Abstract
Mutations in TBC1D24 cause severe epilepsy and DOORS syndrome, but the molecular mechanisms underlying these pathologies are unresolved. We solved the crystal structure of the TBC domain of the Drosophila ortholog Skywalker, revealing an unanticipated cationic pocket conserved among TBC1D24 homologs. Cocrystallization and biochemistry showed that this pocket binds phosphoinositides phosphorylated at the 4 and 5 positions. The most prevalent patient mutations affect the phosphoinositide-binding pocket and inhibit lipid binding. Using in vivo photobleaching of Skywalker-GFP mutants, including pathogenic mutants, we showed that membrane binding via this pocket restricts Skywalker diffusion in presynaptic terminals. Additionally, the pathogenic mutations cause severe neurological defects in flies, including impaired synaptic-vesicle trafficking and seizures, and these defects are reversed by genetically increasing synaptic PI(4,5)P2 concentrations through synaptojanin mutations. Hence, we discovered that a TBC domain affected by clinical mutations directly binds phosphoinositides through a cationic pocket and that phosphoinositide binding is critical for presynaptic function.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Corbett, M.A. et al. A focal epilepsy and intellectual disability syndrome is due to a mutation in TBC1D24. Am. J. Hum. Genet. 87, 371–375 (2010).
Falace, A. et al. TBC1D24, an ARF6-interacting protein, is mutated in familial infantile myoclonic epilepsy. Am. J. Hum. Genet. 87, 365–370 (2010).
Guven, A. & Tolun, A. TBC1D24 truncating mutation resulting in severe neurodegeneration. J. Med. Genet. 50, 199–202 (2013).
Milh, M. et al. Novel compound heterozygous mutations in TBC1D24 cause familial malignant migrating partial seizures of infancy. Hum. Mutat. 34, 869–872 (2013).
Muona, M. et al. A recurrent de novo mutation in KCNC1 causes progressive myoclonus epilepsy. Nat. Genet. 47, 39–46 (2015).
Poulat, A.-L. et al. Homozygous TBC1D24 mutation in two siblings with familial infantile myoclonic epilepsy (FIME) and moderate intellectual disability. Epilepsy Res. 111, 72–77 (2015).
Azaiez, H. et al. TBC1D24 mutation causes autosomal-dominant nonsyndromic hearing loss. Hum. Mutat. 35, 819–823 (2014).
Bakhchane, A. et al. Recessive TBC1D24 mutations are frequent in Moroccan non-syndromic hearing loss pedigrees. PLoS One 10, e0138072 (2015).
Stražišar, B.G., Neubauer, D., Paro Panjan, D. & Writzl, K. Early-onset epileptic encephalopathy with hearing loss in two siblings with TBC1D24 recessive mutations. Eur. J. Paediatr. Neurol. 19, 251–256 (2015).
Zhang, L., Hu, L., Chai, Y. & Pang, X. A dominant mutation in the stereocilia-expressing gene TBC1D24 is a probable cause for non-syndromic hearing impairment. Hum. Mutat. 35, 814–818 (2014).
Doummar, D. et al. A novel homozygous TBC1D24 mutation causing multifocal myoclonus with cerebellar involvement. Mov. Disord. 30, 1431–1432 (2015).
Campeau, P.M. & Hennekam, R.C. DOORS syndrome: phenotype, genotype and comparison with Coffin-Siris syndrome. Am. J. Med. Genet. C. Semin. Med. Genet. 166C, 327–332 (2014).
Campeau, P.M. et al. The genetic basis of DOORS syndrome: an exome-sequencing study. Lancet Neurol. 13, 44–58 (2014).
Mucha, B.E., Hennekam, R.C., Sisodiya, S. & Campeau, P.M. TBC1D24-related disorders. in GeneReviews (eds. Pagon, R.A. et al.) (University of Washington, Seattle, 2015).
Frasa, M.A., Koessmeier, K.T., Ahmadian, M.R. & Braga, V.M. Illuminating the functional and structural repertoire of human TBC/RABGAPs. Nat. Rev. Mol. Cell Biol. 13, 67–73 (2012).
Stenmark, H. Rab GTPases as coordinators of vesicle traffic. Nat. Rev. Mol. Cell Biol. 10, 513–525 (2009).
Fukuda, M. TBC proteins: GAPs for mammalian small GTPase Rab? Biosci. Rep. 31, 159–168 (2011).
Gavriljuk, K. et al. Catalytic mechanism of a mammalian Rab·RabGAP complex in atomic detail. Proc. Natl. Acad. Sci. USA 109, 21348–21353 (2012).
Pan, X., Eathiraj, S., Munson, M. & Lambright, D.G. TBC-domain GAPs for Rab GTPases accelerate GTP hydrolysis by a dual-finger mechanism. Nature 442, 303–306 (2006).
Uytterhoeven, V., Kuenen, S., Kasprowicz, J., Miskiewicz, K. & Verstreken, P. Loss of skywalker reveals synaptic endosomes as sorting stations for synaptic vesicle proteins. Cell 145, 117–132 (2011).
Chaineau, M., Ioannou, M.S. & McPherson, P.S. Rab35: GEFs, GAPs and effectors. Traffic 14, 1109–1117 (2013).
Cherfils, J. Arf GTPases and their effectors: assembling multivalent membrane-binding platforms. Curr. Opin. Struct. Biol. 29, 67–76 (2014).
Chesneau, L. et al. An ARF6/Rab35 GTPase cascade for endocytic recycling and successful cytokinesis. Curr. Biol. 22, 147–153 (2012).
Fernandes, A.C. et al. Reduced synaptic vesicle protein degradation at lysosomes curbs TBC1D24/sky-induced neurodegeneration. J. Cell Biol. 207, 453–462 (2014).
Cremona, O. et al. Essential role of phosphoinositide metabolism in synaptic vesicle recycling. Cell 99, 179–188 (1999).
Verstreken, P. et al. Synaptojanin is recruited by endophilin to promote synaptic vesicle uncoating. Neuron 40, 733–748 (2003).
Park, S.-Y., Jin, W., Woo, J.R. & Shoelson, S.E. Crystal structures of human TBC1D1 and TBC1D4 (AS160) RabGTPase-activating protein (RabGAP) domains reveal critical elements for GLUT4 translocation. J. Biol. Chem. 286, 18130–18138 (2011).
Ashkenazy, H., Erez, E., Martz, E., Pupko, T. & Ben-Tal, N. ConSurf 2010: calculating evolutionary conservation in sequence and structure of proteins and nucleic acids. Nucleic Acids Res. 38, W529–W533 (2010).
Rehman, A.U. et al. Mutations in TBC1D24, a gene associated with epilepsy, also cause nonsyndromic deafness DFNB86. Am. J. Hum. Genet. 94, 144–152 (2014).
Falace, A. et al. TBC1D24 regulates neuronal migration and maturation through modulation of the ARF6-dependent pathway. Proc. Natl. Acad. Sci. USA 111, 2337–2342 (2014).
Di Paolo, G. & De Camilli, P. Phosphoinositides in cell regulation and membrane dynamics. Nature 443, 651–657 (2006).
Czech, M.P. PIP2 and PIP3: complex roles at the cell surface. Cell 100, 603–606 (2000).
Khuong, T.M. et al. Synaptic PI(3,4,5)P3 is required for Syntaxin1A clustering and neurotransmitter release. Neuron 77, 1097–1108 (2013).
Niesen, F.H., Berglund, H. & Vedadi, M. The use of differential scanning fluorimetry to detect ligand interactions that promote protein stability. Nat. Protoc. 2, 2212–2221 (2007).
Bottomley, M.J., Salim, K. & Panayotou, G. Phospholipid-binding protein domains. Biochim. Biophys. Acta 1436, 165–183 (1998).
Bravo, J. et al. The crystal structure of the PX domain from p40(phox) bound to phosphatidylinositol 3-phosphate. Mol. Cell 8, 829–839 (2001).
Saheki, Y. & De Camilli, P. Synaptic vesicle endocytosis. Cold Spring Harb. Perspect. Biol. 4, a005645 (2012).
Salkoff, L. & Kelly, L. Temperature-induced seizure and frequency-dependent neuromuscular block in a ts mutant of Drosophila. Nature 273, 156–158 (1978).
Titus, S.A., Warmke, J.W. & Ganetzky, B. The Drosophila erg K+ channel polypeptide is encoded by the seizure locus. J. Neurosci. 17, 875–881 (1997).
Krans, J.L., Rivlin, P.K. & Hoy, R.R. Demonstrating the temperature sensitivity of synaptic transmission in a Drosophila mutant. J. Undergrad. Neurosci. Educ. 4, A27–A33 (2005).
Rosato, E. & Kyriacou, C.P. Analysis of locomotor activity rhythms in Drosophila. Nat. Protoc. 1, 559–568 (2006).
Harris, T.W., Hartwieg, E., Horvitz, H.R. & Jorgensen, E.M. Mutations in synaptojanin disrupt synaptic vesicle recycling. J. Cell Biol. 150, 589–600 (2000).
Verstreken, P. et al. Tweek, an evolutionarily conserved protein, is required for synaptic vesicle recycling. Neuron 63, 203–215 (2009).
Sklan, E.H. et al. TBC1D20 is a Rab1 GTPase-activating protein that mediates hepatitis C virus replication. J. Biol. Chem. 282, 36354–36361 (2007).
Moravcevic, K., Oxley, C.L. & Lemmon, M.A. Conditional peripheral membrane proteins: facing up to limited specificity. Structure 20, 15–27 (2012).
Kutateladze, T.G. Translation of the phosphoinositide code by PI effectors. Nat. Chem. Biol. 6, 507–513 (2010).
Lemmon, M.A. Membrane recognition by phospholipid-binding domains. Nat. Rev. Mol. Cell Biol. 9, 99–111 (2008).
Slabbaert, J.R., Khuong, T.M. & Verstreken, P. Phosphoinositides at the neuromuscular junction of Drosophila melanogaster: a genetic approach. Methods Cell Biol. 108, 227–247 (2012).
Marley, J., Lu, M. & Bracken, C. A method for efficient isotopic labeling of recombinant proteins. J. Biomol. NMR 20, 71–75 (2001).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Chen, V.B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Dolinsky, T.J. et al. PDB2PQR: expanding and upgrading automated preparation of biomolecular structures for molecular simulations. Nucleic Acids Res. 35, W522–W525 (2007).
Baker, N.A., Sept, D., Joseph, S., Holst, M.J. & Mccammon, J.A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl. Acad. Sci. USA 98, 10037–10041 (2001).
Jo, S., Kim, T., Iyer, V.G. & Im, W. CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comput. Chem. 29, 1859–1865 (2008).
Altschul, S.F., Gish, W., Miller, W., Myers, E.W. & Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Papadopoulos, J.S. & Agarwala, R. COBALT: constraint-based alignment tool for multiple protein sequences. Bioinformatics 23, 1073–1079 (2007).
Di Tommaso, P. et al. T-Coffee: a web server for the multiple sequence alignment of protein and RNA sequences using structural information and homology extension. Nucleic Acids Res. 39, W13–W17 (2011).
Gouet, P., Robert, X. & Courcelle, E. ESPript/ENDscript: extracting and rendering sequence and 3D information from atomic structures of proteins. Nucleic Acids Res. 31, 3320–3323 (2003).
Bischof, J., Maeda, R.K., Hediger, M., Karch, F. & Basler, K. An optimized transgenesis system for Drosophila using germ-line-specific phiC31 integrases. Proc. Natl. Acad. Sci. USA 104, 3312–3317 (2007).
Gibson, D.G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Acknowledgements
We thank the staff at the beamlines Proxima 2 of Soleil (France) and ID23-1 of ESRF (France) for assistance during data collection. We thank E. Lauwers and J. McInnes for advice on lipid biology and members of the Versées and Verstreken laboratories for comments. This work was supported by the Fonds voor Wetenschappelijk Onderzoek (J.P., V.U., P.V. and W.V.), Strategic Research Program Financing from the VUB (W.V.) and KU Leuven (P.V.), the Hercules foundation (W.V. and P.V.), BioStruct-X from the European Community's Seventh Framework Programme (W.V.), the ERC (P.V.), a Methusalem grant from the Flemish government (P.V.), the Belgian Science Policy (P.V.), Agentschap voor Innovatie door Wetenschap en Technologie (I.M.), the Belgian American Educational Foundation (K.L.) and VIB (P.V.).
Author information
Authors and Affiliations
Contributions
B.F. and J.P. solved the structures, produced and purified proteins, performed biochemical experiments, analyzed data and wrote the manuscript; K.L. performed biochemical and in vivo experiments, analyzed data and wrote the manuscript; C.D.K. assisted with biochemistry; I.M., S.K. and V.U. performed in vivo experiments; J.S. performed transmission electron microscopy; P.V. and W.V. conceived and supervised the work, contributed to data interpretation and wrote the manuscript. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Different crystal forms obtained for Sky1–353.
(a) Crystal form 1 obtained in the presence of 20% PEG 3350 and 0.2 M ammonium citrate tribasic pH 7.0.
(b) Crystal form 1 of the selenomethionine-labelled Sky1-353 obtained in the presence of 20% PEG 3350 and 0.2 M ammonium citrate tribasic pH 7.0
(c) Crystal form 2 obtained in the presence of 25% PEG 1500 and 0.1 M succinate/phosphate/glycine pH 7.0. This crystal form was used to obtain the Sky1-353-IP3 crystal structure after soaking with 5 mM IP3.
Supplementary Figure 2 Sequence alignment of Sky1–353 with the TBC domain of human TBC1D24 and representative RabGAP proteins for which the crystal structure has been reported.
The residues corresponding to the arginine and glutamine fingers in conventional TBC Rab-GAP proteins are highlighted in green. The residues corresponding to the cationic pocket residues in Sky are highlighted in blue. Residues corresponding to patient mutations are red, with those located in the cationic pocket additionally indicated by a red star. The α-helices determined from the Sky1-353 structure are indicated above the alignment, with the numbering corresponding to the numbering used in the Gyp1 structure. The sequences used are human TBC1D24 (UniProt: Q9ULP9), Gyp1 from Saccharomyces cerevisiae (UniProt: Q08484), human TBC1D1 (UniProt: Q86TI0), human TBC1D4 (Uniprot: O60343), human TBC1D7 (UniProt: Q9P0N9), human TBC1D11 (UniProt: Q9Y3P9), human TBC1D14 (UniProt: Q9P2M4), human TBC1D18 (UniProt: Q5R372), human TBC1D20 (UniProt: Q96BZ9), human TBC1D22A (UniProt: Q8WUA7), human TBC1D22B (UniProt: Q9NU19), CrfRabGAP from Chlamydomonas reinhardtii (UniProt: A8JCA4).
Supplementary Figure 3 Surface electrostatics of Sky1–353 in comparison to other TBC domains.
The electrostatic potential mapped on the solvent accessible surface of Sky1-353 in the first panel shows the cationic pocket located on the opposite side of the GTPase binding region. The electrostatic potential surfaces of the 11 other TBC domains deposited in the PDB are shown in exactly the same orientation. None of these proteins seems to harbor a well-defined cationic pocket similar to Sky1-353. The structures of the TBC domains shown in this figure are: Gyp1 (pdb 1FKM; Rak, A. et al., EMBO J. 19, 5105–13, 2000), TBC1D1 (pdb 3QYE; Park, S.-Y. et al., J. Biol. Chem. 286, 18130–8, 2011), TBC1D4 (pdb 3QYB; Park, S.-Y. et al., J. Biol. Chem. 286, 18130–8, 2011), TBC1D7 (pdb 3QWL; unpublished), TBC1D11 (pdb 4NC6; unpublished), TBC1D14 (pdb 2QQ8; unpublished), TBC1D18 (pdb 3HZJ; unpublished), TBC1D20 (pdb 4HL4; Gavriljuk, K. et al., Proc. Natl. Acad. Sci. U.S.A. 109, 21348–53, 2012), TBC1D22A (pdb 2QFZ; unpublished), TBC1D22B (pdb 3DZX; unpublished), CrfRabGAP (pdb 4P17; Bhogaraju, S. & Lorentzen, E. Proteins 82, 2282–7, 2014).
Supplementary Figure 4 Influence of PI(4,5)P2 and salt concentrations on the binding of Sky1–353 to liposomes.
(a) Western blots of liposome flotation assays using wild type Sky1-353 and liposomes (PC:PS) enriched with different concentrations of PI(4,5)P2 (0%, 0.5%, 2% and 5%). n = 2 independent experiments.
(b) Western blots of liposome flotation assays using wild type Sky1-353 and liposomes (PC:PS) enriched with 2% of PI(4,5)P2 in the presence of three different NaCl concentrations (30 mM, 150 mM and 500 mM). n = 2 independent experiments.
Original blots can be found in Supplementary Data Set 1.
Supplementary Figure 5 Comparison of the phosphoinositide-binding pocket of Sky1–353 with other typical phosphoinositide-binding domains.
Structures of representatives of well-established phosphoinositide-binding domains (ENTH, PX, FYVE and PH) in complex with a phosphoinositide head group are shown in electrostatic surface representation and are compared to the Sky1-353 structure in complex with IP3. The boxes show a close-up of the phosphoinositide binding pocket in cartoon representation with the phosphoinositide head group and the interacting residues shown as sticks. The structures that are shown are: the TBC domain of Sky bound to IP3 (this study), the ENTH domain of epsin bound to IP3 (pdb 1H0A; Ford, M. G. J. et al., Nature 419, 361–366, 2002), the PX domain of P40phox bound to dibutanoyl IP2 (pdb 1H6H; Bravo, J. et al., Mol. Cell 8, 829–39, 2001), the FYVE domain of EEA1 bound to IP2 (pdb 1JOC; Dumas, J. J. et al., Mol. Cell 8, 947–58, 2001), the PH domain of PLC-δ1 bound to IP3 (pdb 1MAI; Ferguson, K. M. et al., Cell 83, 1037–1046, 1995) and the PH domain of spectrin bound to IP3 (pdb 1BTN; Hyvönen, M. et al., EMBO J. 14, 4676–85, 1995).
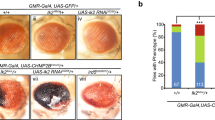
Supplementary Figure 6 Wild-type and mutant Skywalker-GFP traffic to and are present at synaptic terminals
(a) Confocal images of the ventral nerve cord in third instar Drosophila larvae expressing wild type GFP-Sky (SkyWT) or mutant GFP-Sky (SkyR79C, SkyR281C or Sky3Glu). Scale bar for all top panels: 20 μm.
(b) Confocal images of larval neuromuscular junction endplates of animals with genotypes as in (a). (n = 6 animals), scale bar for all panels in (b): 20 μm. GFP intensities between the GFP-Sky mutants is not significantly different.
(c) Quantification of third-instar NMJ GFP intensity at the membrane in single confocal sections. Mean Fluorescence Intensity between the genotypes GFP-SkyWT, GFP-SkyR79C and GFP-Sky3Glu is similar. Error bars: mean ± s.e.m. (n = 6) P = 0.446, ns: not significant by ANOVA, Dunnett’s.
(d) Quantification of Western blot band intensity of GFP-Sky and GFP-Syt in protein isolations from adult fly heads. Intensities are normalized to Syntaxin control levels. GFP-Sky WT and mutant levels are similar. Error bars: mean ± s.e.m. (n = 3) P = 0.681, ns: not significant by ANOVA, Dunnett’s. The levels of Synaptotagmin-GFP (Syt) are shown as a control.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 (PDF 1336 kb)
Supplementary Data Set 1
Uncropped blots for Figure 3 and Supplementary Figure 4. (PDF 3079 kb)
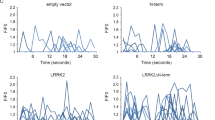
Seizure asssay of Drosophila carrying mutations in the cationic pocket of Sky.
Movie of behavior of adult sky1/2 Drosophila males (yw eyFLP/Y; FRT40A sky1:FRT40A sky2) expressing GFP-SkyWT, GFP-SkyR79C or GFP-Sky3Glu using nSybGal4, following 10 seconds of vortex stimulation. (MP4 1269 kb)
Rights and permissions
About this article
Cite this article
Fischer, B., Lüthy, K., Paesmans, J. et al. Skywalker-TBC1D24 has a lipid-binding pocket mutated in epilepsy and required for synaptic function. Nat Struct Mol Biol 23, 965–973 (2016). https://doi.org/10.1038/nsmb.3297
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3297
This article is cited by
-
Parkinsonism mutations in DNAJC6 cause lipid defects and neurodegeneration that are rescued by Synj1
npj Parkinson's Disease (2023)
-
Addition of an affected family member to a previously ascertained autosomal recessive nonsyndromic hearing loss pedigree and systematic phenotype-genotype analysis of splice-site variants in MYO15A
BMC Medical Genomics (2022)
-
The Putative Drosophila TMEM184B Ortholog Tmep Ensures Proper Locomotion by Restraining Ectopic Firing at the Neuromuscular Junction
Molecular Neurobiology (2022)
-
TBC1D24 emerges as an important contributor to progressive postlingual dominant hearing loss
Scientific Reports (2021)
-
TBC1D24 regulates axonal outgrowth and membrane trafficking at the growth cone in rodent and human neurons
Cell Death & Differentiation (2019)