Abstract
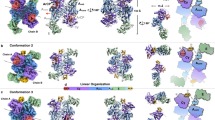
Nonribosomal peptide synthetases (NRPSs) are large, multidomain enzymes that biosynthesize medically important natural products. We report the crystal structure of the free-standing NRPS condensation (C) domain VibH, which catalyzes amide bond formation in the synthesis of vibriobactin, a Vibrio cholerae siderophore. Despite low sequence identity, NRPS condensation enzymes are structurally related to chloramphenicol acetyltransferase (CAT) and dihydrolipoamide acyltransferases. However, although the latter enzymes are homotrimers, VibH is a monomeric pseudodimer. The VibH structure is representative of both NRPS condensation and epimerization domains, as well as the condensation-variant cyclization domains, which are all expected to be monomers. Surprisingly, despite favorable positioning in the active site, a universally conserved histidine important in CAT and in other C domains is not critical for general base catalysis in VibH.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Marahiel, M.A., Stachelhaus, T. & Mootz, H.D. Chem. Rev. 97, 2651–2673 (1997).
Stachelhaus, T. & Walsh, C.T. Biochemistry 39, 5775–5787 (2000).
Marshall, C.G., Hillson, N.J. & Walsh, C.T. Biochemistry 41, 244–250 (2002).
Keating, T.A., Miller, D.A. & Walsh, C.T. Biochemistry 39, 4729–4739 (2000).
Tsai, S.C. et al. Proc. Natl. Acad. Sci. USA 98, 14808–14813 (2001).
Weber, T., Baumgartner, R., Renner, C., Marahiel, M.A. & Holak, T.A. Structure Fold. Des. 8, 407–418 (2000).
Conti, E., Stachelhaus, T., Marahiel, M.A. & Brick, P. EMBO J. 16, 4174–4183 (1997).
Bruner, S.D. et al. Structure 10, 301–310 (2002).
De Crécy-Lagard, V., Marlière, P. & Saurin, W. C. R. Acad. Sci. III 318, 927–936 (1995).
Lewendon, A., Murray, I.A., Shaw, W.V., Gibbs, M.R. & Leslie, A.G. Biochemistry 33, 1944–1950 (1994).
Stachelhaus, T., Mootz, H.D., Bergendahl, V. & Marahiel, M.A. J. Biol. Chem. 273, 22773–22781 (1998).
Bergendahl, V., Linne, U. & Marahiel, M.A. Eur. J. Biochem. 269, 620–629 (2002).
Keating, T.A., Marshall, C.G. & Walsh, C.T. Biochemistry 39, 15513–15521 (2000).
Keating, T.A., Marshall, C.G. & Walsh, C.T. Biochemistry 39, 15522–15530 (2000).
Murzin, A.G., Brenner, S.E., Hubbard, T. & Chothia, C. J. Mol. Biol. 247, 536–540 (1995).
Holm, L. & Sander, C. Trends Biochem. Sci. 20, 478–480 (1995).
Leslie, A.G. J. Mol. Biol. 213, 167–186 (1990).
Leslie, A.G., Moody, P.C. & Shaw, W.V. Proc. Natl. Acad. Sci. USA 85, 4133–4137 (1988).
Mattevi, A., Obmolova, G., Kalk, K.H., Teplyakov, A. & Hol, W.G. Biochemistry 32, 3887–3901 (1993).
Mattevi, A. et al. Science 255, 1544–1550 (1992).
Mattevi, A. et al. J. Mol. Biol. 230, 1183–1199 (1993).
Hendle, J. et al. Biochemistry 34, 4287–4298 (1995).
Niu, X.D., Stoops, J.K. & Reed, L.J. Biochemistry 29, 8614–8619 (1990).
Shaw, W.V. & Leslie, A.G. Annu. Rev. Biophys. Biophys. Chem. 20, 363–386 (1991).
Lewendon, A., Murray, I.A., Kleanthous, C., Cullis, P.M. & Shaw, W.V. Biochemistry 27, 7385–7390 (1988).
Lessard, I.A., Healy, V.L., Park, I.S. & Walsh, C.T. Biochemistry 38, 14006–14022 (1999).
Gulick, A.M., Schmidt, D.M., Gerlt, J.A. & Rayment, I. Biochemistry 40, 15716–15724 (2001).
Otwinowski, Z. Data Collection and Processing (eds Sawer, L., Isaacs, N. & Bailey, S.) 55–62 (SERC, Daresbury Laboratory, Warrington; 1993).
Terwilliger, T.C. & Berendzen, J. Acta Crystallogr. D 55, 849–861 (1999).
Collaborative Computation Project, Number 4. Acta Crystallogr. D 50, 760–763 (1994).
Cowtan, K. & Main, P. Acta Crystallogr. D 54, 487–493 (1998).
Jones, T.A., Zou, J.W., Cowan, S. & Kjeldgaard, M. Acta Crystallogr. A 47, 110–119 (1991).
Brünger, A.T. et al. Acta Crystallogr. D 54, 905–921 (1998).
Thompson, J., Gibson, T., Plewniak, F., Jeanmougin, F. & Higgins, D. Nucleic Acids Res. 25, 4876–4882 (1997).
Cuff, J.A., Clamp, M.E., Siddiqui, A.S., Finlay, M. & Barton, G.J. Bioinformatics 14, 892–893 (1998).
Kraulis, P.J. J. Appl. Crystallogr. 24, 946–950 (1991).
Merritt, E.A. & Bacon, D.J. Methods Enzymol. 277, 505–524 (1997).
Nicholls, A., Sharp, K.A. & Honig, B. Proteins 11, 281–296 (1991).
Acknowledgements
We thank S.D. Bruner, V.N. Malashkevich and L. Luo for suggestions and technical assistance, A.G. Leslie for providing the coordinates of CAT–CoA, and A. Saxena, R. Sweet and the NSLS beamline staff for their generous help. T.A.K. was a fellow of the Damon Runyon-Walter Winchell Foundation. C.G.M. received funding from the NSERC of Canada and the Canadian Institutes of Health Research. This work was funded by the NIH (C.T.W.) and by MIT startup funds (A.E.K.).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Keating, T., Marshall, C., Walsh, C. et al. The structure of VibH represents nonribosomal peptide synthetase condensation, cyclization and epimerization domains. Nat Struct Mol Biol 9, 522–526 (2002). https://doi.org/10.1038/nsb810
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb810
This article is cited by
-
Characterization of protein interaction surface on fatty acyl selectivity of starter condensation domain in lipopeptide biosynthesis
Applied Microbiology and Biotechnology (2020)