Abstract
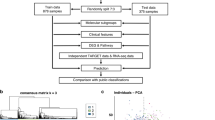
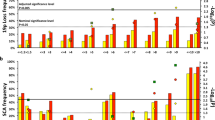
Human neuroblastoma remains enigmatic because it often shows spontaneous regression and aggressive growth. The prognosis of advanced stage of sporadic neuroblastomas is still poor. Here, we investigated whether genomic and molecular signatures could categorize new therapeutic risk groups in primary neuroblastomas. We conducted microarray-based comparative genomic hybridization (array-CGH) with a DNA chip carrying 2464 BAC clones to examine genomic aberrations of 236 neuroblastomas and used in-house cDNA microarrays for gene-expression profiling. Array-CGH demonstrated three major genomic groups of chromosomal aberrations: silent (GGS), partial gains and/or losses (GGP) and whole gains and/or losses (GGW), which well corresponded with the patterns of chromosome 17 abnormalities. They were further classified into subgroups with different outcomes. In 112 sporadic neuroblastomas, MYCN amplification was frequent in GGS (22%) and GGP (53%) and caused serious outcomes in patients. Sporadic tumors with a single copy of MYCN showed the 5-year cumulative survival rates of 89% in GGS, 53% in GGP and 85% in GGW. Molecular signatures also segregated patients into the favorable and unfavorable prognosis groups (P=0.001). Both univariate and multivariate analyses revealed that genomic and molecular signatures were mutually independent, powerful prognostic indicators. Thus, combined genomic and molecular signatures may categorize novel risk groups and confer new clues for allowing tailored or even individualized medicine to patients with neuroblastoma.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Albertson DG, Collins C, McCormick F, Gray JW . (2003). Chromosome aberrations in solid tumors. Nat Genet 34: 369–376.
Attiyeh EF, London WB, Mosse YP, Wang Q, Winter C, Khazi D et al. (2005). Chromosome 1p and 11q deletions and outcome in neuroblastoma. N Engl J Med 353: 2243–2253.
Bagchi A, Papazoglu C, Wu Y, Capurso D, Brodt M, Francis D et al. (2007). CHD5 is a tumor suppressor at human 1p36. Cell 128: 459–475.
Beckwith JB, Perrin EV . (1963). In situ neuroblastomas: a contribution to the natural history of neural crest tumors. Am J Pathol 43: 1089–1101.
Brodeur GM . (2003). Neuroblastoma: biological insight into a clinical enigma. Nat Rev Cancer 3: 203–216.
Brodeur GM, Look AT, Shimada H, Hamilton VM, Maris JM, Hann HW et al. (2001). Biological aspects of neuroblastomas identified by mass screening in Quebec. Med Pediatr Oncol 36: 157–159.
Brodeur GM, Pritchard J, Berthold F, Carlsen NL, Castel V, Castelberry RP et al. (1993). Revisions of the international criteria for neuroblastoma diagnosis, staging, and response to treatment. J Clin Oncol 11: 1466–1477.
Iehara T, Hosoi H, Akazawa K, Matsumoto Y, Yamamoto K, Suita S et al. (2006). MYCN gene amplification is a powerful prognostic factor even in infantile neuroblastoma detected by mass screening. Br J Cancer 94: 1510–1515.
Islam A, Kageyama H, Takada N, Kawamoto T, Takayasu H, Isogai E et al. (2000). High expression of Survivin, mapped to 17q25, is significantly associated with poor prognostic factors and promotes cell survival in human neuroblastoma. Oncogene 19: 617–623.
Jain AN, Tokuyasu TA, Snijders AM, Segraves R, Albertson DG, Pinkel D . (2002). Fully automatic quantification of microarray image data. Genome Res 12: 325–332.
Kaneko M, Tsuchida Y, Mugishima H, Ohnuma N, Yamamoto K, Kawa K et al. (2002). Intensified chemotherapy increases the survival rates in patients with stage 4 neuroblastoma with MYCN amplification. J Pediatr Hematol Oncol 24: 613–621.
Levy IG . (2005). Neuroblastoma, well-designed evaluations, and the optimality of research funding: ask not what your country can do for you. J Natl Cancer Inst 97: 1105–1106.
Look AT, Hayes FA, Nitschke R, McWilliams NB, Green AA . (1984). Cellular DNA content as a predictor of response to chemotherapy in infants with unresectable neuroblastoma. N Engl J Med 311: 231–235.
Maris JM, Matthay KK . (1999). Molecular biology of neuroblastoma. J Clin Oncol 17: 2264–2279.
Nakagawara A . (1998). The NGF story and neuroblastoma. Med Pediatr Oncol 31: 113–115.
Nakagawara A . (2004). Neural crest development and neuroblastoma: the genetic and biological link. Prog Brain Res 146: 233–242.
Nakagawara A, Arima-Nakagawara M, Scavarda NJ, Azar CG, Cantor AB, Brodeur GM . (1993). Association between high levels of expression of the TRK gene and favorable outcome in human neuroblastoma. N Engl J Med 328: 847–854.
Oba S, Tomioka N, Ohira M, Ishii S . (2006). Combfit: a normalization method for array CGH data. IPSJ Trans Bioinformatics 47: 73–82.
Ohira M, Morohashi A, Inuzuka H, Shishikura T, Kawamoto T, Kageyama H et al. (2003). Expression profiling and characterization of 4200 genes cloned from primary neuroblastomas: identification of 305 genes differentially expressed between favorable and unfavorable subsets. Oncogene 22: 5525–5536.
Ohira M, Oba S, Nakamura Y, Isogai E, Kaneko S, Nakagawa A et al. (2005). Expression profiling using a tumor-specific cDNA microarray predicts the prognosis of intermediate risk neuroblastomas. Cancer Cell 7: 337–350.
Pinkel D, Segraves R, Sudar D, Clark S, Poole I, Kowbel D et al. (1998). High resolution analysis of DNA copy number variation using comparative genomic hybridization to microarrays. Nat Genet 20: 207–211.
Sawada T, Hirayama M, Nakata T, Takeda T, Takasugi N, Mori T et al. (1984). Mass screening for neuroblastoma in infants in Japan. Interim report of a mass screening study group. Lancet 2: 271–273.
Schwab M, Westermann F, Hero B, Berthold F . (2003). Neuroblastoma: biology and molecular and chromosomal pathology. Lancet 4: 472–480.
Snijders AM, Nowak N, Segraves R, Blackwood S, Brown N, Conroy J et al. (2001). Assembly of microarrays for genome-wide measurement of DNA copy number. Nat Genet 29: 263–264.
Srivatsan ES, Ying KL, Seeger RC . (1993). Deletion of chromosome 11 and of 14q sequences in neuroblastoma. Genes Chromosomes Cancer 7: 32–37.
Storey JD, Tibshirani R . (2003). Statistical significance for genomewide studies. Proc Natl Acad Sci USA 100: 9440–9445.
Tomioka N, Kobayashi H, Kageyama H, Ohira M, Nakamura Y, Sasaki F et al. (2003). Chromosomes that show partial loss or gain in near-diploid tumors coincide with chromosomes that show whole loss or gain in near-triploid tumors: evidence suggesting the involvement of the same genes in the tumorigenesis of high- and low-risk neuroblastomas. Genes Chromosomes Cancer 36: 139–150.
Tonini GP . (1993). Neuroblastoma: the result of multistep transformation? Stem Cells 11: 276–282.
Vandesompele J, Van Roy N, Van Gele M, Laureys G, Ambros P, Heimann P et al. (1998). Genetic heterogeneity of neuroblastoma studied by comparative genomic hybridization. Genes Chromosomes Cancer 23: 141–152.
Wei JS, Greer BT, Westermann F, Steinberg SM, Son CG, Chen QR et al. (2004). Prediction of clinical outcome using gene expression profiling and artificial neural networks for patients with neuroblastoma. Cancer Res 64: 6883–6891.
White PS, Maris JM, Beltinger C, Sulman E, Marshall HN, Fujimori M et al. (1995). A region of consistent deletion in neuroblastoma maps within human chromosome 1p36.2–36.3. Proc Natl Acad Sci USA 92: 5520–5524.
Woods WG, Gao RN, Shuster JJ, Robison LL, Bernstein M, Weitzman S et al. (2002). Screening of infants and mortality due to neuroblastoma. N Engl J Med 346: 1041–1046.
Acknowledgements
We thank institutions and hospitals for providing tumor specimens (see Supplementary Information). We also thank Shigeru Sakiyama, Hiroki Nagase, Iwao Nozawa, Tadayuki Koda and technical staff, past and present, at Division of Biochemistry, Chiba Cancer Center Research Institute. We acknowledge Hisamitsu Pharmaceutical Co. Inc., the Ministry of Education, Culture, Sports, Science and Technology of Japan, the Ministry of Health, Labour and Welfare of Japan and the Hamaguchi Foundation for the Advancement of Biochemistry for funding this work.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Oncogene web site (http://www.nature.com/onc).
Supplementary information
Rights and permissions
About this article
Cite this article
Tomioka, N., Oba, S., Ohira, M. et al. Novel risk stratification of patients with neuroblastoma by genomic signature, which is independent of molecular signature. Oncogene 27, 441–449 (2008). https://doi.org/10.1038/sj.onc.1210661
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1210661
Keywords
This article is cited by
-
Development and validation of a 21-gene prognostic signature in neuroblastoma
Scientific Reports (2023)
-
A narrative review of radiomics and deep learning advances in neuroblastoma: updates and challenges
Pediatric Radiology (2023)
-
Results of a phase II trial for high-risk neuroblastoma treatment protocol JN-H-07: a report from the Japan Childhood Cancer Group Neuroblastoma Committee (JNBSG)
International Journal of Clinical Oncology (2018)
-
11q deletion in neuroblastoma: a review of biological and clinical implications
Molecular Cancer (2017)
-
Risk stratification and therapeutics of neuroblastoma: the challenges remain
World Journal of Pediatrics (2016)