroblastoma. Cancer Res 66: 3673–3680.
Tusher VG, Tibshirani R, Chu G . (2001). Significance analysis of microarrays applied to the ionizing radiation response. Proc Natl Acad Sci USA 98: 5116–5121.
Wang Q, Diskin S, Rappaport E, Attiyeh E, Mosse Y, Shue D et al. (2006). Integrative genomics identifies distinct molecular classes of neuroblastoma and shows that multiple genes are targeted by regional alterations in DNA copy number. Cancer Res 66: 6050–6062.
Westermann F, Muth D, Benner A, Bauer T, Henrich KO, Oberthuer A et al. (2008). Distinct transcriptional MYCN/c-MYC activities are associated with spontaneous regression or malignant progression in neuroblastomas. Genome Biol 9: R150.
Yoshimoto T, Matsuura K, Karnan S, Tagawa H, Nakada C, Tanigawa M et al. (2007). High-resolution analysis of DNA copy number alterations and gene expression in renal clear cell carcinoma. J Pathol 213: 392–401.
Acknowledgements
We are grateful to Yvonne Kahlert for excellent technical assistance and to Dr Roman Thomas for critical reading of the paper. This work was supported by grants from the Deutsche Krebshilfe (Grant 50-2719), the Bundesministerium für Bildung und Forschung (BMBF) through the National Genome Research Network 2 (NGFN2, Grants 01GS0456 and 01GR0450) and the Competence Network Paediatric Oncology and Hematology (KPOH) as well as the Fördergesellschaft Kinderkrebs-Neuroblastom-Forschung e.V.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Oncogene website (http://www.nature.com/onc)
Supplementary information
Rights and permissions
About this article
Cite this article
Fischer, M., Bauer, T., Oberthür, A. et al. Integrated genomic profiling identifies two distinct molecular subtypes with divergent outcome in neuroblastoma with loss of chromosome 11q. Oncogene 29, 865–875 (2010). https://doi.org/10.1038/onc.2009.390
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2009.390
Keywords
This article is cited by
-
Investigation of major genetic alterations in neuroblastoma
Molecular Biology Reports (2018)
-
MYCN amplification predicts poor prognosis based on interphase fluorescence in situ hybridization analysis of bone marrow cells in bone marrow metastases of neuroblastoma
Cancer Cell International (2017)
-
11q deletion in neuroblastoma: a review of biological and clinical implications
Molecular Cancer (2017)
-
Influence of segmental chromosome abnormalities on survival in children over the age of 12 months with unresectable localised peripheral neuroblastic tumours without MYCN amplification
British Journal of Cancer (2015)
-
FOXP1inhibits cell growth and attenuates tumorigenicity of neuroblastoma
BMC Cancer (2014)