Abstract
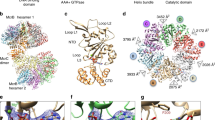
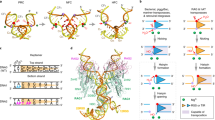
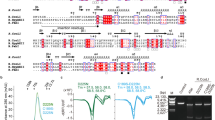
Restriction endonuclease BglII completely encircles its target DNA, making contacts to both the major and minor grooves. To allow the DNA to enter and leave the binding cleft, the enzyme dimer has to rearrange. To understand how this occurs, we have solved the structure of the free enzyme at 2.3 Å resolution, as a complement to our earlier work on the BglII–DNA complex. Unexpectedly, the enzyme opens by a dramatic `scissor-like' motion, accompanied by a complete rearrangement of the α-helices at the dimer interface. Moreover, within each monomer, a set of residues — a ‘lever’ — lowers or raises to alternately sequester or expose the active site residues. Such an extreme difference in free versus complexed structures has not been reported for other restriction endonucleases. This elegant mechanism for capturing DNA may extend to other enzymes that encircle DNA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Winkler, F.K. et al. EMBO J. 12, 1781–1795 (1993).
Newman, M., Strzelecka, T., Dorner, L.F., Schildkraut, I. & Aggarwal, A.K. Structure 2, 439–452 (1994).
Newman, M., Strzelecka, T., Dorner, L.F., Schildkraut, I. & Aggarwal, A.K. Nature 368, 660–664 (1994).
Newman, M., Strzelecka, T., Dorner, L., Schildkraut, I. & Aggarwal, A.K. Science 269, 656–663 (1995).
Athanasiadis, A. et al. Nature Struct. Biol. 1, 469–475 (1994).
Cheng, X., Balendiran, K., Schildkraut, I. & Anderson, J.E. EMBO J. 13, 3927–3935 (1994).
Wah, D.A., Hirsch, J.A., Dorner, L.F., Schildkraut, I. & Aggarwal, A.K. Nature 388, 97–100 (1997).
Wah, D.A., Bitinaite, J., Schildkraut, I. & Aggarwal, A.K. Proc. Natl. Acad. Sci. USA 95, 10564–10569 (1998).
Roberts, R.J. & Halford, S.E. In Nucleases (eds., Linn, S.M., Lloyd, R.S. & Roberts, R.J.) (Cold Spring Harbor, New York; 1993).
Aggarwal, A.K. Curr. Opin. Struct. Biol. 5, 11–19 (1995).
Pingoud, A. & Jeltsch, A. Eur. J. Biochem. 246, 1–22 (1997).
Kim, Y., Grable, J.C., Choi, P.J., Greene, P. & Rosenberg, J.M. Science 249, 1307–1309 (1990).
Deibert, M., Grazulis, S., Janulaitis, A., Siksnys, V. & Huber, R. EMBO J. 18, 5805–5816 (1999).
Lukacs, C.M., Kucera, R., Schildkraut, I. & Aggarwal, A.K. Nature Struct. Biol. 7, 134–140 (2000).
Perona, J.J. & Martin, A.M. J. Mol. Biol. 273, 207–225 (1997).
Anton, B.P. et al. Gene 187, 19–27 (1997).
Nicholls, A., Sharp, K. & Honig, B. Proteins 11, 281–296 (1991).
Janin, J., Miller, S. & Chothia, C. J. Mol. Biol. 204, 155–164 (1988).
Jones, S. & Thornton, J.M. Proc. Natl. Acad. Sci. USA 93, 13–20 (1996).
Chantalat, L. et al. EMBO J. 18, 2930–2940 (1999).
Chantalat, L. et al. Mol. Cell 6, 183–189 (2000).
Heath, P.J., Stephens, K.M., Monnat, R.J., Jr. & Stoddard, B.L. Nature Struct. Biol. 4, 468–476 (1997).
Galburt, E.A. et al. J. Mol. Biol. 300, 877–887 (2000).
Ghosh, G., Van Duyne, G., Ghosh, S. & Sigler, P.B. Nature 373, 303–310 (1995).
Muller, C.W., Rey, F.A., Sodeoka, M., Verdine, G.L. & Harrison, S.C. Nature 373, 311–317 (1995).
Muller, C.W., Rey, F.A. & Harrison, S.C. Nature Struct. Biol. 3, 224–227 (1996).
Ferre-D'Amare, A.R., Prendergast, G.C., Ziff, E.B. & Burley, S.K. Nature 363, 38–45 (1993).
Ellenberger, T., Fass, D., Arnaud, M. & Harrison, S.C. Genes Dev. 8, 970–980 (1994).
Fairman, R., Beran-Steed, R.K. & Handel, T.M. Protein Sci. 6, 175–184 (1997).
Lu, B., Morrow, J. & Weisgraber, K.H. J. Biol. Chem. 275, 20775–20781 (2000).
Sutton, R.B., Fasshauer, D., Jahn, R. & Brunger, A.T. Nature 395, 347–353 (1998).
Viadiu, H. & Aggarwal, A.K. Mol. Cell 5, 889–895 (2000).
Otwinowski, Z. & Minor, W. Methods Enzymol. 276, 307–326 (1997).
Navaza, J. In Molecular replacement (eds., Dodson, E.J., Gover, S. & Wolf, W.) (Science and Engineering Research Council, Daresbury Laboratory, Warrington, UK; 1992).
Brunger, A.T. et al. Acta Crystallogr. D 54, 905 (1998).
Jones, A.T., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Acta Crystallogr. A 47, 110–119 (1991).
Fortelle, d.L. & Bricogne, G. Methods Enzymol. 276, 472–494 (1997).
Abrahams, J.P. & Leslie, A.G.W. Acta Crystallogr. D 52, 32–42 (1996).
Kraulis, P. J. Appl. Crystallogr. 24, 946–950 (1991).
Merritt, E.A. & Murphy, M.E.P. Acta Crystallogr. D 50, 869–873 (1994).
Acknowledgements
We thank L. Berman and H. Lewis for facilitating data collection at NSLS. A.K.A. is supported by a grant from the NIH, and C.M.L. is supported by a Cancer Research Fund of the Damon Runyon-Walter Winchell Foundation Fellowship.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lukacs, C., Kucera, R., Schildkraut, I. et al. Structure of free BglII reveals an unprecedented scissor-like motion for opening an endonuclease. Nat Struct Mol Biol 8, 126–130 (2001). https://doi.org/10.1038/84111
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/84111
This article is cited by
-
2′-O-methyl nucleotide modified DNA substrates influence the cleavage efficiencies of BamHI and BglII
Journal of Biosciences (2014)