Abstract
To fully understand the transport mechanism of Na+/H+ exchangers, it is necessary to clearly establish the global rearrangements required to facilitate ion translocation. Currently, two different transport models have been proposed. Some reports have suggested that structural isomerization is achieved through large elevator-like rearrangements similar to those seen in the structurally unrelated sodium-coupled glutamate-transporter homolog GltPh. Others have proposed that only small domain movements are required for ion exchange, and a conventional rocking-bundle model has been proposed instead. Here, to resolve these differences, we report atomic-resolution structures of the same Na+/H+ antiporter (NapA from Thermus thermophilus) in both outward- and inward-facing conformations. These data combined with cross-linking, molecular dynamics simulations and isothermal calorimetry suggest that Na+/H+ antiporters provide alternating access to the ion-binding site by using elevator-like structural transitions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Brett, C.L., Donowitz, M. & Rao, R. Evolutionary origins of eukaryotic sodium/proton exchangers. Am. J. Physiol. Cell Physiol. 288, C223–C239 (2005).
Fuster, D.G. & Alexander, R.T. Traditional and emerging roles for the SLC9 Na+/H+ exchangers. Pflugers Arch. 466, 61–76 (2014).
Verma, V., Bali, A., Singh, N. & Jaggi, A.S. Implications of sodium hydrogen exchangers in various brain diseases. J. Basic Clin. Physiol. Pharmacol. 26, 417–426 (2015).
Hunte, C. et al. Structure of a Na+/H+ antiporter and insights into mechanism of action and regulation by pH. Nature 435, 1197–1202 (2005).
Lee, C. et al. Crystal structure of the sodium-proton antiporter NhaA dimer and new mechanistic insights. J. Gen. Physiol. 144, 529–544 (2014).
Lee, C. et al. A two-domain elevator mechanism for sodium/proton antiport. Nature 501, 573–577 (2013).
Maes, M., Rimon, A., Kozachkov-Magrisso, L., Friedler, A. & Padan, E. Revealing the ligand binding site of NhaA Na+/H+ antiporter and its pH dependence. J. Biol. Chem. 287, 38150–38157 (2012).
Wöhlert, D., Kühlbrandt, W. & Yildiz, O. Structure and substrate ion binding in the sodium/proton antiporter PaNhaP. eLife 3, e03579 (2014).
Arkin, I.T. et al. Mechanism of Na+/H+ antiporting. Science 317, 799–803 (2007).
Reyes, N., Ginter, C. & Boudker, O. Transport mechanism of a bacterial homologue of glutamate transporters. Nature 462, 880–885 (2009).
Paulino, C., Wöhlert, D., Kapotova, E., Yildiz, Ö. & Kühlbrandt, W. Structure and transport mechanism of the sodium/proton antiporter MjNhaP1. eLife 3, e03583 (2014).
Furrer, E.M., Ronchetti, M.F., Verrey, F. & Pos, K.M. Functional characterization of a NapA Na+/H+ antiporter from Thermus thermophilus . FEBS Lett. 581, 572–578 (2007).
Padan, E. et al. NhaA of Escherichia coli, as a model of a pH-regulated Na+/H+antiporter. Biochim. Biophys. Acta 1658, 2–13 (2004).
Kozachkov, L. & Padan, E. Conformational changes in NhaA Na+/H+ antiporter. Mol. Membr. Biol. 30, 90–100 (2013).
Rimon, A., Kozachkov-Magrisso, L. & Padan, E. The unwound portion dividing helix IV of NhaA undergoes a conformational change at physiological pH and lines the cation passage. Biochemistry 51, 9560–9569 (2012).
Akyuz, N. et al. Transport domain unlocking sets the uptake rate of an aspartate transporter. Nature 518, 68–73 (2015).
West, I.C. & Mitchell, P. Proton/sodium ion antiport in Escherichia coli . Biochem. J. 144, 87–90 (1974).
Taglicht, D., Padan, E. & Schuldiner, S. Overproduction and purification of a functional Na+/H+ antiporter coded by nhaA (ant) from Escherichia coli . J. Biol. Chem. 266, 11289–11294 (1991).
Mager, T., Rimon, A., Padan, E. & Fendler, K. Transport mechanism and pH regulation of the Na+/H+ antiporter NhaA from Escherichia coli: an electrophysiological study. J. Biol. Chem. 286, 23570–23581 (2011).
Caˇlinescu, O., Paulino, C., Kühlbrandt, W. & Fendler, K. Keeping it simple, transport mechanism and pH regulation in Na+/H+ exchangers. J. Biol. Chem. 289, 13168–13176 (2014).
Caˇlinescu, O., Danner, E., Böhm, M., Hunte, C. & Fendler, K. Species differences in bacterial NhaA Na+/H+ exchangers. FEBS Lett. 588, 3111–3116 (2014).
Cao, Y. et al. Crystal structure of a phosphorylation-coupled saccharide transporter. Nature 473, 50–54 (2011).
Luo, P. et al. Crystal structure of a phosphorylation-coupled vitamin C transporter. Nat. Struct. Mol. Biol. 22, 238–241 (2015).
Johnson, Z.L., Cheong, C.G. & Lee, S.Y. Crystal structure of a concentrative nucleoside transporter from Vibrio cholerae at 2.4 Å. Nature 483, 489–493 (2012).
Bolla, J.R. et al. Crystal structure of the Alcanivorax borkumensis YdaH transporter reveals an unusual topology. Nat. Commun. 6, 6874 (2015).
Su, C.C. et al. Structure and function of Neisseria gonorrhoeae MtrF illuminates a class of antimetabolite efflux pumps. Cell Reports 11, 61–70 (2015).
Hu, N.J., Iwata, S., Cameron, A.D. & Drew, D. Crystal structure of a bacterial homologue of the bile acid sodium symporter ASBT. Nature 478, 408–411 (2011).
Zhou, X. et al. Structural basis of the alternating-access mechanism in a bile acid transporter. Nature 505, 569–573 (2014).
Vinothkumar, K.R., Smits, S.H. & Kühlbrandt, W. pH-induced structural change in a sodium/proton antiporter from Methanococcus jannaschii . EMBO J. 24, 2720–2729 (2005).
Paulino, C. & Kühlbrandt, W. pH- and sodium-induced changes in a sodium/proton antiporter. eLife 3, e01412 (2014).
Kozachkov, L. & Padan, E. Site-directed tryptophan fluorescence reveals two essential conformational changes in the Na+/H+ antiporter NhaA. Proc. Natl. Acad. Sci. USA 108, 15769–15774 (2011).
Appel, M., Hizlan, D., Vinothkumar, K.R., Ziegler, C. & Kühlbrandt, W. Conformations of NhaA, the Na+/H+ exchanger from Escherichia coli, in the pH-activated and ion-translocating states. J. Mol. Biol. 388, 659–672 (2009).
Yernool, D., Boudker, O., Jin, Y. & Gouaux, E. Structure of a glutamate transporter homologue from Pyrococcus horikoshii . Nature 431, 811–818 (2004).
Wagner, S. et al. Tuning Escherichia coli for membrane protein overexpression. Proc. Natl. Acad. Sci. USA 105, 14371–14376 (2008).
Lee, C. et al. MemStar: a one-shot Escherichia coli-based approach for high-level bacterial membrane protein production. FEBS Lett. 588, 3761–3769 (2014).
Ishmukhametov, R.R., Galkin, M.A. & Vik, S.B. Ultrafast purification and reconstitution of His-tagged cysteine-less Escherichia coli F1Fo ATP synthase. Biochim. Biophys. Acta 1706, 110–116 (2005).
Wiedenmann, A., Dimroth, P. & von Ballmoos, C. Functional asymmetry of the F(0) motor in bacterial ATP synthases. Mol. Microbiol. 72, 479–490 (2009).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Evans, P.R. & Murshudov, G.N. How good are my data and what is the resolution? Acta Crystallogr. D Biol. Crystallogr. 69, 1204–1214 (2013).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Davis, I.W., Murray, L.W., Richardson, J.S. & Richardson, D.C. MOLPROBITY: structure validation and all-atom contact analysis for nucleic acids and their complexes. Nucleic Acids Res. 32, W615–W619 (2004).
Jones, T.A. & Kjeldgaard, M. Electron-density map interpretation. Methods Enzymol. 277, 173–208 (1997).
Kleywegt, G.J. & Jones, T.A. A super position. Joint CCP4 ESF-EACBM Newsl. Protein Crystallogr. 31, 9–14 (1994).
Landau, M. et al. ConSurf 2005: the projection of evolutionary conservation scores of residues on protein structures. Nucleic Acids Res. 33, W299–W302 (2005).
Pronk, S. et al. GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29, 845–854 (2013).
MacKerell, A.D. et al. All-atom empirical potential for molecular modeling and dynamics studies of proteins. J. Phys. Chem. B 102, 3586–3616 (1998).
Mackerell, A.D. Jr., Feig, M. & Brooks, C.L. III. Extending the treatment of backbone energetics in protein force fields: limitations of gas-phase quantum mechanics in reproducing protein conformational distributions in molecular dynamics simulations. J. Comput. Chem. 25, 1400–1415 (2004).
Best, R.B. et al. Optimization of the additive CHARMM all-atom protein force field targeting improved sampling of the backbone ϕ, ψ and side-chain χ(1) and χ(2) dihedral angles. J. Chem. Theory Comput. 8, 3257–3273 (2012).
Klauda, J.B. et al. Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J. Phys. Chem. B 114, 7830–7843 (2010).
Scott, K.A. et al. Coarse-grained MD simulations of membrane protein-bilayer self-assembly. Structure 16, 621–630 (2008).
Stansfeld, P.J. & Sansom, M.S.P. From coarse grained to atomistic: a serial multiscale approach to membrane protein simulations. J. Chem. Theory Comput. 7, 1157–1166 (2011).
Søndergaard, C.R., Olsson, M.H.M., Rostkowski, M. & Jensen, J.H. Improved treatment of ligands and coupling effects in empirical calculation and rationalization of pKa values. J. Chem. Theory Comput. 7, 2284–2295 (2011).
Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 126, 014101 (2007).
Parrinello, M. & Rahman, A. Polymorphic transitions in single crystals: a new molecular dynamics method. J. Appl. Phys. 52, 7182–7190 (1981).
Essman, U. et al. A smooth particle mesh Ewald method. J. Chem. Phys. 103, 8577–8592 (1995).
Piggot, T.J., Piñeiro, Á. & Khalid, S. Molecular dynamics simulations of phosphatidylcholine membranes: a comparative force field study. J. Chem. Theory Comput. 8, 4593–4609 (2012).
Hess, B. P-LINCS:a parallel linear constraint solver for molecular simulation. J. Chem. Theory Comput. 4, 116–122 (2008).
Miyamoto, S. & Kollman, P.A. SETTLE: an analytical version of the SHAKE and RATTLE algorithms for rigid water models. J. Comput. Chem. 13, 952–962 (1992).
Michaud-Agrawal, N., Denning, E.J., Woolf, T.B. & Beckstein, O. MDAnalysis: a toolkit for the analysis of molecular dynamics simulations. J. Comput. Chem. 32, 2319–2327 (2011).
Lomize, M.A., Lomize, A.L., Pogozheva, I.D. & Mosberg, H.I. OPM: orientations of proteins in membranes database. Bioinformatics 22, 623–625 (2006).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38, 27–28 (1996).
Dahl, A.C.E., Chavent, M. & Sansom, M.S.P. Bendix: intuitive helix geometry analysis and abstraction. Bioinformatics 28, 2193–2194 (2012).
Acknowledgements
We are grateful to G. Verdon for discussions and comments. Data were collected at Diamond Light Source with excellent assistance from beamline scientists. This work was supported by the Swedish Research Council (D.D.) and the Knut and Alice Wallenberg Foundation (D.D.). The authors are grateful for the use of the Membrane Protein Laboratory supported by the Wellcome Trust UK (grant 062164/Z/00/Z) at the Diamond Light Source Limited and the Centre for Biomembrane Research supported by the Swedish Foundation for Strategic Research. Computer simulations were partially run on the Extreme Science and Engineering Discovery Environment (XSEDE), which is supported by US National Science Foundation grant OCI-1053575 (allocation TG-MCB130177 to O.B.). M.C. was supported as a Wenner-Gren postdoctoral fellow, and D.D. is supported as a European Molecular Biology Organization (EMBO) Young Investigator.
Author information
Authors and Affiliations
Contributions
D.D. designed the project. Cloning, expression screening, protein purification and crystallization of inward-facing NapA were carried out by M.C. and P.U. LCP crystallization of outward-facing NapA was carried out by E.N. Data collection and structural determination were carried out by M.C. and D.D. with assistance from E.N. and A.D.C. Experiments for functional analysis were carried out by P.U., I.W., S.A.-H. and M.C. MD simulations were carried out by D.L.D. and O.B. D.D. wrote the manuscript with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Assessing disulfide-bond formation in NapA cysteine mutants.
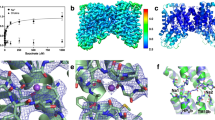
a. Reactivity of purified wild type NapA and single, double, and triple cysteine mutants thereof to the N-[4-(7-diethylamino-4-methyl-3-coumarinyl)phenyl]-maleimide (CPM) dye (see Methods). b. Representative ACMA fluorescence traces of NapA wild type antiport activity in proteoliposomes as outlined in the schematic (insert adapted from ref. 6). The addition of ATP (0.5-3 min), NaCl/LiCl (3rd min) and NH4Cl (4th min) is indicated by arrows and labeled. Black trace (presence of DTT; reducing) and the red trace (after DTT removal; non-reducing). c. SDS-PAGE analysis of a triple cysteine mutant of NapA (used previously for phasing/structure determination ref. 6) and the dimer domain mutant I55C in the presence or absence of DTT as labeled. d. Representative Li+ catalysed antiport activity of the I55C mutant under either reducing (black trace) or non-reducing (red trace) conditions. e. Cartoon representation of the inward-facing NapA structure showing the positions of V71 (red sticks) in the dimer domain (wheat) and I130 (yellow sticks) and L141 (green sticks) in the core domain (blue). f. Reactivity of purified single and double cysteine mutants V71C and V71C L141C (open circle) to mPEG-5K (black circle). g-h. Representative ACMA fluorescence traces for V71C L141C and V71C I130C mutants in the presence of DTT (reducing, black trace), after DTT removal (non-reducing, red trace) and after re-addition of DTT (reducing, blue trace).
Supplementary Figure 2 Electron density maps of the disulfide-locked inward-facing NapA structure and TM5 in both conformations.
a. Stereoview showing 2FoFc electron density calculated after the final refinement (contoured at 1.5σ level) with the final inward-facing structure overlaid (purple core; orange dimer) from the extracellular side (top) and from the side-view (bottom). b NapA structures in the inward-facing and outward-facing conformation as superimposed on their respective dimer domains. The ion-binding D157 (orange) displacement between the two states is shown between the position of their Cα atom or their Cγ atom. c 2FoFc electron density calculated after the final refinement (contoured at 1.0σ level) around TM5 in the outward-facing (left panel) and inward-facing (right panel) conformation.
Supplementary Figure 3 Analysis of the disulfide-trapped structure of NapA in an inward-facing conformation.
a. MD simulations of the disulfide cross-linked crystal structure V31C I130C (red) and the wild-type structure (blue), which was modelled from the cross-linked structure, were compared for protomer A and B. Cα r.m.s.d as a function of time, after r.m.s.d-fitting to the crystal structure. Data were averaged over 10-ns windows and bands indicate the 95% quantile of the data. b. Per-residue root mean square fluctuations (r.m.s.f.), relative to the average of all MD conformations. Dashed lines indicate the positions of the cross-linking residues C31 and C130. Gray boxes show the locations of helices TM 1 to −12. Two yellow boxes indicate putative hinge regions between dimer and core domain. Two independent repeat simulations of the wild type of 300 ns length (not shown) display the same behavior with r.m.s.d values between 2 Å and 2.5 Å and very similar r.m.s.f. profiles. c Superimposition of disulfide-trapped NapA (wheat dimer, light-blue core) and inward-facing MjNhaP1 (grey; PDB 4CZB) structures across both core and dimer domains evenly. Top inset shows the relative positioning of the ion-binding aspartate residue D157 in NapA (orange stick) and equivalent aspartate D161 in MjNhaP1 (grey stick). Bottom inset shows the region around the disulfide formed between C130 (green stick) and C31 (red stick) residues in NapA and the equivalent helices in MjNhaP1. d. Cytoplasmic view showing the highly conserved residues in NapA that becomes accessible upon the cavity opening to the inside in the inward-facing disulfide-trapped structure (below), as compared to the published outward-facing structure (above).
Supplementary Figure 4 Ion coordination as seen in MD simulations.
a–e: Inward facing conformation. a. Representative snapshot of a Na+ ion (magenta) in the putative binding site. b. Distribution of Na+-oxygen distances with oxygen atoms from any protein residues or water molecules; only bound Na+ ions were considered, i.e., ions within 3 Å of the carboxyl oxygen atoms of D157 or D156. Oxygen atoms within 3 Å radius form the first coordination shell of the Na+ ion. c. Distribution of observed Na+ coordination numbers n1 (the number of oxygens in the first coordination shell). d. Average contribution of oxygen ligands to n1 for the most frequently observed coordination numbers. e. Time series of the distance of any Na+ ion to the closest oxygen atom in the carboxyl moiety of D157 or D156. Data for protomer A and B from the first 300 ns of three independent simulations were overlaid and shown in different colors. On average, an ion was bound for 70.8% of the simulation time. f–j: Outward facing conformation. The same quantities as in a–e are displayed; however, on average an ion was only bound for 4.2% of the simulation time (j).
Supplementary Figure 5 Positions of the core and dimer domains in MD simulations.
a. Data from 1 μs MD simulations of the outward facing (top) and inward facing (bottom) conformation. Time series and histograms of the z component of the center of mass of the core (blue) and dimer (orange) domain (left panels) indicate equilibrium fluctuations of the domains relative to the instantaneous center of mass of the lipid bilayer. Only protomer A is shown; time series for protomer B are similar. Snapshots from MD simulations show the relative positions of the domains (blue, orange) relative to the membrane (white POPE and gray POPG lipids). b. The z component of the centre of mass of the core and dimer domain in equilibrium MD simulations of outward facing (OF) and inward facing (IF) NapA was used to characterize domain motions. Protomer A (chain A, solid line) and protomer B (chain B, broken line) are shown separately. Left: distribution of the difference in z coordinate relative to the instantaneous center of mass of the lipid bilayer between OF and IF simulations. Right: distribution of the difference in z coordinate relative to the instantaneous center of mass of the dimer domain between OF and IF simulations. The dashed vertical line indicates the value obtained from comparing the LCP OF structure with the disulfide-linked IF structure. c. Distributions of domain positions from two additional independent repeat simulations of 300 ns length.
Supplementary Figure 6 The mobile hinge regions that link the core and dimer domains in NapA.
a. Ribbon representation of outward-facing NapA (dimer light orange; core light-blue). The linking helix TM6 is shown in grey and the mobile connecting hinge regions shown in pink. The position of flanking hinge glycine residues are labelled and shown as pink spheres. b. Ribbon representation of the high-resolution outward-facing (LCP) NapA structure rendered by b-factors scale (rainbow colour scale and tube diameter). The position of flanking hinge glycine residues are labelled and shown as pink spheres.
Supplementary Figure 7 Location of bound (LCP) lipid and comparison of the extent of core movements between NapA and MjNhaP1 structures.
a. Outward-facing NapA, as viewed on a right-angle from the membrane plane, is represented as an electrostatic surface for the dimer domain and as transparent cartoon helices in blue for the core domain. Non-protein density (2FoFc omit map at 1.5σ level) at core-dimer domain interface was fitted as modified monolein MAG7.7 (green sticks). b. The inward-facing structures of NapA (left: dimer light-orange and core blue) and MjNhaP1 (right: dimer light-orange and core blue; PDB 4CZB) were superimposed on the dimer domain of their respective outward-facing structures (shown in white; PDB 4D0A MjNhaP1 and PDB 5BZ3 NapA). The vertical distance between the respective ion-binding aspartates D157 in NapA and D161 in MjNhaP1 are shown as an inset below.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1 and 2 (PDF 2608 kb)
Supplementary Data Set 1
Uncropped SDS-gels used to assess disulfide-bond formation of NapA cysteine mutants (PDF 8561 kb)
The elevator alternating-access mechanism of the Na+ /H+ antiporter NapA
Video showing the morph between the outwardfacing and inward-facing NapA crystal structures after superposition against the dimer domains. The colouring as in Fig. 1a except for half-helices TM11b and TM4b shown in orange and magenta, respectively. The strictly conserved ion-binding aspartate D157 is shown in stick form. View is from the side. (MP4 8172 kb)
The elevator alternating-access mechanism of the Na+ /H+ antiporter NapA
As in Video 1 viewed from the extracellular surface (MP4 8185 kb)
Rights and permissions
About this article
Cite this article
Coincon, M., Uzdavinys, P., Nji, E. et al. Crystal structures reveal the molecular basis of ion translocation in sodium/proton antiporters. Nat Struct Mol Biol 23, 248–255 (2016). https://doi.org/10.1038/nsmb.3164
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3164
This article is cited by
-
Ion and lipid orchestration of secondary active transport
Nature (2024)
-
Architecture and autoinhibitory mechanism of the plasma membrane Na+/H+ antiporter SOS1 in Arabidopsis
Nature Communications (2023)
-
Crystal structure of the Na+/H+ antiporter NhaA at active pH reveals the mechanistic basis for pH sensing
Nature Communications (2022)
-
Structure and mechanism of the human NHE1-CHP1 complex
Nature Communications (2021)
-
Progress in Structural Biology of Solute Carriers
Current Molecular Biology Reports (2021)