Abstract
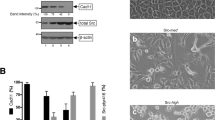
The RelA transcription factor is part of dimeric complexes, most commonly either p50-RelA (NF-κB) heterodimers or RelA homodimers, that control a variety of cellular processes. Immortalized embryonic fibroblasts established from rela knockout mice have previously been shown to be more sensitive to apoptosis induced by tumor necrosis factor (TNF) than are control fibroblasts. In this report, we show that one line of rela−/− fibroblasts has additional phenotypes that distinguish them from control mouse fibroblasts. As compared to normal 3T3 cells, RelA-deficient fibroblasts have a spindled morphology, are less adherent to culture dishes, grow to a higher saturation density, and can form colonies in soft agar. These properties are consistent with a weakly transformed phenotype for rela−/− cells. Furthermore, RelA-deficient fibroblasts can form tumors in immunodeficient mice, but these tumors regress, probably because of the sensitivity of these cells to TNF. The ability of rela−/− fibroblasts to form colonies in soft agar can be reverted by re-expression of wild-type mouse RelA, but not by expression of RelA mutants that cannot form homodimers. There is no clear correlation between the absence of RelA and the levels of expression of other Rel/NF-κB family members or adhesion proteins (ICAM-1 and VCAM-1) whose genes have upstream κB sites. Taken together, these results suggest that RelA has tumor suppressing activity under some circumstances and that RelA complexes are involved in the control of a variety of cellular properties associated with oncogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Bancroft GJ, Sheehan KC, Schreiber RD, Unanue ER . 1989 J. Immunol. 143: 127–130
Barkett M, Gilmore TD . 1999 Oncogene 18: 6910–6924
Beg AA, Baltimore D . 1996 Science 274: 782–784
Beg AA, Sha WC, Bronson RT, Ghosh S, Baltimore D . 1995 Nature 376: 167–170
Capobianco A, Gilmore TD . 1991 Oncogene 6: 2203–2210
Cogswell PC, Guttridge DC, Funkhouser WK, Baldwin Jr AS . 2000 Oncogene 19: 1123–1131
Doi TS, Marino MW, Takahashi T, Sakakaura T, Yoshida T, Old LS, Obata Y . 1999 Proc. Natl. Acad. Sci. USA 96: 2994–2999
Dorshkind K, Pollack SB, Bosma MJ, Phillips RA . 1985 J. Immunol. 134: 3798–3801
Finco TS, Westwick JK, Norris JL, Beg AA, Der CJ, Baldwin Jr AS . 1997 J. Biol. Chem. 272: 24113–24116
Ganchi PA, Sun S-C, Greene WC, Ballard DW . 1993 Mol. Cell. Biol. 13: 7826–7835
Gilmore TD . 1999 Oncogene 18: 6842–6844
Gilmore TD, Cormier C, Jean-Jacques J, Gapuzan M-E . 2001 Oncogene 20: 7098–7103
Gilmore TD, Gapuzan M-E, Kalaitzidis D, Starczynowski D . 2002 Cancer Lett. in press
Hannink M, Temin HM . 1990 Oncogene 5: 1843–1850
Iademarco MF, McQuillan JJ, Rosen GD, Dean DC . 1992 J. Biol. Chem. 267: 16323–16329
Jo H, Zhong R, Zhong H, McKinsey TA, Shao J, Beauchamp RD, Ballard DW, Liang P . 2000 Oncogene 19: 841–849
Morgenstern JP, Land H . 1990 Nucleic Acids Res. 18: 3587–3596
Pahl H . 1999 Oncogene 18: 6853–6866
Rayet B, Gélinas C . 1999 Oncogene 18: 6938–6947
Ricci A, Biroccio A, Trisciuoglio D, Cippitelli M, Zupi G, Bufalo DD . 2001 Br. J. Cancer 85: 1914–1921
Ryan KM, Ernst MK, Rice NR, Vousden KH . 2000 Nature 404: 892–897
Sif S, Gilmore TD . 1993 J. Virol. 67: 7612–7617
Trecca D, Guerrini L, Fracchiolla NS, Pomati M, Baldini L, Maiolo AT, Neri A . 1997 Oncogene 14: 791–799
van de Stolpe A, Caldenhoven E, Stade BG, Koenderman L, Raaijmakers JAM, Johnson JP, van der Saag PT . 1994 J. Biol. Chem. 269: 6185–6192
van Hogerlinden M, Rozell BL, Ährlund-Richter L, Toftgård R . 1999 Cancer Res. 59: 3299–3303
Wang CY, Cusack Jr JC, Liu R, Baldwin Jr AS . 1999 Nat. Med. 5: 412–417
Zong W-X, Bash J, Gélinas C . 1998 Cell Death Differ. 5: 963–972
Acknowledgements
We thank Mark Hannink, Alexander Hoffman, and David Baltimore for the rela−/− fibroblasts and helpful discussions, Youssef Jounaidi for the BOSC23 retrovirus-packaging cells, and Geoffrey Cooper for helpful discussions. We thank Nancy Rice for the anti-RelA antiserum. We thank Youssef Jounaidi and Beverly Keniston for help with mouse tumorigenicity assays. This work was supported by NIH grant CA47763 (TD Gilmore) and a small grant (to PV Yufit) from the Undergraduate Research Opportunities Program of Boston University. PV Yufit was partially supported by an NSF-REU grant and performed research as part of the Undergraduate Honors Program in Biochemistry and Molecular Biology at Boston University.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Gapuzan, ME., Yufit, P. & Gilmore, T. Immortalized embryonic mouse fibroblasts lacking the RelA subunit of transcription factor NF-κB have a malignantly transformed phenotype. Oncogene 21, 2484–2492 (2002). https://doi.org/10.1038/sj.onc.1205333
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1205333
Keywords
This article is cited by
-
RelB inhibits cell proliferation and tumor growth through p53 transcriptional activation
Oncogene (2013)
-
Epstein-Barr virus-encoded EBNA1 inhibits the canonical NF-κB pathway in carcinoma cells by inhibiting IKK phosphorylation
Molecular Cancer (2010)
-
Overexpression of an activated REL mutant enhances the transformed state of the human B-lymphoma BJAB cell line and alters its gene expression profile
Oncogene (2009)
-
Good cop, bad cop: the different faces of NF-κB
Cell Death & Differentiation (2006)
-
Immortalized fibroblasts from NF-κB RelA knockout mice show phenotypic heterogeneity and maintain increased sensitivity to tumor necrosis factor α after transformation by v-Ras
Oncogene (2005)