Abstract
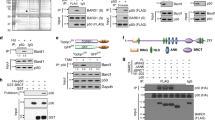
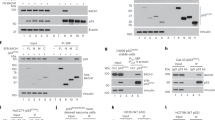
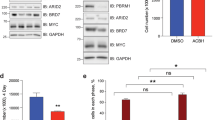
The tumor suppressor protein BARD1 plays a dual role in response to genotoxic stress: DNA repair as a BARD1–BRCA1 heterodimer and induction of apoptosis in a BRCA1-independent manner. We have constructed a series of BARD1 deletion mutants and analysed their cellular distribution and capacity to induce apoptosis. As opposed to previous studies suggesting an exclusively nuclear localization of BARD1, we found, both in tissues and cell cultures, nuclear and cytoplasmic localization of BARD1. Enhanced cytoplasmic localization of BARD1, as well as appearance of a 67 kDa C-terminal proteolytic cleavage product, coincided with apoptosis. BARD1 translocates to the nucleus independently of BRCA1. For recruitment to nuclear dots, however, the BRCA1-interacting RING finger domain is required but not sufficient. Protein levels of N-terminal RING finger deletion mutants were much higher than those of full-length BARD1, despite comparable mRNA levels, suggesting that the N-terminal region comprising the RING finger is important for BARD1 degradation. Sequences required for apoptosis induction were mapped between the ankyrin repeats and the BRCT domains coinciding with two known cancer-associated missense mutations. We suggest that nuclear and cytoplasmic localization of BARD1 reflect its dual function and that the increased cytoplasmic localization of BARD1 is associated with apoptosis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout









Similar content being viewed by others
References
Ayi TC, Tsan JT, Hwang LY, Bowcock AM and Baer R . (1998). Oncogene, 17, 2143–2148.
Baer R and Ludwig T . (2002). Curr. Opin. Genet. Dev., 12, 86–91.
Baer R . (2001). Nat. Struct. Biol., 8, 822–824.
Brzovic PS, Meza JE, King MC and Klevit RE . (2001a). J. Biol. Chem., 276, 41399–41406.
Brzovic PS, Rajagopal P, Hoyt DW, King MC and Klevit RE . (2001b). Nat. Struct. Biol., 8, 833–837.
Chen A, Kleiman FE, Manley JL, Ouchi T and Pan ZQ . (2002). J. Biol. Chem., 277, 22085–22092.
Chiba N and Parvin JD . (2002). Cancer Res., 62, 4222–4228.
Conus S, Rosse T and Borner C . (2000). Cell Death Differ., 7, 947–954.
Dechend R, Hirano F, Lehmann K, Heissmeyer V, Ansieau S, Wulczyn FG, Scheidereit C and Leutz A . (1999). Oncogene, 18, 3316–3323.
Deng CX . (2002). Oncogene, 21, 6222–6227.
Fabbro M and Henderson BR . (2003). Exp. Cell Res., 282, 59–69.
Fabbro M, Rodriguez JA, Baer R and Henderson BR . (2002). J. Biol. Chem., 277, 21315–21324.
Ferrer I and Planas AM . (2003). J. Neuropathol. Exp. Neurol., 62, 329–339.
Gautier F, Irminger-Finger I, Gregoire M, Meflah K and Harb J . (2000). Cancer Res., 60, 6895–6900.
Ghimenti C, Sensi E, Presciuttini S, Brunetti IM, Conte P, Bevilacqua G and Caligo MA . (2002). Genes Chromosomes Cancer, 33, 235–242.
Gu J, Nie L, Wiederschain D and Yuan ZM . (2001). Mol. Cell. Biol., 21, 8533–8546.
Hashizume R, Fukuda M, Maeda I, Nishikawa H, Oyake D, Yabuki Y, Ogata H and Ohta T . (2001). J. Biol. Chem., 276, 14537–14540.
Hsu LC and White RL . (1998). Proc. Natl. Acad. Sci. USA, 95, 12983–12988.
Irminger-Finger I and Leung WC . (2002). Int. J. Biochem. Cell Biol., 34, 582–587.
Irminger-Finger I, Leung WC, Li J, Dubois-Dauphin M, Harb J, Feki A, Jefford CE, Soriano JV, Jaconi M, Montesano R and Krause KH . (2001). Mol. Cell, 8, 1255–1266.
Irminger-Finger I, Soriano JV, Vaudan G, Montesano R and Sappino AP . (1998). J. Cell Biol., 143, 1329–1339.
Jin Y, Xu XL, Yang MC, Wei F, Ayi TC, Bowcock AM and Baer R . (1997). Proc. Natl. Acad. Sci. USA, 94, 12075–12080.
Kleiman FE and Manley JL . (1999). Science, 285, 1576–1579.
Kleiman FE and Manley JL . (2001). Cell, 104, 743–753.
Lohrum MA, Woods DB, Ludwig RL, Balint E and Vousden KH . (2001). Mol. Cell. Biol., 21, 8521–8532.
Lotti LV, Ottini L, D’Amico C, Gradini R, Cama A, Belleudi F, Frati L, Torrisi MR and Mariani-Costantini R . (2002). Genes Chromosomes Cancer, 35, 193–203.
Mallery DL, Vandenberg CJ and Hiom K . (2002). EMBO J., 21, 6755–6762.
Meza JE, Brzovic PS, King MC and Klevit RE . (1999). J. Biol. Chem., 274, 5659–5665.
Morris JR, Keep NH and Solomon E . (2002). J. Biol. Chem., 277, 9382–9386.
Parvin JD . (2001). Proc. Natl. Acad. Sci. USA, 98, 5952–5954.
Rodriguez JA, Schuchner S, Au WW, Fabbro M and Henderson BR . (2003). Oncogene.
Ruffner H, Joazeiro CA, Hemmati D, Hunter T and Verma IM . (2001). Proc. Natl. Acad. Sci. USA, 98, 5134–5139.
Scully R, Chen J, Ochs RL, Keegan K, Hoekstra M, Feunteun J and Livingston DM . (1997a). Cell, 90, 425–435.
Scully R, Chen J, Plug A, Xiao Y, Weaver D, Feunteun J, Ashley T and Livingston DM . (1997b). Cell, 88, 265–275.
Soriano JV, Irminger-Finger I, Uyttendaele H, Vaudan G, Kitajewski J, Sappino AP and Montesano R . (2000). Adv. Exp. Med. Biol., 480, 175–184.
Spahn L, Petermann R, Siligan C, Schmid JA, Aryee DN and Kovar H . (2002). Cancer Res., 62, 4583–4587.
Thai TH, Du F, Tsan JT, Jin Y, Phung A, Spillman MA, Massa HF, Muller CY, Ashfaq R, Mathis JM, Miller DS, Trask BJ, Baer R and Bowcock AM . (1998). Hum. Mol. Genet., 7, 195–202.
Wu LC, Wang ZW, Tsan JT, Spillman MA, Phung A, Xu XL, Yang MC, Hwang LY, Bowcock AM and Baer R . (1996). Nat. Genet., 14, 430–440.
Xia Y, Pao GM, Chen HW, Verma IM and Hunter T . (2003). J. Biol. Chem., 278, 5255–5263.
Yu X and Baer R . (2000). J. Biol. Chem., 275, 18541–18549.
Acknowledgements
This work was supported by Swiss National Science Foundation grant to IIF and KHK, and Johnson & Johnson and AETAS awards to IIF. We are grateful to Dr Gilbert LeNoir and Dr Anne Vincent for the generous gift of the BRCA1 antibody, Dr Monica Sauer for discussing with us unpublished observations, Dr Wai-Choi Leung for critical comments on the manuscript, and Aurélie Caillon for excellent technical support.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jefford, C., Feki, A., Harb, J. et al. Nuclear–cytoplasmic translocation of BARD1 is linked to its apoptotic activity. Oncogene 23, 3509–3520 (2004). https://doi.org/10.1038/sj.onc.1207427
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1207427
Keywords
This article is cited by
-
A synergetic effect of BARD1 mutations on tumorigenesis
Nature Communications (2021)
-
RNF19A-mediated ubiquitination of BARD1 prevents BRCA1/BARD1-dependent homologous recombination
Nature Communications (2021)
-
Expression of an Oncogenic BARD1 Splice Variant Impairs Homologous Recombination and Predicts Response to PARP-1 Inhibitor Therapy in Colon Cancer
Scientific Reports (2016)
-
BARD1 mediates TGF-β signaling in pulmonary fibrosis
Respiratory Research (2015)
-
Alternative splicing in cancer: implications for biology and therapy
Oncogene (2015)