Abstract
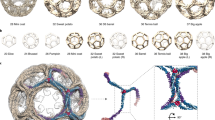
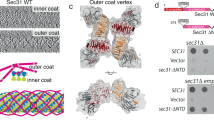
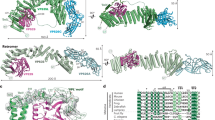
COPII vesicles transport proteins from the endoplasmic reticulum to the Golgi apparatus. Previous COPII-cage cryo-EM structures lacked the resolution necessary to determine the residues of Sec13 and Sec31 that mediate assembly and flexibility of the COPII cage. Here we present a 12-Å structure of the human COPII cage, where the tertiary structure of Sec13 and Sec31 is clearly identifiable. We employ this structure and a homology model of the Sec13–Sec31 complex to create a reliable pseudoatomic model of the COPII cage. We combined this model with hydrogen/deuterium-exchange MS analysis to characterize four distinct contact regions at the vertices of the COPII cage. Furthermore, we found that the two-fold symmetry of the Sec31 dimeric region in Sec13–Sec31 is broken upon cage formation and that the resulting hinge is essential to form the proper edge geometry in COPII cages.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Zanetti, G., Pahuja, K.B., Studer, S., Shim, S. & Schekman, R. COPII and the regulation of protein sorting in mammals. Nat. Cell Biol. 14, 20–28 (2011).
Lee, M.C. et al. Sar1p N-terminal helix initiates membrane curvature and completes the fission of a COPII vesicle. Cell 122, 605–617 (2005).
Stagg, S.M. et al. Structure of the Sec13/31 COPII coat cage. Nature 439, 234–238 (2006).
Fath, S., Mancias, J.D., Bi, X. & Goldberg, J. Structure and organization of coat proteins in the COPII cage. Cell 129, 1325–1336 (2007).
Bacia, K. et al. Multibudded tubules formed by COPII on artificial liposomes. Sci Rep 1, 17 (2011).
Elrod-Erickson, M.J. & Kaiser, C.A. Genes that control the fidelity of endoplasmic reticulum to Golgi transport identified as suppressors of vesicle budding mutations. Mol. Biol. Cell 7, 1043–1058 (1996).
Copic, A., Latham, C.F., Horlbeck, M.A., D'Arcangelo, J.G. & Miller, E.A. ER cargo properties specify a requirement for COPII coat rigidity mediated by Sec13p. Science 335, 1359–1362 (2012).
Jin, L. et al. Ubiquitin-dependent regulation of COPII coat size and function. Nature 482, 495–500 (2012).
Engen, J.R. Analysis of protein conformation and dynamics by hydrogen/deuterium exchange MS. Anal. Chem. 81, 7870–7875 (2009).
Kastner, B. et al. GraFix: sample preparation for single-particle electron cryomicroscopy. Nat. Methods 5, 53–55 (2008).
Ludtke, S.J., Baldwin, P.R. & Chiu, W. EMAN: semiautomated software for high-resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999).
Frank, J. et al. SPIDER and WEB: processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996).
Stagg, S.M. et al. Structural basis for cargo regulation of COPII coat assembly. Cell 134, 474–484 (2008).
Eswar, N. et al. Comparative protein structure modeling using MODELLER. in Curr. Protoc. Protein Sci. Ch. 2, 2.9.1–2.9.31 (Wiley, 2007).
Trabuco, L.G., Villa, E., Mitra, K., Frank, J. & Schulten, K. Flexible fitting of atomic structures into electron microscopy maps using molecular dynamics. Structure 16, 673–683 (2008).
Marshall, A.G., Hendrickson, C.L. & Jackson, G.S. Fourier transform ion cyclotron resonance mass spectrometry: a primer. Mass Spectrom. Rev. 17, 1–35 (1998).
Chalmers, M.J. et al. Probing protein ligand interactions by automated hydrogen/deuterium exchange mass spectrometry. Anal. Chem. 78, 1005–1014 (2006).
Zhang, Q. et al. Epitope mapping of a 95 kDa antigen in complex with antibody by solution-phase amide backbone hydrogen/deuterium exchange monitored by Fourier transform ion cyclotron resonance mass spectrometry. Anal. Chem. 83, 7129–7136 (2011).
Zhang, H.M., Bou-Assaf, G.M., Emmett, M.R. & Marshall, A.G. Fast reversed-phase liquid chromatography to reduce back exchange and increase throughput in H/D exchange monitored by FT-ICR mass spectrometry. J. Am. Soc. Mass Spectrom. 20, 520–524 (2009).
Cravello, L., Lascoux, D. & Forest, E. Use of different proteases working in acidic conditions to improve sequence coverage and resolution in hydrogen/deuterium exchange of large proteins. Rapid Commun. Mass Spectrom. 17, 2387–2393 (2003).
Zhang, X. et al. Near-atomic resolution using electron cryomicroscopy and single-particle reconstruction. Proc. Natl. Acad. Sci. USA 105, 1867–1872 (2008).
Weis, D.D., Engen, J. & Kass, I. Semi-automated data processing of hydrogen exchange mass spectra using HX-Express. J. Am. Soc. Mass Spectrom. 17, 1700–1703 (2006).
Pascal, B.D., Chalmers, M.J., Busby, S.A. & Griffin, P.R.H.D. Desktop: an integrated platform for the analysis and visualization of H/D exchange data. J. Am. Soc. Mass Spectrom. 20, 601–610 (2009).
Kazazic, S. et al. Automated data reduction for hydrogen/deuterium exchange experiments, enabled by high-resolution Fourier transform ion cyclotron resonance mass spectrometry. J. Am. Soc. Mass Spectrom. 21, 550–558 (2010).
Zhang, Z., Zhang, A. & Xiao, G. Improved protein hydrogen/deuterium exchange mass spectrometry platform with fully automated data processing. Anal. Chem. 84, 4942–4949 (2012).
Schaub, T.M. et al. High-performance mass spectrometry: Fourier transform ion cyclotron resonance at 14.5 Tesla. Anal. Chem. 80, 3985–3990 (2008).
Bi, X., Mancias, J.D. & Goldberg, J. Insights into COPII coat nucleation from the structure of Sec23.Sar1 complexed with the active fragment of Sec31. Dev. Cell 13, 635–645 (2007).
Brohawn, S.G., Leksa, N.C., Spear, E.D., Rajashankar, K.R. & Schwartz, T.U. Structural evidence for common ancestry of the nuclear pore complex and vesicle coats. Science 322, 1369–1373 (2008).
Whittle, J.R.R. & Schwartz, T.U. Structure of the Sec13-Sec16 edge element, a template for assembly of the COPII vesicle coat. J. Cell Biol. 190, 347–361 (2010).
Bhattacharya, N., Donnell, O.J. & Stagg, S.M. The structure of the Sec13/31 COPII cage bound to Sec23. J. Mol. Biol. 420, 324–334 (2012).
Lederkremer, G.Z. et al. Structure of the Sec23p/24p and Sec13p/31p complexes of COPII. Proc. Natl. Acad. Sci. USA 98, 10704–10709 (2001).
O'Donnell, J., Maddox, K. & Stagg, S. The structure of a COPII tubule. J. Struct. Biol. 173, 358–364 (2011).
Coombs, D.H. & Watts, N.R. Generating sucrose gradients in three minutes by tilted tube rotation. Anal. Biochem. 148, 254–259 (1985).
Suloway, C. et al. Automated molecular microscopy: the new Leginon system. J. Struct. Biol. 151, 41–60 (2005).
Lander, G.C. et al. Appion: an integrated, database-driven pipeline to facilitate EM image processing. J. Struct. Biol. 166, 95–102 (2009).
Mallick, S.P., Carragher, B., Potter, C.S. & Kriegman, D.J. ACE: automated CTF estimation. Ultramicroscopy 104, 8–29 (2005).
Scheres, S.H.W., Núñez-Ramírez, R., Sorzano, C.O.S., Carazo, J.M. & Marabini, R. Image processing for electron microscopy single-particle analysis using XMIPP. Nat. Protoc. 3, 977–990 (2008).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Wilcox, B.E., Hendrickson, C. & Marshall, A. Improved ion extraction from a linear octopole ion trap: SIMION analysis and experimental demonstration. J. Am. Soc. Mass Spectrom. 13, 1304–1312 (2002).
Beu, S.C. & Laude, D.A. Elimination of axial ejection during excitation with a capacitively coupled open trapped-ion cell for Fourier transform ion cyclotron resonance mass spectrometry. Anal. Chem. 64, 177–180 (1992).
Schwartz, J.C., Senko, M. & Syka, J. A two-dimensional quadrupole ion trap mass spectrometer. J. Am. Soc. Mass Spectrom. 13, 659–669 (2002).
Marshall, A.G. & Guan, S. Advantages of high magnetic field for Fourier transform ion cyclotron resonance mass spectrometry. Rapid Commun. Mass Spectrom. 10, 1819–1823 (1996).
Acknowledgements
This work was supported by the US National Institutes of Health through grant R01GM086892 (S.M.S.), the American Heart Association through grant 0835300N (S.M.S.), the US National Science Foundation Division of Materials Research through grant DMR-06-54118 (A.G.M.) and the State of Florida (A.G.M.). We thank W. Balch (Scripps Research Institute, La Jolla, California, USA) for providing the Sec13–31 expression construct. We thank C.L. Hendrickson and F. Xian for helpful discussions.
Author information
Authors and Affiliations
Contributions
J.O., H.H. and N.B. performed the specimen preparation and biochemistry. J.O. collected the cryo-EM data. S.M.S. performed the cryo-EM reconstructions. Q.Z. collected the MS data and analyzed it with the help of A.G.M. A.J.N. performed the modeling studies. A.J.N., Q.Z., A.G.M. and S.M.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3, 5, Supplementary Table 1 and Supplementary Note (PDF 2222 kb)
Supplementary Data set
Supplementary Figure 4a (PDF 908 kb)
Supplementary Data set
Supplementary Figure 4b (PDF 516 kb)
Supplementary Movie 1
12 Å reconstruction of the Sec13-31 COPII cage, radially colored. The human Sec13-31 homology model, colored by chain, is then docked into the density map. An asymmetric unit of the density map is segmented out and MDFF is applied to the homology model. Octahedral symmetry is imposed when introducing 23 additional copies of the Sec13-31 atomic model. The 24 Sec13-31 atomic models are again subjected to MDFF. Multiple views of the resulting atomic model are shown at a vertex, colored as in Figure 6. At the end of the movie, an individual flexibly fitted Sec13- 31 unit is shown. The two Sec13 chains are aligned, superimposed, and colored as in Figure 2. The resulting hinge with a range of 17° in the plane normal to the 4-fold axis is shown. (MOV 30685 kb)
Rights and permissions
About this article
Cite this article
Noble, A., Zhang, Q., O'Donnell, J. et al. A pseudoatomic model of the COPII cage obtained from cryo-electron microscopy and mass spectrometry. Nat Struct Mol Biol 20, 167–173 (2013). https://doi.org/10.1038/nsmb.2467
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2467