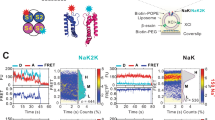
Abstract
Crystallography has provided invaluable insights regarding ion-channel selectivity and gating, but to advance understanding to a new level, dynamic views of channel structures within membranes are essential. We labeled tetrameric KirBac1.1 potassium channels with single donor and acceptor fluorophores at different sites and then examined structural dynamics within lipid membranes by single-molecule fluorescence resonance energy transfer (FRET). We found that the extracellular region is structurally rigid in both closed and open states, whereas the N-terminal slide helix undergoes marked conformational fluctuations. The cytoplasmic C-terminal domain fluctuates between two major structural states, both of which become less dynamic and move away from the pore axis and away from the membrane in closed channels. Our results reveal mobile and rigid conformations of functionally relevant KirBac1.1 channel motifs, implying similar dynamics for similar motifs in eukaryotic Kir channels and in cation channels in general.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Neher, E. & Sakmann, B. Single-channel currents recorded from membrane of denervated frog muscle fibres. Nature 260, 799–802 (1976).
Colquhoun, D. & Sigworth, F. in Single-channel Recording 483–587 (Springer, 1995).
Cao, E., Liao, M., Cheng, Y. & Julius, D. TRPV1 structures in distinct conformations reveal activation mechanisms. Nature 504, 113–118 (2013).
Whorton, M.R. & MacKinnon, R. Crystal structure of the mammalian GIRK2 K+ channel and gating regulation by G proteins, PIP2, and sodium. Cell 147, 199–208 (2011).
Cuello, L.G., Jogini, V., Cortes, D.M. & Perozo, E. Structural mechanism of C-type inactivation in K+ channels. Nature 466, 203–208 (2010).
Cuello, L.G. et al. Structural basis for the coupling between activation and inactivation gates in K+ channels. Nature 466, 272–275 (2010).
Jiang, Y. et al. Crystal structure and mechanism of a calcium-gated potassium channel. Nature 417, 515–522 (2002).
Zhou, Y., Morais-Cabral, J.H., Kaufman, A. & MacKinnon, R. Chemistry of ion coordination and hydration revealed by a K+ channel–Fab complex at 2.0 A resolution. Nature 414, 43–48 (2001).
Nordengren, J. et al. Differential localization and expression of urokinase plasminogen activator (uPA), its receptor (uPAR), and its inhibitor (PAI-1) mRNA and protein in endometrial tissue during the menstrual cycle. Mol. Hum. Reprod. 10, 655–663 (2004).
Bernèche, S. & Roux, B. Energetics of ion conduction through the K+ channel. Nature 414, 73–77 (2001).
Nichols, C.G. KATP channels as molecular sensors of cellular metabolism. Nature 440, 470–476 (2006).
Enkvetchakul, D., Jeliazkova, I., Bhattacharyya, J. & Nichols, C.G. Control of inward rectifier K channel activity by lipid tethering of cytoplasmic domains. J. Gen. Physiol. 130, 329–334 (2007).
Männikkö, R. et al. Interaction between mutations in the slide helix of Kir6.2 associated with neonatal diabetes and neurological symptoms. Hum. Mol. Genet. 19, 963–972 (2010).
Kuo, A., Domene, C., Johnson, L.N., Doyle, D.A. & Vénien-Bryan, C. Two different conformational states of the KirBac3.1 potassium channel revealed by electron crystallography. Structure 13, 1463–1472 (2005).
Lu, Z., Klem, A.M. & Ramu, Y. Coupling between voltage sensors and activation gate in voltage-gated K+ channels. J. Gen. Physiol. 120, 663–676 (2002).
Long, S.B., Campbell, E.B. & Mackinnon, R. Voltage sensor of Kv1.2: structural basis of electromechanical coupling. Science 309, 903–908 (2005).
Hansen, S.B., Tao, X. & MacKinnon, R. Structural basis of PIP2 activation of the classical inward rectifier K+ channel Kir2.2. Nature 477, 495–498 (2011).
Bavro, V.N. et al. Structure of a KirBac potassium channel with an open bundle crossing indicates a mechanism of channel gating. Nat. Struct. Mol. Biol. 19, 158–163 (2012).
Hibino, H. et al. Inwardly rectifying potassium channels: their structure, function, and physiological roles. Physiol. Rev. 90, 291–366 (2010).
Cheng, W.W., Enkvetchakul, D. & Nichols, C.G. KirBac1.1: it’s an inward rectifying potassium channel. J. Gen. Physiol. 133, 295–305 (2009).
Kuo, A. et al. Crystal structure of the potassium channel KirBac1.1 in the closed state. Science 300, 1922–1926 (2003).
Wang, S., Lee, S.J., Heyman, S., Enkvetchakul, D. & Nichols, C.G. Structural rearrangements underlying ligand-gating in Kir channels. Nat. Commun. 3, 617 (2012).
Suh, B.C. & Hille, B. PIP2 is a necessary cofactor for ion channel function: how and why? Annu. Rev. Biophys. 37, 175–195 (2008).
Xie, L.H., John, S.A., Ribalet, B. & Weiss, J.N. Activation of inwardly rectifying potassium (Kir) channels by phosphatidylinosital-4,5-bisphosphate (PIP2): interaction with other regulatory ligands. Prog. Biophys. Mol. Biol. 94, 320–335 (2007).
Logothetis, D.E., Jin, T., Lupyan, D. & Rosenhouse-Dantsker, A. Phosphoinositide-mediated gating of inwardly rectifying K(+) channels. Pflugers Arch. 455, 83–95 (2007).
Enkvetchakul, D., Jeliazkova, I. & Nichols, C.G. Direct modulation of Kir channel gating by membrane phosphatidylinositol 4,5-bisphosphate. J. Biol. Chem. 280, 35785–35788 (2005).
Ha, T. et al. Probing the interaction between two single molecules: fluorescence resonance energy transfer between a single donor and a single acceptor. Proc. Natl. Acad. Sci. USA 93, 6264–6268 (1996).
Roy, R., Hohng, S. & Ha, T. A practical guide to single-molecule FRET. Nat. Methods 5, 507–516 (2008).
Zhao, Y. et al. Single-molecule dynamics of gating in a neurotransmitter transporter homologue. Nature 465, 188–193 (2010).
Akyuz, N., Altman, R.B., Blanchard, S.C. & Boudker, O. Transport dynamics in a glutamate transporter homologue. Nature 502, 114–118 (2013).
Erkens, G.B., Hänelt, I., Goudsmits, J.M., Slotboom, D.J. & van Oijen, A.M. Unsynchronised subunit motion in single trimeric sodium-coupled aspartate transporters. Nature 502, 119–123 (2013).
Wang, Y. et al. Single molecule FRET reveals pore size and opening mechanism of a mechano-sensitive ion channel. eLife 3, e01834 (2014).
Vafabakhsh, R., Levitz, J. & Isacoff, E.Y. Conformational dynamics of a class C G-protein-coupled receptor. Nature 524, 497–501 (2015).
Roux, B. Ion conduction and selectivity in K(+) channels. Annu. Rev. Biophys. Biomol. Struct. 34, 153–171 (2005).
Raghuraman, H., Islam, S.M., Mukherjee, S., Roux, B. & Perozo, E. Dynamics transitions at the outer vestibule of the KcsA potassium channel during gating. Proc. Natl. Acad. Sci. USA 111, 1831–1836 (2014).
Payandeh, J., Scheuer, T., Zheng, N. & Catterall, W.A. The crystal structure of a voltage-gated sodium channel. Nature 475, 353–358 (2011).
Xiao, J., Zhen, X.G. & Yang, J. Localization of PIP2 activation gate in inward rectifier K+ channels. Nat. Neurosci. 6, 811–818 (2003).
Lee, S.J. et al. Secondary anionic phospholipid binding site and gating mechanism in Kir2.1 inward rectifier channels. Nat. Commun. 4, 2786 (2013).
McCusker, E.C. et al. Structure of a bacterial voltage-gated sodium channel pore reveals mechanisms of opening and closing. Nat. Commun. 3, 1102 (2012).
Bollepalli, M.K. et al. State-dependent network connectivity determines gating in a K+ channel. Structure 22, 1037–1046 (2014).
Clarke, O.B. et al. Domain reorientation and rotation of an intracellular assembly regulate conduction in Kir potassium channels. Cell 141, 1018–1029 (2010).
Joo, C. & Ha, T. Prism-type total internal reflection microscopy for single-molecule FRET. Cold Spring Harb. Protoc. 2012 Pdb.prot072041 (2012).
Enkvetchakul, D. et al. Functional characterization of a prokaryotic Kir channel. J. Biol. Chem. 279, 47076–47080 (2004).
Wang, S., Alimi, Y., Tong, A., Nichols, C.G. & Enkvetchakul, D. Differential roles of blocking ions in KirBac1.1 tetramer stability. J. Biol. Chem. 284, 2854–2860 (2009).
Joo, C. & Ha, T. Preparing sample chambers for single-molecule FRET. Cold Spring Harb. Protoc. 2012, 1104–1108 (2012).
Acknowledgements
Financial support was provided by US National Institutes of Health (NIH) grant HL54171 (C.G.N.) and US National Science Foundation grant PHY1430124 (T.H.). T.H. is supported as an investigator of the Howard Hughes Medical Institute. W.F.B. was supported by NIH grants T32 HL007275 and T32 HL125241.
Author information
Authors and Affiliations
Contributions
S.W., R.V., T.H. and C.G.N. conceived and designed the experiments; S.W. and W.F.B. performed the electrophysiological studies; S.W. designed, constructed, purified and labeled the protein samples with fluorophore; W.F.B. analyzed the single-channel recordings; and S.W. and R.V. collected and analyzed the smFRET data. The paper was written by S.W. and C.G.N. and edited by R.V., W.F.B. and T.H.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Purified KirBac1.1 mutants for smFRET experiments.
(A) Gel filtration profiles of KirBac1.1 mutants labeled with Alexa Fluor 555 and Alexa Fluor 647 c2 maleimide. Purified KirBac1.1 proteins (1 mg/ml) were reacted with 1:1 mixer of Alexa Fluor 555 and Alexa Fluor 647 c2 maleimide at protein/fluorophore ratio of 1:4 for 2 hrs under room temperature. The labeled mixture was loaded onto a metal affinity column and the non-covalently attached fluorophores were removed by extensive washes. The labeled proteins were eluted using imidazole and then loaded onto a gel filtration column. The central fractions (between the dash lines) of tetramer peaks were collected for functional study (rubidium flux assay) or smFRET experiments. (B) Channel function assay of KirBac1.1 mutants labeling with Alexa Fluoro 555 and Alexa Fluoro 647 by rubidium flux. Mutant proteins were labeled with 1:1 mixer of Alexa Fluoro 555 and Alexa Fluoro 647 and reconstituted into liposomes at a protein:lipid ratio of 1:100. The lipids used for reconstitution were POPE and POPG (3:1), with or without 1% PIP2. The intraliposomal solution contained 20 mM HEPES, 450 mM KCl, 1 mM EDTA, 1 mM EGTA, 4 mM NMDG, pH7.5. The extraliposomal solution contained 20 mM HEPES, 400 mM Sorbitol, 1 mM EDTA, 1 mM EGTA, 4 mM NMDG, pH7.5. The rubidium assay was performed following the protocol described previously (19), and all rubidium uptakes were normalized against WT. The data are presented as mean±se, n=3. The labeling efficiencies of protein samples were determined by a Nanodrop 1000 spectrophotometer according to the protocol provided with the Alexa Fluor 555 and 647 dye package. The labeling efficiencies were location-dependent and changed slightly for different batches. For construct A45C-WT and T120C-WT constructs, the typical labeling efficiencies were 0.6~0.7; for W48C-WT construct, the labeling efficiency was ~0.5; for A167C-WT and A273C-WT, the labeling efficiencies were typically >0.9. (C) The specificity of fluorophore labeling in KirBac1.1 cysteine mutants examined by SDS-PAGE. The labeled protein samples were separated by 4-20% SDS-PAGE, and fluorescence images were acquired with a Kodak fluorescence scanner. For visualizing Alexa Fluor 555, the excitation light was 535 nm and emission images were collected with a 600nm long-pass filter; for Alexa Fluor 647, the excitation light was 637 nm, and emission images were collected with a 670 nm long-pass filter. The protein was visualized by Coomassie Brilliant Blue R-250 staining and the images were acquired with a document scanner.
Supplementary Figure 2 Representative smFRET traces.
Representative smFRET traces of T120C-WT (A), A45C-WT (B) and A273C-WT (C) sample without or with PIP2.
Supplementary Figure 3 smFRET histograms and contour plots of KirBac1.1 mutants at 30-ms and 100-ms time resolution.
(A) FRET histograms and contour plots of T120C-WT (n=53 and 40 for control and PIP2 conditions, respectively) and A45C-WT (n=218 and 235 for control and PIP2 conditions, respectively) mutants with single molecule imaging data of 30 ms time resolution. (B) FRET histograms and contour plots of W48C-WT (n=100 and 46 for control and PIP2 conditions, respectively) and F167C-WT mutants (n=333 and 254 for control and PIP2 conditions, respectively).
Supplementary Figure 4 Ensemble FRET measurements of KirBac1.1 mutants labeled at different structural motifs in liposomes with or without PIP2.
KirBac1.1 mutants were labeled with EDANS/DABCYL-plus (A) or AlexaFluor-488/QSY7 FRET pair (B) and reconstituted into liposomes (POPE:POPG = 3:1) with or without 1% PIP2. Ensemble FRET measurements were performed following the protocols as described previously (Wang, S. et al., 2012, Nature communications 3, 617). The data were presented as mean± se, n = 6 (*p<0.05, **p<0.01).
Supplementary Figure 5 Representative smFRET traces.
Representative smFRET traces of T120C/A270C-WT-WT-WT (A), T120C-A270C-WT-WT (B) and T120C-WT-A270C -WT (C) samples without or with PIP2.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Tables 1 and 2 (PDF 1077 kb)
Rights and permissions
About this article
Cite this article
Wang, S., Vafabakhsh, R., Borschel, W. et al. Structural dynamics of potassium-channel gating revealed by single-molecule FRET. Nat Struct Mol Biol 23, 31–36 (2016). https://doi.org/10.1038/nsmb.3138
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3138
This article is cited by
-
Conformational plasticity of NaK2K and TREK2 potassium channel selectivity filters
Nature Communications (2023)
-
Membrane Cholesterol Interactions with Proteins in Hypercholesterolemia-Induced Endothelial Dysfunction
Current Atherosclerosis Reports (2023)
-
cAMP binding to closed pacemaker ion channels is non-cooperative
Nature (2021)
-
Conformational equilibrium shift underlies altered K+ channel gating as revealed by NMR
Nature Communications (2020)
-
New Structural insights into Kir channel gating from molecular simulations, HDX-MS and functional studies
Scientific Reports (2020)